Volume 20, Number 5—May 2014
CME ACTIVITY - Synopsis
Outbreaks of Kingella kingae Infections in Daycare Facilities
Cite This Article
Citation for Media
Introduction

Medscape, LLC is pleased to provide online continuing medical education (CME) for this journal article, allowing clinicians the opportunity to earn CME credit.
This activity has been planned and implemented in accordance with the Essential Areas and policies of the Accreditation Council for Continuing Medical Education through the joint sponsorship of Medscape, LLC and Emerging Infectious Diseases. Medscape, LLC is accredited by the ACCME to provide continuing medical education for physicians.
Medscape, LLC designates this Journal-based CME activity for a maximum of 1 AMA PRA Category 1 Credit(s)TM. Physicians should claim only the credit commensurate with the extent of their participation in the activity.
All other clinicians completing this activity will be issued a certificate of participation. To participate in this journal CME activity: (1) review the learning objectives and author disclosures; (2) study the education content; (3) take the post-test with a 70% minimum passing score and complete the evaluation at www.medscape.org/journal/eid; (4) view/print certificate.
Release date: April 9, 2014; Expiration date: April 9, 2015
Learning Objectives
Upon completion of this activity, participants will be able to:
• Distinguish anatomic sites of invasive Kingella kingae infections
• Analyze the clinical presentation of invasive K. kingae infections
• Evaluate the diagnostic process for invasive K. kingae infections
• Assess the treatment of contacts of children with invasive K. kingae infections.
CME Editor
Jean Michaels Jones, Technical Writer/Editor, Emerging Infectious Diseases. Disclosure: Jean Michaels Jones has disclosed no relevant financial relationships.
CME Author
Charles P. Vega, MD, Associate Professor and Residency Director, Department of Family Medicine, University of California, Irvine. Disclosure: Charles P. Vega, MD, has disclosed no relevant financial relationships.
Author
Disclosure: Pablo Yagupsky, MD, has disclosed no relevant financial relationships.
Abstract
During the past decade, transmission of the bacterium Kingella kingae has caused clusters of serious infections, including osteomyelitis, septic arthritis, bacteremia, endocarditis, and meningitis, among children in daycare centers in the United States, France, and Israel. These events have been characterized by high attack rates of disease and prevalence of the invasive strain among asymptomatic classmates of the respective index patients, suggesting that the causative organisms benefitted from enhanced colonization fitness, high transmissibility, and high virulence. After prophylactic antibacterial drugs were administered to close contacts of infected children, no further cases of disease were detected in the facilities, although test results showed that some children still carried the bacterium. Increased awareness of this public health problem and use of improved culture methods and sensitive nucleic acid amplification assays for detecting infected children and respiratory carriers are needed to identify and adequately investigate outbreaks of K. kingae disease.
During the past 3 decades, Western countries have reported a rising number of mothers entering the workforce and, consequently, a growing number of children receiving care outside the home (1). This trend has substantial public health consequences because the incidence of infectious diseases in general, and of those caused by respiratory pathogens in particular, has substantially increased among daycare center attendees (1,2). These organisms are usually spread within daycare centers by child-to-child transmission; they colonize the upper respiratory tract surfaces, from which they can disseminate to other attendees. From the upper airways, pathogens may invade adjacent structures such as the lungs, middle ear, or nasal sinuses, and may penetrate into the bloodstream, causing invasive diseases (1). Most bacterial pathogens responsible for such infections are enclosed by polysaccharide capsules that protect them from phagocytosis and complement-mediated killing, ensuring their persistence on the respiratory mucosae and survival in the bloodstream and deep body tissues (3). Maturation of the T-cell independent arm of the immune system in humans is delayed until the age of 2–4 years; thus, young children are prone to colonization and infection by encapsulated bacteria (3).
