Volume 11, Number 7—July 2005
Dispatch
Yersinia pseudotuberculosis Septicemia and HIV
Cite This Article
Citation for Media
Abstract
Two cases of community-acquired septicemia caused by serotype-O1 Yersinia pseudotuberculosis were diagnosed in middle-aged, HIV-positive, immunodeficient patients during an 8-month period. Bacterial isolates were genetically indistinguishable, but no epidemiologic link between the 2 patients was established. HIV-related immunosuppression should be regarded as a risk factor for Y. pseudotuberculosis septicemia.
Yersinia pseudotuberculosis is a rare cause of disease in humans. Animals, food, and the abiotic environment are Y. pseudotuberculosis reservoirs from which epizootic and human infection may arise (1). A geographic gradient of Y. pseudotuberculosis isolation rates has been reported in Europe (1,2), with a 0.05% recovery rate from stools of patients with acute enteritis in Italy (3). The organism can also cause mesenteric lymphadenitis, which mimics appendicitis, or infection at other body sites that occasionally leads to postinfectious sequelae such as reactive arthritis and erythema nodosum (1). About 60 cases of Y. pseudotuberculosis septicemia have been reported thus far, mainly in patients with underlying conditions such as hepatic cirrhosis, malignancy, diabetes, aplastic anemia, thalassemia, and iron overload (1,4,5). We recently reported the first case of Y. pseudotuberculosis septicemia in a severely immunocompromised, HIV-positive patient (6). Here, a second case of Y. pseudotuberculosis septicemia in an HIV-infected outpatient attending the same hospital is described. The unique molecular type of both Y. pseudotuberculosis isolates and the atypical clinical course of infection will be comparatively discussed.
The first case has recently been reported (6) and is briefly reviewed here. Patient 1 was a 42-year-old woman with HIV infection since 1987 (Centers for Disease Control and Prevention [CDC] Classification C3). In June 2003, she was admitted to the National Institute for Infectious Diseases, Rome, from prison because of high fever and confusion. Physical examination showed a temperature of 39.5°C, abnormal mental status, and oral candidiasis, but no gastrointestinal symptoms. Her history included lack of response to highly active antiretroviral therapy (HAART), and she exhibited HIV viremia of 413,624 copies/mL, a low CD4+ cell count (5/mm3), and leukopenia (3.0 × 103/mm3). Laboratory values were altered for aminotransferases (aspartate aminotransferase, 273 U/L; alanine aminotransferase, 77 U/L), hemoglobin (7.0 g/dL), erythrocyte sedimentation rate (136 mm in the first hour), and platelet count (34 × 103/mm3). Multiple blood cultures yielded growth of Y. pseudotuberculosis. Stool cultures and test results for antibodies against Y. pseudotuberculosis were negative. Intravenous ceftriaxone therapy was started at admission, with total remission of symptoms in 4 days. No recurrence of Y. pseudotuberculosis infection was observed during a 1-year follow-up period.
Patient 2 was a 54-year-old man who was admitted to the same hospital in February 2004 because of sudden fever and confusion. He tested HIV-positive in 1993 (CDC Classification B3), and his CD4+ cell count nadir was 121 cells/mm3. His recent antiretroviral therapy was a combination of stavudine, lamivudine, and nelfinavir. HCV-related liver cirrhosis was diagnosed 3 years before this admission. His condition was routinely followed up in the outpatient unit, and he had been hospitalized 1 month earlier for culture-negative pneumonia in the left lower lobe. On admission, the patient had a high fever (41°C), chills, and abnormal mental status with somnolence. Results of a chest radiograph, nuclear magnetic resonance imaging of the brain, and echocardiogram were normal. HIV viremia level was <50 copies/mL and CD4+ cell count was 204 cells/mm3. Laboratory values were notable for leukocytosis (leukocytes, 14 × 103/mm3), anemia (hemoglobin, 10 g/dL), and thrombocytopenia (platelets, 69 × 103/mm3). Levels of C reactive protein, aminotransferases, blood glucose, creatinine, urea nitrogen, and electrolytes were within the normal range. Liver cirrhosis was classified as Child-Pugh group B, score 7. Urine was negative for opioids and cocaine metabolites. On second day postadmission, multiple blood cultures became positive for Y. pseudotuberculosis, while stool cultures were negative. Intravenous ceftriaxone therapy was begun at admission, with total remission of symptoms within 3 days. He was discharged on day 10 and continued intravenous ceftriaxone therapy for 4 weeks (2 g daily) as an outpatient. At the final clinical observation, 6 months later, the patient was free of clinical symptoms and had no recurrence of Y. pseudotuberculosis infection.
Blood culture isolates from both patients were identified with >99% confidence as Y. pseudotuberculosis by using both the API20E-Rapid test (bioMérieux, Marcy l'Etoile, France) and the BD Phoenix system (Becton Dickinson, Sparks MD, USA). Isolates showed an extended spectrum of antimicrobial susceptibility (sensitive to β-lactams, monobactams, aminoglycosides, quinolones, and sulfonamides). O-serotyping with commercial antisera (Denka Seiken, Tokyo, Japan) identified both organisms as Y. pseudotuberculosis serotype O1, consistent with the high frequency of this serotype among strains isolated from patients with septicemia in Europe (2).
