Volume 11, Number 5—May 2005
Dispatch
Human T-cell Leukemia Virus Type 1 Molecular Variants, Vanuatu, Melanesia
Figure A1
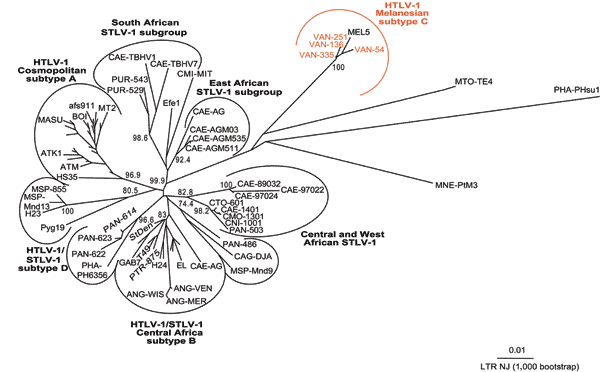
Figure A1. . Unrooted phylogenetic tree generated by the neighbor-joining method by using the complete fragment of the long terminal repeat (LTR) (755 bp). Distance matrices were generated with the DNADIST program, using the Kimura 2-parameter method and 5.17 as the transition/transversion ratio. Bootstrap analysis was carried out with 1,000 datasets. The values on the branches indicate frequencies of occurrence for 1,000 trees. The 4 new human T-cell leukemia virus type 1 (HTLV-1) sequences (VAN 54, VAN 136, VAN 251, VAN 335; GenBank Accession nos. AY549875, AY549876, AY549877, AY549878) were analyzed with 75 HTLV-1/STLV-1 sequences available from the GenBank database. The branch lengths are proportional to the evolutionary distance between the taxa.