Volume 20, Number 5—May 2014
Dispatch
PCR for Detection of Oseltamivir Resistance Mutation in Influenza A(H7N9) Virus
Cite This Article
Citation for Media
Abstract
Sensitive molecular techniques are needed for rapid detection of the R292K oseltamivir-resistant mutant of influenza A(H7/N9) virus strain to monitor its transmission and guide antiviral treatment. We developed a real-time reverse transcription PCR and single nucleotide polymorphism probes to differentiate this mutant strain in mixed virus populations in human specimens.
An outbreak of human infections with a novel reassortant avian-origin influenza A(H7N9) virus occurred in several provinces of China during March 2013 (1), This outbreak caused 137 laboratory-confirmed cases and 45 deaths as of October 2013 (www.who.int/csr/don/2013_10_24a/en/index.html). An unusually high proportion of severe cases and a high case-fatality rate have been observed for patients infected with this virus (2).
We reported emergence of an influenza virus with a mutation in the neuraminidase (NA) gene (R292K) and its association with severe clinical outcome in infected persons (3). Studies have shown that the NA R292K mutation can cause a high level of resistance to oseltamivir in influenza A(H7N9) virus (4,5). Thus, sensitive molecular techniques are needed for rapid detection of influenza virus with this mutation to monitor its circulation and transmission and guide antiviral treatment. In this study, we developed a single-nucleotide polymorphism (SNP) real-time reverse transcription PCR (RT-PCR) to differentiate NA 292K mutant virus from R292 wild-type virus in clinical samples.
The NA R292K assay has 2 reactions with 1 pair of primers. One reaction contained a FAM-labeled SNP probe specific for the 292K mutant strain and a second reaction contained a VIC-labeled probe specific for the R292 wild-type strain (Technical Appendix).
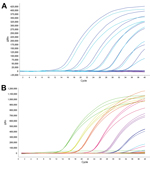
To assess the sensitivity of the assay, we constructed 2 plasmids that contained R292 wild-type virus or 292K mutant virus, respectively. Fragments of the NA gene inserted into the plasmids were amplified from nasopharyngeal swab specimens from 2 patients infected with influenza A(H7N9) virus and confirmed by using Sanger sequencing (Technical Appendix). The 2 plasmids were serially diluted 10-fold (101–1011 copies) in sterile water and used to test the assay. The linear range of sensitivity was 102–108 copies. The lower limit of detection was 100 copies/reaction (3/3 reactions detected) for wild-type and mutant virus (Figure). However, the sensitivity of the duplex reaction containing both probes was 100-fold lower than that of each separate reaction.
Of 35 respiratory samples tested, 6 were infected with influenza A(H3N2), 2 with influenza A(H1N1) virus, 6 with influenza A(H1N1)pdm09 virus, 4 with parainfluenza virus, 4 with human rhinovirus, 4 with human coronavirus, 5 with influenza B virus, and 4 with respiratory syncytial virus. In addition, 6 other respiratory samples were virus negative. Cross-amplification was not observed during sample testing. Thus, the assay is highly specific for detecting the mutant NA gene of influenza A(H7N9) virus.
To test the performance of the assay when 292K mutant and R292 wild-type viruses were present in 1 sample, a series of mixtures containing the 292K plasmid and the R292 plasmid at copy numbers of 104 copies/reaction were prepared at the following ratios of mutant virus to wild-type virus: 2:98, 5:95, 10:90, 20:80, 30:70, 40:60, 50:50, 60:40, 70:30, 80:20, 90:10, 95:5, and 98:2. The ΔCtK – R of the mixture a ratio of 50:50 was used as the assay-specific normalization value in determination of the percentage of 292K mutant in mixed population as described by Liu et al. (6). The assay detected the 292K mutant in the mixture at a proportion of 2% of the 104 copies/reaction, and correct estimation of its proportion ranged from 10% to 98% (Technical Appendix).
To validate the assay with clinical samples, we tested 11 paired nasopharyngeal swab specimens and sputum specimens obtained from 9 patients infected with influenza A(H7N9) virus who had various disease outcomes (Table). The time of sampling (mean 12.6 days, range 7–20 days) was at the end of treatment with an NA inhibitor (oseltamivir or peramivir) or afterwards. Eleven of 22 samples were positive for influenza A(H7N9) virus by a quantitative real-time RT-PCR described in a previous study (3). Seven of 11 samples had positive results in the R292K assay: 5 samples positive in the 292K assay and 2 samples positive in both assays. Four of 11 samples were negative in both assays. All 292K-positive samples were further confirmed as positive by Sanger sequencing of NA genes.
