Volume 19, Number 11—November 2013
Research
Migration and Persistence of Human Influenza A Viruses, Vietnam, 2001–2008
Figure 5
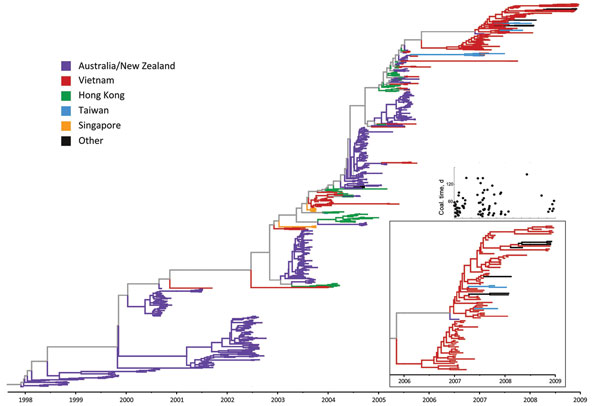
Figure 5. . Maximum clade credibility tree for regional influenza (H3N2) hemagglutinin data, generated by BEAST version 1.6.2 (26) under a constant population model; these are the same sequences as shown on Figure 1, except 2 sequences from Vietnam were removed because of missing sampling dates (n = 785 sequences). Inset on the bottom right shows a magnification of the tree for the 2007–2008 Vietnam sequences, to highlight persistence during this time. The smaller inset above shows coalescent (coal.) times for the Vietnam sequences in the larger inset. The 2 x-axes on the insets coincide, and each black circle showing a coalescence time corresponds to the tip of a branch of Vietnam virus in the magnified-tree inset.
References
- Rambaut A, Pybus OG, Nelson MI, Viboud C, Taubenberger JK, Holmes EC. The genomic and epidemiological dynamics of human influenza A virus. Nature. 2008;453:615–9. DOIPubMedGoogle Scholar
- Russell CA, Jones TC, Barr IG, Cox NJ, Garten RJ, Gregory V, The global circulation of seasonal influenza A (H3N2) viruses. Science. 2008;320:340–6. DOIPubMedGoogle Scholar
- Bedford T, Cobey S, Beerli P, Pascual M. Global migration dynamics underlie evolution and persistence of human influenza A (H3N2). PLoS Pathog. 2010;6:e1000918. DOIPubMedGoogle Scholar
- Bahl J, Nelson MI, Chan KH, Chen R, Vijaykrishna D, Halpin RA, Temporally structured metapopulation dynamics and persistence of influenza A H3N2 virus in humans. Proc Natl Acad Sci U S A. 2011;108:19359–64 and. DOIPubMedGoogle Scholar
- Cheng X, Tan Y, He M, Lam TT-Y, Lu X, Viboud C, Epidemiological dynamics and phylogeography of influenza virus in southern China. J Infect Dis. 2013;207:106–14. DOIPubMedGoogle Scholar
- Nguyen HT, Dharan NJ, Le MTQ, Nguyen NB, Nguyen CT, Hoang DV, National influenza surveillance in Vietnam 2006–2007. Vaccine. 2009;28:398–402. DOIPubMedGoogle Scholar
- Oxford University Clinical Research Unit. Influenza-like illness (ILI) in Ho Chi Minh City. 2013 [cited 2013 Mar 4]. www.ili.vn.
- Horby P, Mai Le Q, Fox A, Thai PQ, Thi Thu Yen N. Thanh le T, et al. The epidemiology of interpandemic and pandemic influenza in Vietnam, 2007–2010: The Ha Nam Household Cohort Study I. Am J Epidemiol. 2012;175:1062–74.
