Volume 13, Number 9—September 2007
Dispatch
Hantavirus in Northern Short-tailed Shrew, United States
Cite This Article
Citation for Media
Abstract
Phylogenetic analyses, based on partial medium- and large-segment sequences, support an ancient evolutionary origin of a genetically distinct hantavirus detected by reverse transcription–PCR in tissues of northern short-tailed shrews (Blarina brevicauda) captured in Minnesota in August 1998. To our knowledge, this is the first evidence of hantaviruses harbored by shrews in the Americas.
Rodents and their ectoparasites serve as reservoirs and vectors of myriad viruses and other pathogenic microbes. In contrast, the role of insectivores (or soricomorphs) in the transmission and ecology of zoonoses is largely unknown. Because some soricomorphs share habitats with rodents, shrews might also be involved in the maintenance of the enzootic cycle and contribute to the evolutionary history and genetic diversity of hantaviruses.
Hantavirus antigens have been detected in the Eurasian common shrew (Sorex araneus), alpine shrew (Sorex alpinus), Eurasian water shrew (Neomys fodiens), and common mole (Talpa europea) in Russia and the former Yugoslavia (1–3). More than 20 years ago, when Prospect Hill virus was discovered in meadow voles (Microtus pennsylvanicus) captured in Frederick, Maryland, USA, serologic evidence suggestive of hantavirus infection was found in the northern short-tailed shrew (Blarina brevicauda) (4). However, virus isolation attempts were unsuccessful, and molecular tools such as PCR were unavailable. Empowered by robust gene-amplification techniques and the complete genome of Thottapalayam virus (TPMV) isolated from the Asian house shrew (Suncus murinus) (5,6), we have identified a genetically distinct hantavirus in the northern short-tailed shrew.
After obtaining approval from the University of Hawaii Institutional Animal Care and Use Committee, we retrieved lung and liver tissues of 30 northern short-tailed shrews captured from several regions within the United States during 1994–1999 (Table 1) from deep-freeze storage at the University of New Mexico Museum of Southwestern Biology. Total RNA was extracted from shrew tissues by using the PureLink Micro-to-Midi Total RNA Purification Kit (Invitrogen, San Diego, CA, USA). cDNA was then prepared by using the SuperScript III First-Strand Synthesis System (Invitrogen) for reverse transcription–PCR (RT-PCR) with oligonucleotide primers designed from TPMV and other hantaviruses: medium (M) (outer: 5′-GGACCAGGTGCADCTTGTGAAGC-3′, 5′-GAACCCCADGCCCCNTCYAT-3′; inner: 5′-TAAVTTCAMCAACATGTCT-3′, 5′-CATGAYATCTCCAGGGTCHCC-3′) and large (L) (outer: 5′-CAGTCWACARTTGGTGCAAGTGG-3′, 5′-TCCATKATWGACATBGMRCCA-3′; inner: 5′-YTMATGTATGTTAGTGCAGATGC-3′, 5′-GRTTAAACATACTCTTCCACATCTC-3′). For confirmation, RNA extraction and RT-PCR were performed independently in a laboratory in which hantaviruses had never been handled. Amplicons were sequenced directly by using an ABI Prism 377XL Genetic Analyzer (Applied Biosystems, Foster City, CA, USA).
Of the 30 northern short-tailed shrews tested, hantavirus M-segment sequences were amplified from lung tissues of 3 of 12 animals captured in Camp Ripley (46.185°N, 94.4337°W), a 53,000-acre, state-owned military and civilian training center near Little Falls, in Morrison County, Minnesota, USA, in August 1998 (Table 1). Pairwise alignment and comparison of the 1,390-nt region (463 aa) spanning the Gn and Gc glycoprotein–encoding M segment indicated differences of 33.6%–41.9% and 32.7%–47.4% at the nucleotide and amino acid levels, respectively, from representative hantaviruses harbored by Murinae, Arvicolinae, Neotominae, and Sigmodontinae rodents (Table 2). No insertions or deletions were found in the regions sequenced compared with sequences of other hantaviruses. The new hantavirus, designated Camp Ripley virus (RPLV), showed sequence similarity of 98.1%–98.5% among the 3 strains.