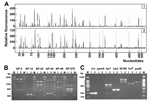
The 16S ribosomal DNA (rDNA) sequences were identical in the 2 isolates and optimally matched (>99% identity) the 16S rDNA of the Y. pseudotuberculosis type strain ATCC 29833 (GenBank accession no. AF366375). To differentiate the strains at the genome level, amplified fragment length polymorphism and arbitrarily primed polymerase chain reaction (PCR) were performed (7–9). While these methods are highly discriminatory and detect DNA polymorphisms to the strain level, no differences between the 2 clinical isolates were observed (Figure, panels A and B).
To gain insight into the pathogenic potential of the 2 isolates, we assessed the presence of pYV (70-kb virulence plasmid), high pathogenicity island (HPI), and Y. pseudotuberculosis–derived mitogen (YPM) genetic markers by PCR (2,6). Both isolates harbored a characteristic repertoire of virulence determinants. Correctly sized PCR products were obtained for inv (chromosome-borne), irp1, irp2, IS100 (HPI-borne), lcrF, and yadA (plasmidborne) genes (Figure, panel C). The identity of PCR products was confirmed by direct DNA sequencing. Interestingly, both isolates were negative for the ypmA (chromosome-borne) gene, consistent with the genetic instability of this marker which is typically absent in strains from Western countries (2,10). Reducing PCR stringency for ypmA yielded a 237-bp amplicon, which resulted from mispriming of oligonucleotides on the Y. pseudotuberculosis hmp flavohemoprotein gene (data not shown).
Molecular typing and virulence gene probing indicate that both cases resulted from indistinguishable Y. pseudotuberculosis strains, which raises the suspicion of a common source of infection. However, no other cases of Y. pseudotuberculosis infection were diagnosed in our institute or reported to the National Center for Enteropathogenic Bacteria, Istituto Superiore di Sanità, Rome, from January 2003 to June 2004 (I. Luzzi, pers. comm.). In September 2004, patients were interviewed about their lifestyle, illness, food and fluid consumption, or animal exposure in the 2 weeks preceding hospitalization. Nosocomial infection and direct contact between patients were ruled out, which suggested that infection had been independently acquired in the community. Both patients denied any contact with wild or domestic animals. Patient 1 had not been released from detention in the 6 months preceding hospitalization and regularly consumed collective meals in prison. However, prison infirmary records did not show an increased frequency of abdominal symptoms or fever among ≈500 inmates from May to July 2003. Patient 2 was a heavy smoker who had been abusing alcohol and illicit drugs until 2001. He lived alone and had a seafood meal 4 days before admission. Thus, although infection may have been acquired from contaminated food or fluid, questions regarding the actual source of bacteria and the extent of exposure remain unanswered.
AIDS is a known risk factor for Y. enterocolitica infection (11), but no link between HIV and Y. pseudotuberculosis infection has yet been proposed. Yersiniae have an impaired iron metabolism, and for this reason, they rarely cause sepsis in patients without iron overload, which is often secondary to alcoholism, asplenia, hemochromatosis, thalassemia major, or tobacco smoking (1). In our 2 patients, no clinical evidence indicated iron overload, and a diagnosis of hemochromatosis was excluded. The most important risk factor was HIV-related severe immunodeficiency. Patient 1 was severely immunocompromised despite HAART, while patient 2 responded poorly to HAART and had hepatitis C–related liver cirrhosis.
Although the clinical management of Y. pseudotuberculosis septicemia is often difficult and mortality rates are high (≈75%), despite antimicrobial drug therapy (1), both our patients responded unexpectedly well to ceftriaxone therapy and promptly recovered. During infection, Y. pseudotuberculosis directly manipulates lymphocyte signaling and activation by expressing different virulence factors (12). The pYV plasmid-encoded Yop proteins have been implicated in lymphocyte suppression through down-regulation of co-stimulatory molecules (13) and impairment of nitric oxide, tumor necrosis factor (TNF)-α, and proinflammatory cytokine production (14). Conversely, YPM superantigen(s) contribute to systemic illness by activating a large proportion of T cells (essentially CD4+) and inducing proinflammatory cytokines such as TNF-α, TNF-β, γ-interferon, and interleukins 1 and 6, as in toxic shock syndrome (15). Our experimental data show that the 2 clinical strains were positive for all the virulence genes tested, except for ypmA. Even considering the different degree of immunodeficiency between the 2 patients, we speculate that impairment of immune response secondary to HIV infection may have increased the susceptibility to Y. pseudotuberculosis infection while mitigating the septic shock sequelae. Accordingly, the inflammatory response consequent to T-cell activation may have been attenuated by the deficiency of CD4+ cells in both patients, concomitant with the lack of YPM expression by both Y. pseudotuberculosis isolates. In conclusion, these 2 cases indicate that Y. pseudotuberculosis is an emerging pathogen in HIV patients and remind us that septicemia in these patients can exist without prodromic symptoms. They also alert us to the local circulation of a pathogenic Y. pseudotuberculosis strain whose natural reservoir remains so far unknown.
