Volume 20, Number 12—December 2014
Letter
Peste des Petits Ruminants Virus, Tunisia, 2012–2013
Cite This Article
Citation for Media
To the Editor: Peste des petits ruminants (PPR) is a viral disease of sheep and goats caused by peste des petits ruminants virus (PPRV), a negative-sense, single-stranded RNA virus of the genus Morbillivirus. Illness and death can be high (>90%) when PPR occurs in populations of immunologically naive sheep and goats (1). Mortality rates are ≈10%–40% in disease-endemic areas (2). Because of its economic effects and ability to spread rapidly, PPR has been included among reportable diseases by the World Organization for Animal Health.
In the past 20 years, PPR has shown rapid spread throughout large areas of Africa and Asia (2). A unique serotype of PPRV circulates and is classified into 4 genetically distinct lineages (3). The geographic distribution of lineages I and II is restricted mainly to western and central Africa and that of lineage III to eastern Africa. Lineage IV is more widely distributed throughout eastern Africa (4,5), the Near and Middle East, and large areas of Asia (3).
In 2008, PPR occurred in Morocco, and 257 cases were reported in goats and sheep over a 6-month period (3). In 2011, PPR was officially reported in Algeria (3). Genetic analysis showed that PPRV strains isolated in Morocco and Algeria belonged to lineage IV (4–6). Although PPR has been reported in Tunisia since 2011 (7), no data are available on the molecular characterization of PPRV circulating in this country.
During September 2012–January 2013, clinical signs compatible with PPR in ovine and caprine flocks were reported to the Tunisian veterinary service. Ocular, nasal, oral, and rectal swab specimens were obtained from animals showing clinical signs of this disease. Swab specimens were sent to the Institute de la Recherche Vétérinaire de Tunisie in Tunis for laboratory confirmation. Total RNA from swab samples was extracted by using the NucleoSpin RNA Virus Kit (Macherey-Nagel, Düren, Germany) according to the manufacturer’s instructions. The presence of the PPR viral RNA was determined in samples by using a specific reverse transcription PCR reported by Polci et al. (8). Laboratory tests confirmed circulation of PPRV in farms near Kairouan and Sidi Bouzid.
Aliquots of RNA samples were shipped to the Istituto Zooprofilattico Sperimentale dell’Abruzzo e del Molise in Teramo, Italy, for genetic characterization. RNA was amplified by using the reverse transcription PCR reported by Couacy-Hymann et al. (9). Amplicons from virus- positive samples were purified by using the QIAquick PCR Purification Kit (QIAGEN, Valencia, CA, USA) and used for direct sequencing. Sequencing reactions were performed by using the Big Dye Terminator Kit (Applied Biosystems, Foster City, CA, USA), and nucleotide sequences were determined by using the ABI PRISM 3100 DNA sequencer (Applied Biosystems). Amplification and sequencing were repeated twice to avoid introduction of artificial substitutions. Raw sequence data were assembled by using Contig Express (Vector NTI suite 9.1; Invitrogen, Carlsbad, CA, USA), and a 351-nt fragment of the nucleoprotein coding sequence was obtained after deletion of primer sequences.
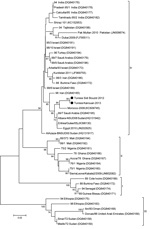
Two sequences were obtained, 1 from the Kairouan outbreak and 1 from the Sidi Bouzid outbreak. The 2 sequences generated in this study were submitted to GenBank (accession nos. KM068121 and KM068122). Sequences showed nearly complete identity at nucleotide and amino acid levels; there was 1 nucleotide substitution. The BLAST (http://www.ncbi.nlm.nih.gov) was used to detect homologous regions in sequence databases. Sequences were aligned by using ClustalW (http://www.genome.jp/tools/clustalw/) and MEGA version 6 (10). Phylogenetic analysis was performed with a 255-nt sequence of the PPRV nucleoprotein gene by using the neighbor-joining method with bootstrap support (1,000 replicates) in MEGA version 6 (10) and reference strains representing the 4 lineages of PPRV that have been isolated in different years or countries (Figure).
Our results indicate that lineage IV of PPRV is present in Tunisia. PPRV isolates from the outbreaks in Sidi Bouzid in 2012 and Kairouan in 2013 are closely related to viruses responsible for PPR outbreaks in Morocco in 2008 (4) and Algeria in 2010 (6). These isolates are closely related to strains from Saudi Arabia, which were detected in Eritrea in 2005 (5) and in Sudan in 2008 (4). These data suggest that a unique PPRV strain is circulating across this area of the Maghreb. PPRV circulation is maintained probably by the abundant trade in ruminants between Tunisia and neighboring countries. This information highlights the need for a regional approach to control PPR in northern Africa.
Acknowledgment
The Istituto Zooprofilattico Sperimentale dell’Abruzzo e del Molise G. Caporale is supported by the Italian Ministry of Health.
References
- Albina E, Kwiatek O, Minet C, Lancelot R, Servan de Almeida R, Libeau G. Peste des petits ruminants, the next eradicated animal disease? Vet Microbiol. 2013;165:38–44 . DOIPubMedGoogle Scholar
- Baron MD, Parida S, Oura CA. Peste des petits ruminants: a suitable candidate for eradication? Vet Rec. 2011;169:16–21. DOIPubMedGoogle Scholar
- Munir M, Zohari S, Berg M. Molecular biology and pathogenesis of peste des petits ruminants virus. New York: Springer; 2013.
- Kwiatek O, Ali YH, Saeed IK, Khalafalla AI, Mohamed OI, Obeida AA, Asian lineage of peste des petits ruminants virus, Africa. Emerg Infect Dis. 2011;17:1223–31. DOIPubMedGoogle Scholar
- Cosseddu GM, Pinoni C, Polci A, Sebhatu T, Lelli R, Monaco F. Characterization of peste des petits ruminants virus, Eritrea, 2002–2011. Emerg Infect Dis. 2013;19:160–1. DOIPubMedGoogle Scholar
- De Nardi M, Lamin Saleh SM, Batten C, Oura C, Di Nardo A, Rossi D. First evidence of peste des petits ruminants (PPR) virus circulation in Algeria (Sahrawi territories): outbreak investigation and virus lineage identification. Transbound Emerg Dis. 2012;59:214–22. DOIPubMedGoogle Scholar
- Office International des Epizooties (OIE). WAHID Interface, 2011. Event summary: peste des petits ruminants, Tunisia [cited 2014 Jun 21]. http://www.oie.int/wahis_2/public/wahid.php/Reviewreport/Review/viewsummary?fupser=&dothis=&reportid=10579.
- Polci A, Cosseddu GM, Ancora M, Pinoni C, El Harrak M, Sebhatu TT, Development and preliminary evaluation of a new real-time RT-PCR assay for detection of peste des petits ruminants virus genome. Transbound Emerg Dis. 2013;19:•••; Epub ahead of print and.PubMedGoogle Scholar
- Couacy-Hymann E, Roger F, Hurard C, Guillou JP, Libeau G, Diallo A. Rapid and sensitive detection of peste des petits ruminants virus by a polymerase chain reaction assay. J Virol Methods. 2002;100:17–25. DOIPubMedGoogle Scholar
- Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol. 2013;30:2725–9 . DOIPubMedGoogle Scholar
Figure
Cite This Article1These authors contributed equally to this article.
Related Links
Table of Contents – Volume 20, Number 12—December 2014
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Gian Mario Cosseddu, Istituto Zooprofilattico Sperimentale dell’Abruzzo e del Molise G. Caporale, Campo Boario, 64100 Teramo, Italy
Top