Volume 17, Number 6—June 2011
Dispatch
Novel Reassortant Highly Pathogenic Avian Influenza (H5N5) Viruses in Domestic Ducks, China
Cite This Article
Citation for Media
Abstract
In China, domestic ducks and wild birds often share the same water, in which influenza viruses replicate preferentially. Isolation of 2 novel reassortant highly pathogenic avian influenza (H5N5) viruses from apparently healthy domestic ducks highlights the role of these ducks as reassortment vessels. Such new subtypes of influenza viruses may pose a pandemic threat.
Aquatic birds are considered the natural reservoir for influenza A viruses of all known 16 hemagglutinin (HA) and 9 neuraminidase (NA) subtypes (1). Influenza viruses replicate preferentially in the cells lining the intestinal tracts of wild waterfowl, usually causing no clinical signs. Excretion of substantial amounts of virus in the feces can infect wild and domestic birds by waterborne transmission (1). In the People’s Republic of China, domestic ducks raised in the traditional free-range system often share water with wild aquatic birds. Moreover, domestic ducks are often in close contact with poultry, livestock, and humans in the same village or farm. Therefore, domestic ducks play a major role in the ecology of influenza viruses (2) and can act as potential vessels for genetic reassortment (3). Systematic surveillance of influenza viruses in domestic ducks could provide timely and valuable epidemiologic information and should be continued.
As part of routine surveillance for avian influenza viruses from December 2008 through January 2009 in eastern China, tracheal and cloacal swab samples from apparently healthy domestic ducks in live poultry markets were collected for virus isolation and identification as described (4). From these samples, 2 influenza (H5N5) viruses—A/duck/eastern China/008/2008 (008 [H5N5]) and A/duck/eastern China/031/2009 (031 [H5N5])—were detected in mallard ducks (Anas platyrhynchos).
These 2 viruses grew efficiently in eggs and in MDCK cells, each with virus titers >8 log10 50% egg infectious dose (EID50)/mL or 8 log10 50% tissue culture infectious dose/mL (Table 1). The intravenous pathogenicity index for chickens and 50% lethal dose for mice were 2.6 and 104.0 EID50 for 008 (H5N5), 2.5 and 105.4 EID50 for 031 (H5N5), respectively (Table 1). Therefore, both novel influenza subtype H5N5 viruses were assumed to be highly pathogenic for chickens and moderately virulent for mice (5).
When mice were inoculated with a sublethal dose of 103.5 EID50, each influenza subtype H5N5 virus was able to replicate without prior adaptation. The highest virus titers were detected in the mouse lung. The viruses were able to spread to the brain and heart. Furthermore, influenza virus 008 (H5N5) was isolated from the spleen and liver, and influenza virus 031 (H5N5) was detected in the kidney (Table 1). Microscopic findings in infected mice were interstitial pneumonia with various amounts of erythrocytes in alveolar lumens, hyperemia and lymphocyte infiltration of meningeal veins and cardiac muscles, low numbers of lymphocytes in periarterial lymphatic sheaths and macrophage recruitment in the spleen, lymphocyte infiltration in the liver, and slight congestion in the renal cortex and glomerulus (data not shown).
Genomic analysis showed that the influenza viruses 008 (H5N5) and 031 (H5N5) were highly homologous with each other, sharing 99.2%–99.7% nt identities among the 8 gene segments except for the polymerase acidic protein (PA) gene (94.1%). Another 3 viruses isolated from the same surveillance study— A/duck/eastern China/108/2008 of H5N1 subtype (108 [H5N1]), A/duck/eastern China/909/2009 of H5N1 subtype (909 [H5N1]), and A/duck/Yangzhou/013/2008 of H6N5 subtype (013 [H6N5])—were more closely related to the novel influenza subtype H5N5 viruses than were those in GenBank (Table 2).
