Volume 16, Number 9—September 2010
CME ACTIVITY - Synopsis
Recurrent Granulibacter bethesdensis Infections and Chronic Granulomatous Disease
Figure 3
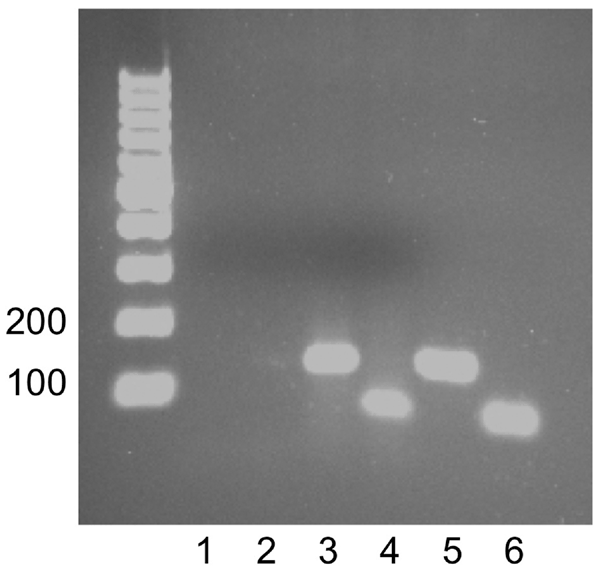
Figure 3. Detection of Granulibacter sp. DNA by PCR of the spleen of a patient with chronic granulomatous disease. Lanes 1 and 2, no template controls for each primer set; lane 3, 400 ng of spleen DNA amplifying Granulibacter sp. 16S rRNA gene; lane 4, 400 ng of spleen DNA amplifying the Granulibacter sp. methanol dehydrogenase gene; lane 5, 100 ng DNA from the G. bethesdensis type strain amplifying the 16S rRNA gene (positive control); lane 6, 100 ng DNA from the G. bethesdensis type strain amplifying the methanol dehydrogenase gene (positive control). Left lane, molecular mass ladder. Expected PCR product sizes were 137 bp for the 16S rRNA gene and 63 bp for the methanol dehydrogenase gene. Values on the left are in bp.
Page created: August 28, 2011
Page updated: August 28, 2011
Page reviewed: August 28, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.