Volume 19, Number 3—March 2013
Letter
Cryptococcus gattii, Florida, USA, 2011
Figure
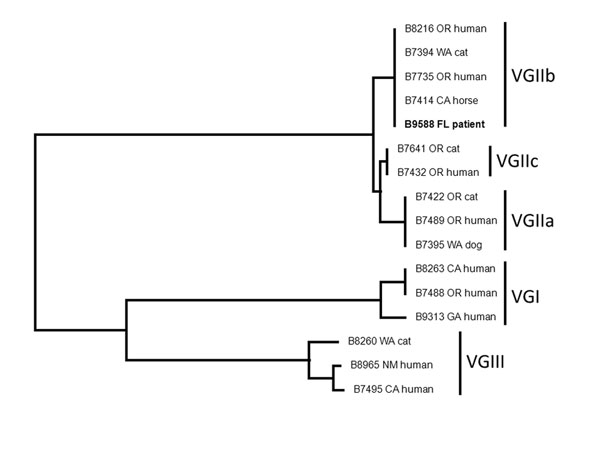
Figure. . . Neighbor-joining dendrogram of FL isolate (B9588, in boldface) with other US isolates showing that the FL isolate is identical to the VGIIb isolates from the US Pacific Northwest. The dendrogram was constructed by using multilocus sequence typing (3). FL, Florida; OR, Oregon; WA, Washington; CA, California; GA, Georgia; NM, New Mexico.
References
- Datta K, Bartlett KH, Baer R, Byrnes E, Galanis E, Heitman J, Spread of Cryptococcus gattii into Pacific Northwest region of the United States. Emerg Infect Dis. 2009;15:1185–91. DOIPubMedGoogle Scholar
- Brandt ME, Hutwagner LC, Klug LA, Baughman WS, Rimland D, Graviss EA, Molecular subtype distribution of Cryptococcus neoformans in four areas of the United States. Cryptococcal Disease Active Surveillance Group. J Clin Microbiol. 1996;34:912–7 .PubMedGoogle Scholar
- Meyer W, Aanensen DM, Boekhout T, Cogliati M, Diaz MR, Esposto MC, Consensus multi-locus sequence typing scheme for Cryptococcus neoformans and Cryptococcus gattii. Med Mycol. 2009;47:561–70. DOIPubMedGoogle Scholar
- Harris J, Lockhart S, Chiller T. Cryptococcus gattii: where do we go from here? Med Mycol. 2012;50:113–29. DOIPubMedGoogle Scholar
- Bovers M, Hagen F, Kuramae EE, Boekhout T. Six monophyletic lineages identified within Cryptococcus neoformans and Cryptococcus gattii by multi-locus sequence typing. Fungal Genet Biol. 2008;45:400–21. DOIPubMedGoogle Scholar
- Kidd SE, Hagen F, Tscharke RL, Huynh M, Bartlett KH, Fyfe M, A rare genotype of Cryptococcus gattii caused the cryptococcosis outbreak on Vancouver Island (British Columbia, Canada). Proc Natl Acad Sci U S A. 2004;101:17258–63. DOIPubMedGoogle Scholar
- Byrnes EJ III, Bildfell R, Frank SA, Mitchell TG, Marr KA, Heitman J. Molecular evidence that the range of the Vancouver Island outbreak of Cryptococcus gattii infection expanded into the Pacific Northwest in the United States. J Infect Dis. 2009;199:1081–6. DOIPubMedGoogle Scholar
- Duncan C, Stephen C, Campbell J. Clinical characteristics and predictors of mortality for Cryptococcus gattii infection in dogs and cats of southwestern British Columbia. Can Vet J. 2006;47:993–8.PubMedGoogle Scholar
- Chan ED, Kong PM, Fennelly K, Dwyer AP, Iseman MD. Vertebral osteomyelitis due to infection with nontuberculous Mycobacterium species after blunt trauma to the back: 3 examples of the principle of locus minoris resistentiae. Clin Infect Dis. 2001;32:1506–10. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Emergence of Cryptococcus gattii—Pacific Northwest, 2004–2010. MMWR Morb Mortal Wkly Rep. 2010;59:865–8.PubMedGoogle Scholar
Page created: February 13, 2013
Page updated: February 13, 2013
Page reviewed: February 13, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.