Volume 19, Number 7—July 2013
Letter
Usutu Virus in Migratory Song Thrushes, Spain
Figure
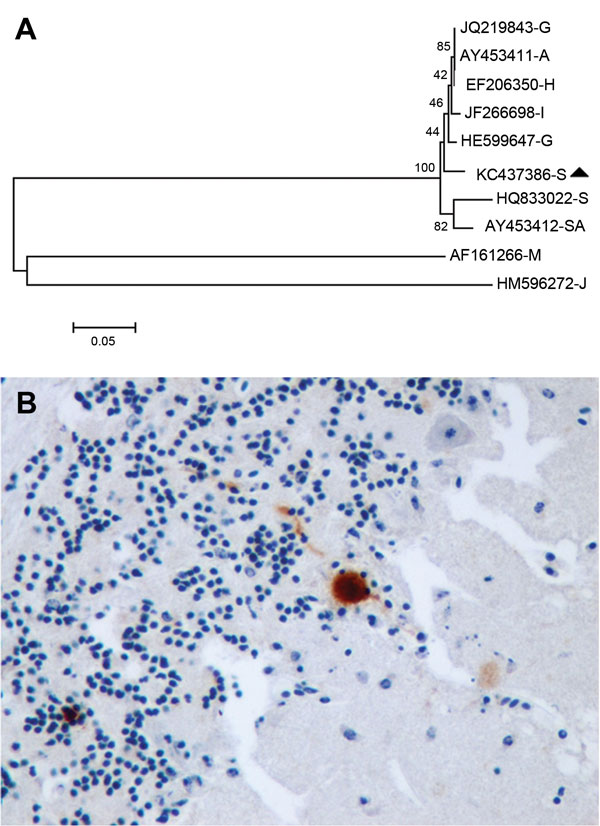
Figure. . . Usutu virus (USUV) in migratory song thrushes in Spain. A) Phylogenetic analysis of European and African USUV strains. The evolutionary history was inferred using the neighbor-Joining method. The optimal tree with the sum of branch length = 1.18014408 is shown. The percentages of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) are shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed by using the maximum composite likelihood method and are in the units of the number of base substitutions per site. The analysis involved 10-nt sequences. All ambiguous positions were removed for each sequence pair. There were a total of 929 positions in the final dataset. Evolutionary analyses were conducted in MEGA5 (www.megasoftware.net). USUV strains are identified by their GenBank accession numbers and a capital letter indicating the country of origin (A, Austria; G, Germany; I, Italy; H, Hungary; S, Spain; SA, South Africa); the sequences of the last 2 branches correspond to outgroup viruses used to root the tree (M, Murray Valley encephalitis virus [GenBank AF161266]; J, Japanese encephalitis virus [GenBank HM596272]). The song thrush strain from Spain 2012 is highlighted with a black triangle. Scale bar indicates nucleotide changes per site. B) Immunohistochemical staining with cross-reacting antibody showing USUV antigen labeling in a Purkinje cell of the cerebellum of a song thrush that died from encephalitis. Original magnification x400.