As outlined by the World Health Organization/World Organisation for Animal Health/Food and Agriculture Organization unified nomenclature system for subtype H5N1 highly pathogenic avian influenza (HPAI) viruses (6), the HA genes of the 2 subtype H5N5 viruses were classified into clade 2.3.4 (Figure 1, panel A), which has been the prevalent lineage in southern China since 2005 (7–9). In addition to the typical residues Q226 and G228 in HA, which confer receptor preference for SAα2,3Gal, influenza viruses 008 (H5N5) and 031 (H5N5) simultaneously carried an S227R mutation in the receptor-binding pocket. A recent report (10) indicated that the S227N substitution accompanied with deglycosylation at residue 158 could substantially increase the affinity of HA for SAα2,6Gal without reducing its binding affinity for SAα2,3Gal. Whether this S227R variation with the changed residual polarity affects the receptor-binding property deserves further investigation.
To construct the NA tree, we retrieved 90 complete N5 sequences from GenBank, including the only 2 influenza subtype H5N5 viruses from the United States: A/mallard/MN/105/2000 and A/duck/Massachusetts/sg-00440/2005. The N5 viruses were grouped into 2 lineages—North American and Eurasian—in accordance with their geographic distribution (Figure 1, panel B). The 2 subtype H5N5 isolates from China belonged to the Eurasian lineage, whereas the 2 from the United States clustered within the North American lineage. In addition, 1 aa deletion at residue 42, located in the stalk region of NA, was identified in all Eurasian, but not in the North American, strains.
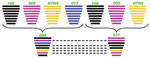
Although the PA genes of influenza viruses 008 (H5N5) and 031 (H5N5) diverged to assemble respectively with influenza viruses 013 (H6N5) and 108 (H5N1), the novel influenza subtype H5N5 viruses aggregated closely with recent Eurasian subtype H5N1 viruses, especially influenza viruses 108 (H5N1) and 909 (H5N1) in the trees of HA (Figure 1, panel A), polymerase basic protein (PB) 2, PB1, nucleocapsid protein, matrix protein, and nonstructural protein genes (Technical Appendix). For NA, spatiotemporal correlation indicates that influenza virus 013 (H6N5), rather than its phylogenetically equidistant counterpart A/mallard/Switzerland/WV4060167/2006(H3N5), might be the N5 donor (Figure 1, panel B). However, because of the relatively low (95%) sequence similarity, it is also possible that Eurasian viruses of an unidentified HA subtype (H?N5), not detected in our epidemiologic survey, could provide the NA genes. Therefore, we speculate that influenza viruses 008 (H5N5) and 031 (H5N5) may be reassortants between contemporary Eurasian subtype H5N1 and some subtype H?N5 and/or H6N5 avian influenza viruses with distant evolutionary relationship with the 2 subtype H5N5 viruses from the United States (Figure 1; Technical Appendix). In addition, regarding the especially high sequence identities of PB1, HA, nucleocapside protein, NA, and matrix protein genes exclusively between influenza viruses 008 (H5N5) and 031 (H5N5), the possibility that the 2 subtype H5N5 viruses donated some gene segments to each other cannot be excluded (Figure 2).
The 2 novel HPAI (H5N5) viruses isolated and characterized in this study are most likely reassortants of recent Eurasian viruses sharing approximate spatiotemporal distribution. It is less likely that they were introduced through intercontinental transmission of subtype H5N5 strains from North America. Considering the endemicity that clade 2.3.4 subtype H5N1 viruses have gained in China since 2005 (7–9), it is plausible that subtype H5N1 viruses have provided the backbone for generating the novel subtype H5N5 viruses instead of the opposite gene flow.
Ducks have been considered “Trojan horses” for influenza (H5N1) because of their pivotal role in virus propagation and evolution (11–13). In our study, the 2 reassortant influenza viruses (008 [H5N5] and 031 [H5N5]) and their 3 possible parent viruses (108 [H5N1], 909 [H5N1], and 013 [H6N5]) were all isolated from apparently healthy domestic ducks. We speculate that domestic ducks may serve as reassortant vessels for creating new subtypes of influenza viruses. In view of the practice of raising ducks in a free-range system, these novel strains could be transmitted to other domestic poultry and even humans. There is evidence that these subtype H5N5 viruses have been transmitted to terrestrial poultry (Zhao et al., unpub. data). Thus, the role of domestic ducks in the influenza virus ecosystem should not be neglected. Systematic surveillance should be instituted to identify emerging HPAI (H5N5) viruses and to reduce their potential threat to animal and human health.