Analysis of a 490-nt (163-aa) region of the L genomic segment amplified from 2 of the 3 shrews indicated that RPLV differed from rodentborne hantaviruses by 26.8%–34.1% at the nucleotide level and 20.7%–34.5% at the amino acid level (Table 2). RPLV differed from Tanganya virus (TGNV), a hantavirus detected recently in the Therese shrew (Crocidura theresae) in Guinea (7), by 27.3%–28.8% and 24.0%–25.0%, respectively. The higher degree of sequence similarity in the L segment between RPLV and other hantaviruses probably signifies the limits of functional preservation of the RNA-dependent RNA polymerase.
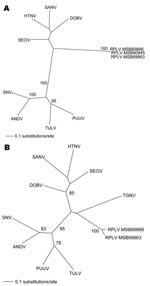
Repeated and exhaustive phylogenetic analyses based on nucleotide and deduced amino acid sequences of the M and L genomic segments generated by the maximum-likelihood method indicated that RPLV was distinct from rodentborne hantaviruses (with high bootstrap support based on 100 maximum likelihood replicates) (Figure). Similar topologies were consistently derived by using various algorithms and different taxa (including La Crosse virus) and combinations of taxa, which suggested an ancient evolutionary origin. However, definitive conclusions about the molecular phylogeny of RPLV and its relationship to TGNV and other soricidborne hantaviruses must await complete-genome sequence analyses.
As we had previously encountered in sequencing the entire genome of TPMV (J.-W. Song, R. Yanagihara, unpub. data), the divergent genome of RPLV presented challenges in designing suitable primers for RT-PCR. We were also constrained by the limited availability of tissues from the 3 infected shrews and the need to retain small portions of tissues for future virus isolation attempts. Consequently, we have been hitherto unable to obtain the full-length sequence of RPLV.
The northern short-tailed shrew (family Soricidae, subfamily Soricinae), 1 of 2 poisonous mammals in North America (8), inhabits forests and grasslands within the central and eastern half of the United States, extending north to Canada, west to Montana, and south to Tennessee and Georgia. Cytochrome b mitochondrial DNA and 16S rRNA sequence analyses support a monophyletic origin for the genus Blarina, with phylogeographic structuring of northern short-tailed shrews into well-defined groups to the east and west of the Mississippi River basin (9). Current studies will examine whether RPLV is harbored by the eastern haplogroup of northern short-tailed shrews and by the southern short-tailed shrew (Blarina carolinensis), a closely related species, which inhabits the southeastern United States, extending as far north as southern Illinois and south-central Virginia and as far south as central Florida.
Given the sympatric and synchronistic coexistence of northern short-tailed shrews with Neotominae and Arvicolinae rodents (such as Peromyscus leucopus and Microtus pennsylvanicus) and their ferocious territorial behavior, hantavirus spillover may be possible. Viruses closely related antigenically to Hantaan virus have been isolated from the Asian house shrew (Suncus murinus), greater white-toothed shrew (Crocidura russula), and Chinese mole shrew (Anourosorex squamipes) (10–12), which suggests that shrews are capable of serving as incidental hosts for hantaviruses typically harbored by rodents.
Shrews that harbor genetically distinct hantaviruses pose a compelling conceptual framework that challenges the long-accepted dogma that rodents are the sole reservoirs of hantaviruses. Viewed within the context of the recent detection of TGNV in the Therese shrew in Guinea, the identification of RPLV in the northern short-tailed shrew in the United States indicates that renewed efforts, facilitated by the rapidly expanding sequence database of shrew-borne hantaviruses, will lead to the discovery of additional hantaviruses in soricids throughout Eurasia, Africa, and the Americas. Our preliminary studies indicate 3 other novel soricid-borne hantaviruses in the Republic of Korea, Vietnam, and Switzerland. To establish if >1 of these newly identified hantaviruses is pathogenic for humans will require development of robust serologic assays (13) and application of other sensitive technologies, such as microarray analysis (14,15), for rapid detection of shrewborne hantavirus RNA in human tissues and bodily fluids.
Dr Arai is a researcher at the Infectious Disease Surveillance Center of the National Institute of Infectious Diseases in Tokyo, Japan. He is currently a guest researcher at the University of Hawaii at Manoa. His research interests include the epizootiology and prevention of vectorborne and zoonotic diseases.
Acknowledgments
We thank the staff of the Greenwood Molecular Biology Facility, Pacific Biosciences Research Center, for excellent technical assistance with DNA sequencing.
This work was supported in part by grants P20RR018727 (Centers of Biomedical Research Excellence) and G12RR003061 (Research Centers in Minority Institutions) from the National Center for Research Resources, National Institutes of Health, Bethesda, MD, USA.
