Volume 19, Number 11—November 2013
Letter
Seoul Virus in Rats (Rattus norvegicus), Hyesan, North Korea, 2009–2011
Figure
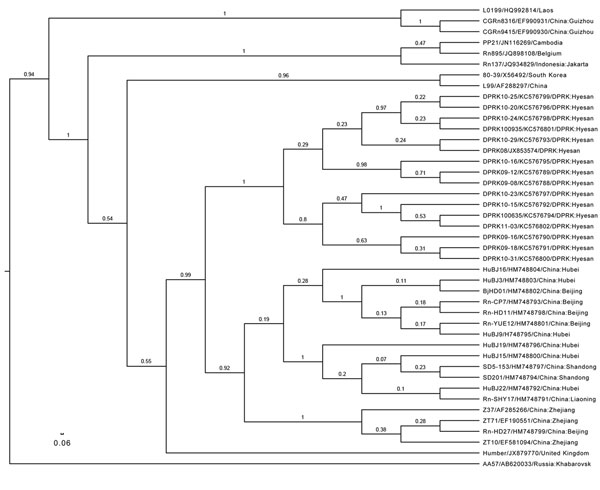
Figure. . . Phylogenetic tree, based on a 330-bp amplicon of the Seoul virus (SEOV) RNA-dependent RNA polymerase gene, depicted in FigTree1.4.0 (http://www.molecularevolution.org/software/phylogenetics/figtree). The tree was generated by using the uncorrelated lognormal distribution relaxed molecular clock model and SRD06 substitution model in BEAST1.74 (7). SEOV strain name/GenBank accession no/country: The location is shown in taxa. The posterior number is shown for each branch. Clades A and D were established as described (2) Scale bar represents number of nucleotide changes per site.
References
- Kariwa H, Yoshimatsu K, Arikawa J. Hantavirus infection in East Asia. Comp Immunol Microbiol Infect Dis. 2007;30:341–56. DOIPubMedGoogle Scholar
- Lin XD, Guo WP, Wang W, Zou Y, Hao ZY, Zhou DJ, Migration of Norway rats resulted in the worldwide distribution of Seoul hantavirus today. J Virol. 2012;86:972–81. DOIPubMedGoogle Scholar
- Kim HC, Klein TA, Chong ST, Collier BW, Usa M, Yi SC, Seroepidemiological survey of rodents collected at a U.S. military installation, Yongsan Garrison, Seoul, Republic of Korea. Mil Med. 2007;172:759–64 .PubMedGoogle Scholar
- Yao LS, Qin CF, Pu Y, Zhang XL, Liu YX, Liu Y, Complete genome sequence of Seoul virus isolated from Rattus norvegicus in the Democratic People's Republic of Korea. J Virol. 2012;86:13853. DOIPubMedGoogle Scholar
- Wang H, Yoshimatsu K, Ebihara H, Ogino M, Araki K, Kariwa H, Genetic diversity of hantaviruses isolated in China and characterization of novel hantaviruses isolated from Niviventer confucianus and Rattus rattus. Virology. 2000;278:332–45. DOIPubMedGoogle Scholar
- Klempa B, Fichet-Calvet E, Lecompte E, Auste B, Aniskin V, Meisel H, Hantavirus in African wood mouse, Guinea. Emerg Infect Dis. 2006;12:838–40. DOIPubMedGoogle Scholar
- Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29:1969–73. DOIPubMedGoogle Scholar
- Shapiro B, Rambaut A, Drummond AJ. Choosing appropriate substitution models for the phylogenetic analysis of protein-coding sequences. Mol Biol Evol. 2006;23:7–9. DOIPubMedGoogle Scholar
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28:2731–9. DOIPubMedGoogle Scholar
- Yan Y, Yao L, Hu G, Du Z, Li M. Zhang. Y. Genetic subtypes and distribution of Seoul virus in Jilin. Chinese Journal of Vector Biology and Control. 2006;17:324–6.
1These authors contributed equally to this article.
Page created: October 23, 2013
Page updated: October 23, 2013
Page reviewed: October 23, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.