- Boni MF, Manh BH, Thai PQ, Farrar J, Hien TT, Hien NT, Modelling the progression of pandemic influenza A(H1N1) and the opportunities for reassortment with other influenza viruses. BMC Med. 2009;7:43. DOIPubMedGoogle Scholar
- Horby P, Thai PQ, Hens N, Yen NTT, Mai LQ, Thoang DD, Social contact patterns in Vietnam and implications for the control of infectious diseases. PLoS ONE. 2011;6:e16965. DOIPubMedGoogle Scholar
- Nelson MI, Simonsen L, Viboud C, Miller MA, Holmes EC. Phylogenetic analysis reveals the global migration of seasonal influenza A viruses. PLoS Pathog. 2007;3:1220–8. DOIPubMedGoogle Scholar
- Nelson MI, Simonsen L, Viboud C, Miller MA, Taylor J, St. George K, Stochastic processes are key determinants of short-term evolution in influenza A virus. PLoS Pathog. 2006;2:e125. DOIPubMedGoogle Scholar
- Tang JW, Ngai KLK, Lam WY, Chan PKS. Seasonality of influenza A (H3N2) virus: a Hong Kong perspective (1997–2006). PLoS ONE. 2008;3:e2768. DOIPubMedGoogle Scholar
- Doraisingham S, Goh KT, Ling AE, Yu M. Influenza surveillance in Singapore: 1972–86. Bull World Health Organ. 1988;66:57–63 .PubMedGoogle Scholar
- Simmerman JM, Uyeki TM. The burden of influenza in East and South-East Asia: a review of the English language literature. Influenza Other Respi Viruses. 2008;2:81–92. DOIPubMedGoogle Scholar
- Wong CM, Yang L, Chan KP, Leung GM, Chan KH, Guan Y, Influenza-associated hospitalization in a subtropical city. PLoS Med. 2006;3:e121. DOIPubMedGoogle Scholar
- Yap FHY, Ho P-L, Lam K-F, Chan PKS, Cheng Y-H, Peiris JSM. Excess hospital admissions for pneumonia, chronic obstructive pulmonary disease, and heart failure during influenza seasons in Hong Kong. J Med Virol. 2004;73:617–23. DOIPubMedGoogle Scholar
- Shek LPC, Lee BW. Epidemiology and seasonality of respiratory tract virus infections in the tropics. Paediatr Respir Rev. 2003;4:105–11. DOIPubMedGoogle Scholar
- Finkelman BS, Viboud C, Koelle K, Ferrari MJ, Bharti N, Grenfell BT. Global patterns in seasonal activity of influenza A/H3N2, A/H1N1, and B from 1997 to 2005: viral coexistence and latitudinal gradients. PLoS ONE. 2007;2:e1296. DOIPubMedGoogle Scholar
- Viboud C, Alonso WJ, Simonsen L. Influenza in tropical regions. PLoS Med. 2006;3:e89. DOIPubMedGoogle Scholar
- Alonso WJ, Viboud C, Simonsen L, Hirano EW, Daufenbach LZ, Miller MA. Seasonality of influenza in Brazil: a traveling wave from the Amazon to the subtropics. Am J Epidemiol. 2007;165:1434–42. DOIPubMedGoogle Scholar
- Ghedin E, Sengamalay NA, Shumway M, Zaborsky J, Feldblyum T, Subbu V, Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution. Nature. 2005;437:1162–6. DOIPubMedGoogle Scholar
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7 . DOIPubMedGoogle Scholar
- Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol. 2008;57:758–71. DOIPubMedGoogle Scholar
- Stamatakis A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 2006;22:2688–90. DOIPubMedGoogle Scholar
- Drummond AJ, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007;7:214. DOIPubMedGoogle Scholar
- Swofford DL, Maddison WP. Reconstructing ancestral character states under Wagner parsimony. Math Biosci. 1987;87:199–229. DOIGoogle Scholar
- Swofford DL. PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4. Sunderland (MA): Sinauer Associates; 2002.
- Boni MF, de Jong MD, van Doorn HR, Holmes EC. Guidelines for identifying homologous recombination events in influenza A virus. PLoS ONE. 2010;5:e10434. DOIPubMedGoogle Scholar
- Bastian M, Heymann S, Jacomy M. Gephi: an open source software for exploring and manipulating networks. Third International AAAI Conference on Weblogs and Social Media; 2009 May 17– 20; San Jose, California [cited 2013 Mar 4]. http://www.aaai.org/Library/ICWSM/icwsm09contents.php
- Boni MF, Gog JR, Andreasen V, Feldman MW. Epidemic dynamics and antigenic evolution in a single season of influenza A. Proc Biol Sci. 2006;273:1307–16. DOIPubMedGoogle Scholar
- Adams B, McHardy AC. The impact of seasonal and year-round transmission regimes on the evolution of influenza A virus. Proc Biol Sci. 2011;278:2249–56. DOIPubMedGoogle Scholar
- Harvey PH, Pagel MD. The comparative method in evolutionary biology. Oxford (UK): Oxford University Press; 1991.
- Pybus OG, Suchard MA, Lemey P, Bernardin FJ, Rambaut A, Crawford FW, Unifying the spatial epidemiology and molecular evolution of emerging epidemics. Proc Natl Acad Sci U S A. 2012;109:15066–71. DOIPubMedGoogle Scholar
- Lemey P, Rambaut A, Drummond AJ, Suchard MA. Bayesian phylogeography finds its roots. PLOS Comput Biol. 2009;5:e1000520 . DOIPubMedGoogle Scholar
- Rabaa MA, Hang VTT, Wills B, Farrar J, Simmons CP, Holmes EC. Phylogeography of recently emerged DENV-2 in southern Viet Nam. PLoS Negl Trop Dis. 2010;4:e766 . DOIPubMedGoogle Scholar
- Raghwani J, Rambaut A, Holmes EC, Hang VT, Hien TT, Farrar J, Endemic dengue associated with the co-circulation of multiple viral lineages and localized density-dependent transmission. PLoS Pathog. 2011;7:e1002064 . DOIPubMedGoogle Scholar
Page created: October 31, 2013
Page updated: October 31, 2013
Page reviewed: October 31, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.