Volume 18, Number 4—April 2012
Letter
Methicillin-Susceptible Staphylococcus aureus ST398, New York and New Jersey, USA
Cite This Article
Citation for Media
To the Editor: Clinical infections with livestock-associated Staphylococcus aureus sequence type (ST) 398 have been reported in Europe, Canada, and the People’s Republic of China (1), as well as the Caribbean (2,3), and Colombia (4). Most reports describe infection with methicillin-resistant S. aureus; relatively few describe infection with methicillin-susceptible S. aureus (MSSA). In the United States, colonization of healthy adults by ST398 has been reported in Iowa (5) and in New York, New York (2); MSSA infections have been anecdotally reported in St. Louis, Missouri (6), and The Bronx, New York (7). We describe 8 infections with MSSA ST398 in the New York City area during a 7-year period (2004–2010). Five infections with a related ST (ST291) from clonal complex (CC) 398 also were identified. These findings highlight the emergence of clinical infections with 2 distinct CC398 sequence types in the New York City area.
Retrospective typing of 4,167 clinical S. aureus isolates from various studies involving inpatients and outpatients in the New York City area identified 13 mecA-negative isolates with CC398-associated spa types (Table). Nine isolates were obtained from cultures of outpatients with skin and soft tissue infections; samples were submitted by physicians in the community. One isolate was associated with recurring skin and soft tissue infections in multiple body sites (BK21466); another was associated with genital infection (BK21732). Of the 4 ST398 isolates derived from bloodstream infections in hospitalized patients, 3 were recovered from intravenous drug users, 1 of whom died 1 day after admission for variceal bleeding (BK26722). Unlike the multidrug-resistant ST398 MSSA recently described in Colombia (4), most isolates in this study were susceptible to all antimicrobial drugs tested except penicillin, although several strains exhibited resistance to clindamycin and erythromycin. One isolate (BK23527) was submitted as oxacillin resistant (MIC ≥4 μg/mL) but lacked the mecA gene, which suggested that another mechanism was contributing to the resistance phenotype.
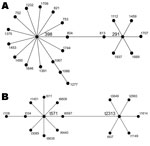
Multilocus sequence typing confirmed 8 isolates as ST398 (3–35–19–2–20–26–39); 5 isolates were assigned to ST291 (3–37–19–2–20–26–32), a double-locus variant of ST398 (Figure A1, panel A). Most of the ST398 strains were spa type 109 (t571), described in MSSA carriage isolates from New York City (2) and MSSA infections from China (1), France (8), Martinique (3), the Dominican Republic (2,3), and Colombia (4). BURP (based upon repeat pattern) analysis clustered all of the spa types into spa-CC t571 (Figure A1, panel B); ST398 isolates clustered with spa type 109 (t571), whereas ST291 isolates clustered with spa type 865 (t2313). Pulsed-field gel electrophoresis was also performed on the 11 available isolates. Although the ST291 isolates were sensitive to digestion with SmaI, pulsed-field gel electrophoresis was performed with Cfr9I to compare all isolates simultaneously. As expected, the ST398 and ST291 isolates clustered separately (data not shown); 4 distinct patterns were observed within each cluster (Table). Only the ST398 isolates were positive for a CC398 lineage-specific PCR that targets the unique restriction-modification system sau1-hsdS1 (9), further highlighting the differences between ST291 and ST398. None of the isolates harbored the genes coding for Panton-Valentine leukocidin.
Because of the retrospective nature of the findings, epidemiologic information for each isolate was limited. One patient (BK19382) reported travel to the Dominican Republic; Caribbean nationality was reported for BK27037 (Puerto Rico) and BK31274 (Trinidad). The cases described here occurred in urban and suburban settings, reflecting the likelihood that exposure to livestock was relatively low; however, travel history was unknown for most of the patients. Previous reports have linked ST398 transmission to other reservoirs, including companion animals, live animal food markets, and commercial meat products (1,2). However, data from a recent genome sequencing study suggest that MSSA ST398 is human in origin (10); other evidence suggests that certain lineages, particularly spa type 109 (t571), might circulate at low levels in humans in the absence of livestock exposure (8).