References
- Gavrilovskaya IN, Apekina NS, Myasnikov YA, Bernshtein AD, Ryltseva EV, Gorbachkova EA, Features of circulation of hemorrhagic fever with renal syndrome (HFRS) virus among small mammals in the European U.S.S.R. Arch Virol. 1983;75:313–6. DOIPubMedGoogle Scholar
- Tkachenko EA, Ivanov AP, Donets MA, Miasnikov YA, Ryltseva EV, Gaponova LK, Potential reservoir and vectors of haemorrhagic fever with renal syndrome (HFRS) in the U.S.S.R. Ann Soc Belg Med Trop. 1983;63:267–9.PubMedGoogle Scholar
- Gligic A, Stojanovic R, Obradovic M, Hlaca D, Dimkovic N, Diglisic G, Hemorrhagic fever with renal syndrome in Yugoslavia: epidemiologic and epizootiologic features of a nationwide outbreak in 1989. Eur J Epidemiol. 1992;8:816–25. DOIPubMedGoogle Scholar
- Lee PW, Amyx HL, Yanagihara R, Gajdusek DC, Goldgaber D, Gibbs CJ Jr. Partial characterization of Prospect Hill virus isolated from meadow voles in the United States. J Infect Dis. 1985;152:826–9.PubMedGoogle Scholar
- Carey DE, Reuben R, Panicker KN, Shope RE, Myers RM. Thottapalayam virus: a presumptive arbovirus isolated from a shrew in India. Indian J Med Res. 1971;59:1758–60.PubMedGoogle Scholar
- Zeller HG, Karabatsos N, Calisher CH, Digoutte J-P, Cropp CB, Murphy FA, Electron microscopic and antigenic studies of uncharacterized viruses. II. Evidence suggesting the placement of viruses in the family Bunyaviridae. Arch Virol. 1989;108:211–27. DOIPubMedGoogle Scholar
- Klempa B, Fichet-Calvet E, Lecompte E, Auste B, Aniskin V, Meisel H, Novel hantavirus sequences in shrew, Guinea. Emerg Infect Dis. 2007;13:520–2. DOIPubMedGoogle Scholar
- Kita M, Nakamura Y, Okumura Y, Ohdachi SD, Oba Y, Yoshikuni M, Blarina toxin, a mammalian lethal venom from the short-tailed shrew Blarina brevicauda: isolation and characterization. Proc Natl Acad Sci U S A. 2004;101:7542–7. DOIPubMedGoogle Scholar
- Brant SV, Ortı G. Molecular phylogeny of short-tailed shrews, Blarina (Insectivora: Soricidae). Mol Phylogenet Evol. 2002;22:163–73. DOIPubMedGoogle Scholar
- Tang YW, Xu ZY, Zhu ZY, Tsai TF. Isolation of haemorrhagic fever with renal syndrome virus from Suncus murinus, an insectivore. Lancet. 1985;1:513–4. DOIPubMedGoogle Scholar
- Yan DY, Xie YJ, Zhang CA, McCormick JB, Sanchez A, Engelman HM, New isolates of HFRS virus in Sichuan, China, and characterization of antigenic differences by monoclonal antibodies. Lancet. 1986;1:1328. DOIPubMedGoogle Scholar
- Tang YW, Ruo SL, Sanchez A, Fisher-Hoch SP, McCormick JB, Xu ZY. Hantavirus strains isolated from rodentia and insectivora in rural China differentiated by polymerase chain reaction. Arch Virol. 1990;115:37–46. DOIPubMedGoogle Scholar
- Okumura M, Yoshimatsu K, Kumperasart S, Nakamura I, Ogino M, Taruishi M, Development of serological assays for Thottapalayam virus, an insectivore-borne hantavirus. Clin Vaccine Immunol. 2007;14:173–81. DOIPubMedGoogle Scholar
- Wang D, Coscoy L, Zylberberg M, Avila PC, Boushey HA, Ganem D, Microarray-based detection and genotyping of viral pathogens. Proc Natl Acad Sci U S A. 2002;99:15687–92. DOIPubMedGoogle Scholar
- Nordström H, Johansson P, Li Q-G, Lundkvist A, Nilsson P, Elgh F. Microarray technology for identification and distinction of hantaviruses. J Med Virol. 2004;72:646–55. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 13, Number 9—September 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Richard Yanagihara, Pacific Center for Emerging Infectious Diseases Research, John A. Burns School of Medicine, University of Hawaii at Manoa, 651 Ilalo St, BSB320L, Honolulu, HI 96813, USA:
Top