Acknowledgment
The study was supported by Major State Basic Research Development Program of China (973 Program) (grant nos. 2011CB505001 and 2011CB505003), Important National Science & Technology Specific Projects (grant no. 2008ZX10004-013), and the Earmarked Fund for Modern Agro-Industry Technology Research System (nycytx-41-G07).
References
- Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y. Evolution and ecology of influenza A viruses. Microbiol Rev. 1992;56:152–79.PubMedGoogle Scholar
- Huang K, Bahl J, Fan XH, Vijaykrishna D, Cheung CL, Webby RJ, Establishment of an H6N2 influenza virus lineage in domestic ducks in southern China. J Virol. 2010;84:6978–86. DOIPubMedGoogle Scholar
- Kim HR, Park CK, Oem JK, Bae YC, Choi JG, Lee OS, Characterization of H5N2 influenza viruses isolated in South Korea and their influence on the emergence of a novel H9N2 influenza virus. J Gen Virol. 2010;91:1978–83. DOIPubMedGoogle Scholar
- Zhang P, Tang Y, Liu X, Peng D, Liu W, Liu H, Characterization of H9N2 influenza viruses isolated from vaccinated flocks in an integrated broiler chicken operation in eastern China during a 5-year period (1998–2002). J Gen Virol. 2008;89:3102–12. DOIPubMedGoogle Scholar
- Katz JM, Lu X, Tumpey TM, Smith CB, Shaw MW, Subbarao K. Molecular correlates of influenza A H5N1 virus pathogenesis in mice. J Virol. 2000;74:10807–10. DOIPubMedGoogle Scholar
- WHO/OIE/FAO H5N1 Evolution Working Group. Toward a unified nomenclature system for highly pathogenic avian influenza virus (H5N1). Emerg Infect Dis. 2008;14:e1.PubMedGoogle Scholar
- Chen H, Smith GJ, Zhang SY, Qin K, Wang J, Li KS, Avian flu: H5N1 virus outbreak in migratory waterfowl. Nature. 2005;436:191–2. DOIPubMedGoogle Scholar
- Smith GJ, Fan XH, Wang J, Li KS, Qin K, Zhang JX, Emergence and predominance of an H5N1 influenza variant in China. Proc Natl Acad Sci U S A. 2006;103:16936–41. DOIPubMedGoogle Scholar
- Smith GJ, Vijaykrishna D, Ellis TM, Dyrting KC, Leung YH, Bahl J, Characterization of avian influenza viruses A (H5N1) from wild birds, Hong Kong, 2004–2008. Emerg Infect Dis. 2009;15:402–7. DOIPubMedGoogle Scholar
- Yen HL, Aldridge JR, Boon AC, Ilyushina NA, Salomon R, Hulse-Post DJ, Changes in H5N1 influenza virus hemagglutinin receptor binding domain affect systemic spread. Proc Natl Acad Sci U S A. 2009;106:286–91. DOIPubMedGoogle Scholar
- Hulse-Post DJ, Sturm-Ramirez KM, Humberd J, Seiler P, Govorkova EA, Krauss S, Role of domestic ducks in the propagation and biological evolution of highly pathogenic H5N1 influenza viruses in Asia. Proc Natl Acad Sci U S A. 2005;102:10682–7. DOIPubMedGoogle Scholar
- Kim JK, Negovetich NJ, Forrest HL, Webster RG. Ducks: the “Trojan horses” of H5N1 influenza. Influenza Other Respir Viruses. 2009;3:121–8. DOIPubMedGoogle Scholar
- Takakuwa H, Yamashiro T, Le MQ, Phuong LS, Ozaki H, Tsunekuni R, Possible circulation of H5N1 avian influenza viruses in healthy ducks on farms in northern Vietnam. Microbiol Immunol. 2010;54:58–62. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 17, Number 6—June 2011
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Xiufan Liu, Animal Infectious Disease Laboratory, College of Veterinary Medicine, Yangzhou University, 48 East Wenhuei Rd, Yangzhou, Jiangsu, 225009, People’s Republic of China
Top