The 7 samples that contained the 292K mutant were obtained from 4 patients: 2 patients who died (patients 2 and 3) and 2 patients who recovered (patients 8 and 10). In our previous study, sequencing of the NA gene was not successful for the first throat swab specimens from patients 8 and 10 because of low viral load (3). However, in this study, the 292K mutant was found in sputum specimens from these 2 patients on days 7 or 15, respectively, after initiation of antiviral treatment. This finding suggested that influenza A(H7N9) virus mutated under the pressure of antiviral treatment, which led to failure of the virus to clear the lower respiratory tracts. These 2 patients, who were infected with the drug-resistant mutant virus, recovered from their diseases, which suggested that host immune response might play a major role in controlling the mutant virus.
Higher viral load in sputum samples indicated that there might be factors, including hemagglutinin (HA) binding preference, which favor greater replication in the lower respiratory tract. A similar phenomenon was observed in patients infected with the HA D222G mutant of influenza A(H1N1)pdm09 virus; this virus showed preferential replication in the lower respiratory tract and this infection was correlated with severe outcomes or deaths (7).
We have developed an SNP real-time RT-PCR for detection of a drug-resistant NA R292K mutant of influenza A(H7N9) virus. The sensitivity of the assay is lower than that of an HA7-specific real-time RT-PCR (i.e., 4 samples positive for influenza A(H7N9) virus were not detected by this RT-PCR). However, as a screening tool, this assay is sensitive, specific, fast, and inexpensive.
This assay had a detection threshold of 10% for a mutant strain in a mixed viral population, which is more sensitive than Sanger sequencing (detection threshold of 25% for a minor component in a mixed viral population) (6). This assay will help clinicians monitor emergence of drug-resistant virus strains during treatment of patients with NA inhibitors to prevent persistent viral replication and severe inflammatory reactions.
Dr Wang is an associate professor in the Pathogen Diagnosis and Biosafety Department, Shanghai Public Health Clinical Center affiliated with Fudan University, Shanghai, China. Her primary research interest is genomic features of respiratory pathogens.
Acknowledgment
This study was supported by the National Megaprojects of China for Infectious Diseases (2012ZX10004211 and 2014ZX10004002-005), the Ministry of Science and Technology (KJYJ-2013-01-01), and the Shanghai Municipal Health and Family Planning Commission (2013QLG002).
References
- Gao R, Cao B, Hu Y, Feng Z, Wang D, Hu W, Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med. 2013;368:1888–97. DOIPubMedGoogle Scholar
- Gao HN, Lu HZ, Cao B, Du B, Shang H, Gan JH, Clinical findings in 111 cases of influenza A (H7N9) virus infection. N Engl J Med. 2013;368:2277–85. DOIPubMedGoogle Scholar
- Hu Y, Lu S, Song Z, Wang W, Hao P, Li J, Association between adverse clinical outcome in human disease caused by novel influenza A H7N9 virus and sustained viral shedding and emergence of antiviral resistance. Lancet. 2013;381:2273–9. DOIPubMedGoogle Scholar
- Sleeman K, Guo Z, Barnes J, Shaw M, Stevens J, Gubareva LV. R292K substitution and drug susceptibility of influenza A(H7N9) viruses. Emerg Infect Dis. 2013;19:1521–4. DOIPubMedGoogle Scholar
- Watanabe T, Kiso M, Fukuyama S, Nakajima N, Imai M, Yamada S, Characterization of H7N9 influenza A viruses isolated from humans. Nature. 2013;501:551–5. DOIPubMedGoogle Scholar
- Liu CM, Driebe EM, Schupp J, Kelley E, Nguyen JT, McSharry JJ, Rapid quantification of single-nucleotide mutations in mixed influenza A viral populations using allele-specific mixture analysis. J Virol Methods. 2010;163:109–15. DOIPubMedGoogle Scholar
- Liu Y, Childs RA, Matrosovich T, Wharton S, Palma AS, Chai W, Altered receptor specificity and cell tropism of D222G hemagglutinin mutants isolated from fatal cases of pandemic A(H1N1) 2009 influenza virus. J Virol. 2010;84:12069–74. DOIPubMedGoogle Scholar
Figure
Table
Cite This Article1These authors contributed equally to this article.
Table of Contents – Volume 20, Number 5—May 2014
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Yunwen Hu, Pathogen Diagnosis and Biosafety Department, Shanghai Public Health Clinical Center, Fudan University, No. 2901 Caolang Rd, Jinshan District, Shanghai, ChinaYunwen Hu, Pathogen Diagnosis and Biosafety Department, Shanghai Public Health Clinical Center, Fudan University, No. 2901 Caolang Rd, Jinshan District, Shanghai, China
Top