Besides the microorganisms’ virulence and the hosts’ age-related immunologic immaturity, many other factors contribute to the enhanced colonization, transmission, and illness rates observed among children in daycare centers, including the number of children present, the degree of crowding, efficacy of ventilation, time spent in day care, length of time from enrollment, frequency of enrollment, age group mixing, and occurrence of seasonal viral infections (1,4–6). Because of age stratification, child-care groups comprise attendees of approximately the same age who have similar degrees of immunologic immaturity and susceptibility to infectious agents. This epidemiologic setting substantially differs from that of large families in that the latter include children of different ages and therefore, at any given time, only a fraction of siblings belong to the age group at enhanced risk for bacterial colonization and invasion, which limits the chances to acquire and transmit the organism (7). In daycare centers, respiratory organisms spread easily through large droplet transmission among young children with poor hygienic habits who share toys contaminated with respiratory secretions or saliva. Under these circumstances, introduction of a virulent bacterium in a crowded daycare facility attended by immunologically naïve children may result in prompt dissemination of the organism and initiate outbreaks of disease such as those caused by pneumococci, Haemophilus influenzae type b, or Neisseria meningitidis (1,2,6).
Because of the improved culture methods (8,9) and sensitive nucleic acid amplification assays (NAAAs) (10–14) developed in recent years, Kingella kingae, a gram-negative coccobacillus of the Neisseriaceae family, is increasingly recognized as an invasive pathogen of early childhood. The organism is a frequent source of childhood bacteremia and the most common agent of skeletal system infections in children 6 months–3 years of age; it is also a cause of bacterial endocarditis in children and adults (8–16). Because of the fastidious nature of K. kingae, many illnesses caused by this bacterium are, probably, overlooked. Although most cases of invasive K. kingae infections are sporadic, clusters of invasive disease have been detected among attendees of daycare centers in Israel, Europe, and the United States.
Misdiagnosis
The bacteriologic identification of K. kingae relies on the following: typical Gram stain results, showing pairs or short chains of plump, gram-negative bacilli with tapered ends; β-hemolysis; pitting of the agar surface; failure to grow on MacConkey medium; weak oxidase activity and a negative catalase reaction; production of acid from glucose; and, with rare exceptions, production of acid maltose (8,9). However, K. kingae tends to retain crystal violet dye and, therefore, it may appear to be gram-positive, and laboratories unfamiliar with its cultural and staining features may misidentify the bacterium altogether or dismiss invasive isolates as culture contaminants. Identification of K. kingae, however, is not difficult, and many commercial instruments and technologies such as VITEK 2 (bioMérieux, Marcy-l’Etoile, France), matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF), or 16S rDNA gene sequencing, correctly identify the organism.
Characteristics and Manifestations
Recent studies have demonstrated the presence of a polysaccharide capsule on the surface of K. kingae that exhibits between-strains chemical variation and, most probably, antigenic variation (17,18). These characteristics may assist K. kingae in evading the immune response by facilitating the successive colonization of the host by strains harboring different capsular types. The polysaccharide capsule probably enables mucosal colonization, the survival of the organism in the bloodstream, and invasion of deep body sites (17,18). In addition, all K. kingae strains produce and secrete a potent repeats-in-toxin (RTX) that exhibits a wide range of cytotoxic activity and is particularly deleterious to macrophage-like cells, leukocytes, and synovial cells and, to a lesser degree, to respiratory epithelial cells that improve the organism’s chances of surviving in the host and of invading skeletal tissues (12,19).
Similar to meningococci, K. kingae is carried on the oropharyngeal epithelium (20,21), and the colonized mucosa is the portal of entry of the organism to the bloodstream from which it may disseminate to 3 areas for which the bacterium shows particular tropism: joints, bones, or the endocardium (22,23). Damage to the upper respiratory surfaces by previous or concurrent viral infections or stomatitis appears to facilitate bloodstream invasion by K. kingae (16).
K. kingae isolates show remarkable genomic diversity and, to date, 37 multilocus sequence typing (MLST) and 74 pulsed field gel electrophoresis (PFGE) clones have been identified (24–26). Carried K. kingae organisms differ in their invasive capabilities (27), and simultaneous carriage of ≥1 genotype is unusual (28). Whereas some strains, which are frequently isolated from healthy carriers, are seldom if ever detected in patients with clinical disease, others are rarely carried asymptomatically, but are responsible for a high proportion of invasive disease (24,27,28). However, a few strains appear to possess an optimal balance between transmissibility and invasiveness and are common among healthy carriers and among infected patients (24,27,28). A recent study has found that certain virulent K. kingae clones, characterized by a distinct combination of PFGE and MLST profiles, are substantially associated with bacteremia with no focal infection, skeletal system invasion, or endocarditis, which suggests biological specialization for invading specific host tissues (25).