Our findings seem to support the hypothesis of low-level ST398 MSSA prevalence, and further surveillance might uncover additional cases of colonization or infection with ST398- and ST291-related strains in the New York City area. For example, active surveillance cultures performed at one of the 3 hospitals during January–March 2009 detected 7 additional ST398 and 3 additional ST291 isolates among 260 MSSA carriage strains (data not shown). In addition to the intrinsic virulence exhibited by ST398 MSSA in previous studies, the potential to acquire resistance to multiple classes of antimicrobial drugs (1,4,10), as well as virulence factors such as Panton-Valentine leukocidin (8), warrants continued surveillance in light of recent ST398 methicillin-resistant S. aureus outbreaks in health care settings (1).
Acknowledgments
We thank Bruce F. Farber for administrative support and clinical guidance and John Gattringer, Nancy Musa, and Michael Henry for gathering data related to this project.
This work was partially funded by the New York State Department of Health Hospital-Acquired Infection Reporting Program (D.A., M.E.S.) and Infection Prevention Project Program (K.S., B.K.) and by a grant from the New York Community Trust (B.N.K.).
References
- Smith TC, Pearson N. The emergence of Staphylococcus aureus ST398. Vector Borne Zoonotic Dis. 2011;11:327–39. DOIPubMedGoogle Scholar
- Bhat M, Dumortier C, Taylor BS, Miller M, Vasquez G, Yunen J, Staphylococcus aureus ST398, New York City and Dominican Republic. Emerg Infect Dis. 2009;15:285–7. DOIPubMedGoogle Scholar
- Uhlemann AC, Dumortier C, Hafer C, Taylor BS, Sanchez EJ, Rodriguez-Taveras C, Molecular characterization of Staphylococcus aureus from outpatients in the Caribbean reveals the presence of pandemic clones. Eur J Clin Microbiol Infect Dis. 2011; Epub ahead of print.DOIPubMedGoogle Scholar
- Jiménez JN, Vélez LA, Mediavilla JR, Ocampo AM, Vanegas JM, Rodríguez EA, Livestock-associated methicillin-susceptible Staphylococcus aureus ST398 in woman, Colombia. Emerg Infect Dis. 2011;17:1970–1.PubMedGoogle Scholar
- Smith TC, Male MJ, Harper AL, Kroeger JS, Tinkler GP, Moritz ED, Methicillin-resistant Staphylococcus aureus (MRSA) strain ST398 is present in midwestern U.S. swine and swine workers. PLoS ONE. 2009;4:e4258. DOIPubMedGoogle Scholar
- Orscheln RC, Hunstad DA, Fritz SA, Loughman JA, Mitchell K, Storch EK, Contribution of genetically restricted, methicillin-susceptible strains to the ongoing epidemic of community-acquired Staphylococcus aureus infections. Clin Infect Dis. 2009;49:536–42. DOIPubMedGoogle Scholar
- Varshney AK, Mediavilla JR, Robiou N, Guh A, Wang X, Gialanella P, Diverse enterotoxin gene profiles among clonal complexes of Staphylococcus aureus isolates from the Bronx, New York. Appl Environ Microbiol. 2009;75:6839–49. DOIPubMedGoogle Scholar
- Davies PR, Wagstrom EA, Bender JB. Lethal necrotizing pneumonia caused by an ST398 Staphylococcus aureus strain. Emerg Infect Dis. 2011;17:1152–3. DOIPubMedGoogle Scholar
- Stegger M, Lindsay JA, Moodley A, Skov R, Broens EM, Guardabassi L. Rapid PCR detection of Staphylococcus aureus clonal complex 398 by targeting the restriction-modification system carrying sau1-hsdS1. J Clin Microbiol. 2011;49:732–4. DOIPubMedGoogle Scholar
- Price LB, Stegger M, Hasman H, Aziz M, Larsen J, Andersen PS, Staphylococcus aureus CC398: host adaptation and emergence of methicillin resistance in livestock. MBio. 2012;3:pii:e00305-11. DOIGoogle Scholar
Figure
Table
Cite This ArticleRelated Links
Table of Contents – Volume 18, Number 4—April 2012
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Barry N. Kreiswirth, Public Health Research Institute Tuberculosis Center, University of Medicine and Dentistry of New Jersey, 225 Warren St, ICPH W210M, Newark, NJ 07103, USA
Top