Volume 18, Number 12—December 2012
Letter
Hepatitis E Virus Genotype 3 in Shellfish, United Kingdom
Figure
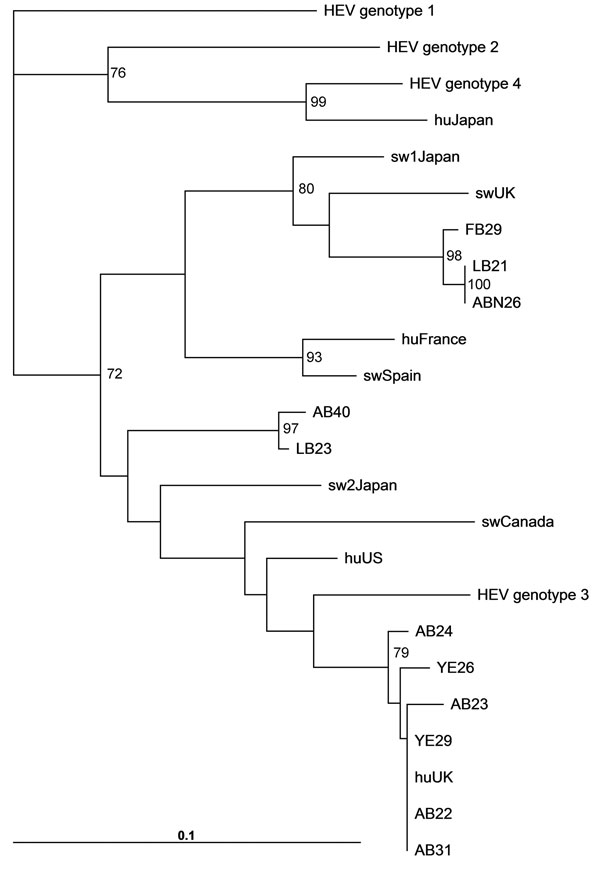
Figure. . Phylogenetic analysis of HEV open reading frame 2 sequences isolated from Mytilus spp. RNA was isolated from 50–100 mg of digestive gland or gill. Tissue was homogenized in 300 μL phosphate-buffered saline, and viral RNA was isolated by using a viral RNA kit (QIAGEN, Crawley, UK), and PCR was conducted by amplifying nucleotides 6332–6476 as described (8). The nucleotide sequences were aligned and bootstrapped, and phylogenetic neighbor-joining trees were constructed by using the ClustalW software (www.ebi.ac.uk/Tools/msa/clustalw2/). Phylogenetic trees were visualized by using FigTree (http://tree.bio.ed.ac.uk/software/figtree/). Bootstrap values >70% are indicated. Scale bar indicates nucleotide substitutions per site. Sample site codes: AB, Ardrossan Beach; LB, Lunderston Bay; ABN, Aberdeen; FB, Ferrybridge; YE, Ythan Estuary. Sequences: Sw, swine; hu, human (followed by country of origin). GenBank accession numbers for reference sequences: HEV genotype 1, B73218; HEV genotype 2, M74506; HEV genotype 3, CO31008; HEV genotype 4, C272108; huUK (KernowC1), HQ389543; HuUS, JN837481; swUK AF503512; huFrance, JN906974; swCanada, AY115488; swSpain, JQ522948; sw2Japan AB248521, huJapan AB161719.
References
- Kamar N, Bendall R, Legrand-Abravanel F, Xia NS, Ijaz S, Izopet J, Hepatitis E. Lancet. 2012;379:2477–88. DOIPubMedGoogle Scholar
- Said B, Ijaz S, Kafatos G, Booth L, Thomas HL, Walsh A, Hepatitis E outbreak on cruise ship. Emerg Infect Dis. 2009;15:1738–44. DOIPubMedGoogle Scholar
- McCreary C, Martelli F, Grierson S, Ostanello F, Nevel A, Banks M. Excretion of hepatitis E virus by pigs of different ages and its presence in slurry stores in the United Kingdom. Vet Rec. 2008;163:261–5. DOIPubMedGoogle Scholar
- Emerson SU, Arankalle VA, Purcell RH. Thermal stability of hepatitis E virus. J Infect Dis. 2005;192:930–3. DOIPubMedGoogle Scholar
- Lowther JA, Gustar NE, Powell AL, Hartnell RE, Lees DN. A two-year systematic study to assess norovirus contamination in oysters from commercial harvesting areas in the United Kingdom. Appl Environ Microbiol. 2012;78:5812–7. DOIPubMedGoogle Scholar
- Baker K, Morris J, McCarthy N, Saldana L, Lowther J, Collinson A, An outbreak of norovirus infection linked to oyster consumption at a UK restaurant, February 2010. J Public Health (Oxf). 2011;33:205–11. DOIPubMedGoogle Scholar
- McLeod C, Hay B, Grant C, Greening G, Day D. Inactivation and elimination of human enteric viruses by Pacific oysters. J Appl Microbiol. 2009;107:1809–18. DOIPubMedGoogle Scholar
- Erker JC, Desai SM, Mushahwar IK. Rapid detection of hepatitis E virus RNA by reverse transcription–polymerase chain reaction using universal oligonucleotide primers. J Virol Methods. 1999;81:109–13. DOIPubMedGoogle Scholar
- Li TC, Miyamura T, Takeda N. Detection of hepatitis E virus RNA from the bivalve Yamato-Shijimi (Corbicula japonica) in Japan. Am J Trop Med Hyg. 2007;76:170–2 .PubMedGoogle Scholar
- Shuval H. Estimating the global burden of thalassogenic diseases: human infectious diseases caused by wastewater pollution of the marine environment. J Water Health. 2003;1:53–64 .PubMedGoogle Scholar