Dr. Paglia is senior scientist at the National Institute for Infectious Diseases "Lazzaro Spallanzani," Rome, Italy. Her research interests include the development of DNA-based microbial identification tools and molecular epidemiology.
Acknowledgments
We are grateful to E. Nebuloso for technical assistance and to P. Marconi and L. Alba for clinical contributions.
This work was supported by grants from Ricerca Corrente and Ricerca Finalizzata from the Italian Ministry of Health to A.A., A.F., and P.V.
References
- Butler T. Yersinia species, including plague. In: Mandell GL, Douglas RG, Bennett JE, editors. Principles and practice of infectious diseases. 5th ed. New York: Churchill Livingstone; 2000. p. 2406–14.
- Fukushima H, Matsuda Y, Seki R, Tsubokura M, Takeda N, Shubin FN, Geographical heterogeneity between Far Eastern and Western countries in prevalence of the virulence plasmid, the superantigen Yersinia pseudotuberculosis–derived mitogen, and the high pathogenicity island among Yersinia pseudotuberculosis strains. J Clin Microbiol. 2001;39:3541–7. DOIPubMedGoogle Scholar
- Chiesa C, Pacifico L, Nanni F, Renzi AM, Ravagnan G. Yersinia pseudotuberculosis in Italy. Attempted recovery from 37,666 samples. Microbiol Immunol. 1993;37:391–4.PubMedGoogle Scholar
- Ljungberg P, Valtonen M, Harjola VP, Kaukoranta-Tolvanen SS, Vaara M. Report of four cases of Yersinia pseudotuberculosis septicemia and a literature review. Eur J Clin Microbiol Infect Dis. 1995;14:804–10. DOIPubMedGoogle Scholar
- Deacon AG, Hay A, Duncan J. Septicemia due to Yersinia pseudotuberculosis—a case report. Clin Microbiol Infect. 2003;9:1118–9. DOIPubMedGoogle Scholar
- Antinori A, Paglia MG, Marconi P, Festa A, Alba L, Boumis E, Short communication: Yersinia pseudotuberculosis septicemia in an HIV-infected patient failed HAART. AIDS Res Hum Retroviruses. 2004;20:709–10. DOIPubMedGoogle Scholar
- Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res. 1995;23:4407–14. DOIPubMedGoogle Scholar
- Makino S, Okada Y, Maruyama T, Kaneko S, Sasakawa C. PCR-based random amplified polymorphic DNA fingerprinting of Yersinia pseudotuberculosis and its practical applications. J Clin Microbiol. 1994;32:65–9.PubMedGoogle Scholar
- Tabacchioni S, Visca P, Chiarini L, Bevivino A, Di Serio C, Fancelli S, Molecular characterization of rhizosphere and clinical isolates of Burkholderia cepacia. Res Microbiol. 1995;146:531–42. DOIPubMedGoogle Scholar
- Carnoy C, Floquet S, Marceau M, Sebbane F, Haentjens-Herwegh S, Devalckenaere A, The superantigen gene ypm is located in an unstable chromosomal locus of Yersinia pseudotuberculosis. J Bacteriol. 2002;184:4489–99. DOIPubMedGoogle Scholar
- Cannon CG, Linnemann CC Jr. Yersinia enterocolitica infections in hospitalized patients: the problem of hospital-acquired infections. Infect Control Hosp Epidemiol. 1992;13:139–43. DOIPubMedGoogle Scholar
- Cornelis GR. Yersinia type III secretion: send in the effectors. J Cell Biol. 2002;158:401–8. DOIPubMedGoogle Scholar
- Yao T, Mecsas J, Healy JI, Falkow S, Chien Y. Suppression of T and B lymphocyte activation by a Yersinia pseudotuberculosis virulence factor, yopH. J Exp Med. 1999;190:1343–50. DOIPubMedGoogle Scholar
- Monnazzi LG, Carlos IZ, de Medeiros BM. Influence of Yersinia pseudotuberculosis outer proteins (Yops) on interleukin-12, tumor necrosis factor alpha and nitric oxide production by peritoneal macrophages. Immunol Lett. 2004;94:91–8. DOIPubMedGoogle Scholar
- Abe J, Onimaru M, Matsumoto S, Noma S, Baba K, Ito Y, Clinical role for a superantigen in Yersinia pseudotuberculosis infection. J Clin Invest. 1997;99:1823–30. DOIPubMedGoogle Scholar
Figure
Cite This ArticleTable of Contents – Volume 11, Number 7—July 2005
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Paolo Visca, Molecular Microbiology Unit, National Institute for Infectious Diseases "Lazzaro Spallanzani" I.R.C.C.S., Via Portuense 292, 00149 Rome, Italy; fax: 39-6-55176321
Top