Most young children in whom an invasive K. kingae disease developed have been otherwise healthy. In contrast, children >4 years of age and adults who become infected frequently have underlying conditions such as congenital heart diseases, chronic renal failure, or a variety of primary immunodeficiencies (16).
The prevalence rate in healthy children during the second year of life ranges between 10% and 12% (7,28), which coincides with the peak attack rate of invasive infections (16). The colonization rate drops substantially in older children and adults (7,28,29). Pharyngeal carriage of K. kingae and occurrence of disease before a child is 6 months of age are exceptions, indicating that maternal immunity and limited social contact provide protection (7,16,30).
Diagnosis of K. kingae Infections
The clinical features of invasive K. kingae infections (other than endocarditis) are usually mild, and diagnosis requires a high level of suspicion by clinicians. Many patients with K. kingae–associated joint or bone disease are afebrile and blood leukocyte counts, C-reactive protein levels, and erythrocyte sedimentation rates are frequently normal (16,31,32). Although bacteremia with no focus is the second most frequent manifestation of invasive K. kingae infections, the condition is probably unsuspected and the diagnosis is likely missed in a large number of cases. The current guidelines for managing illness in young, febrile children with no apparent source of infection, which use body temperature and leukocyte count as criteria for obtaining blood cultures (33), are not sensitive enough for detecting occult K. kingae bacteremia because many infected children have a low-grade fever and may or may not have leukocytosis.
Detection of K. kingae infections by culture is highly dependent on the use of adequate laboratory techniques. The recovery of K. kingae from synovial fluid and bone exudates seeded onto routine solid culture media is suboptimal (8). The yield of cultures can be substantially improved by inoculating clinical specimens into aerobic blood culture vials (BCVs) from a variety of commercial systems (8). Attempts to isolate the organism from synovial fluid or bone exudates on routine solid media succeeded in 2 of 25 patients, whereas inoculation of these specimens into aerobic BACTEC (Becton Dickinson, Cockeysville, MD, USA) BCVs yielded the organism in all cases after a median incubation of 4 days (8). When specimens from BCVs determined to be positive by the automated blood culture instrument were subcultured onto a blood–agar plate of trypticase soy agar with 5% sheep blood hemoglobin or chocolate agar, K. kingae grew readily, indicating that routine solid media are able to support the nutritional requirements of the organism. This observation suggests that skeletal system exudates exert a detrimental effect on this fastidious bacterium, and dilution of purulent material in a large volume of broth decreases the concentration of inhibitory factors, improving its recovery (8). In studies conducted in Israel and France in which BCVs were routinely inoculated with synovial fluid aspirates from young children who had arthritis, K. kingae was isolated in 48% of the patients with culture-proven disease (8,9). Conversely, when BCVs are not used, many K. kingae infections will be overlooked and labeled as culture-negative septic arthritis (34).
In recent years, development of NAAAs has further improved the diagnosis of K. kingae from skeletal system exudates. Use of this novel technique facilitates detection of difficult-to-culture organisms, enables bacteriologic diagnosis in patients already treated with antibacterial drugs, reduces time to detection, and facilitates precise identification of unusual species (10–14). The procedure consists of extraction of bacterial DNA from the synovial fluid sample, a PCR amplification step in which primers target the 16S rDNA, the 23S rDNA, or the rpo genes that are ubiquitous to all bacteria, then sequencing of the amplicon, and comparison of results with those kept in a broad database curator (such as GenBank) to enable precise species identification. Alternately, the specimen may be subjected to amplification by PCR by using species-specific primers that recognize the most plausible pathogens. Use of NAAAs has confirmed that in countries where this topic has been studied, K. kingae is the most common etiologic agent of septic arthritis in children <3 years of age. NAAA use detected the presence of K. kingae–specific DNA sequences, including in cases in which seeding of synovial fluid specimens into BCVs failed to recover the organism and shortened the time required to detect and identify the bacterium from 3–4 days to <24 hours (10–14). It was, then, natural that this sensitive approach was consequently adopted to study respiratory colonization by K. kingae and its connection to invasive disease.
In an outbreak investigated by Bidet et al., the cause of the skeletal infections was determined by sequential use of real-time PCR targeting the K. kingae–specific toxin rtxA and the cpn60 genes (35). The same method was used to investigate the prevalence of K. kingae among attendees of the index daycare facility. The organism was recovered by culture in 6 asymptomatic carriers, whereas the NAAs detected 5 additional carriers (35). MLST and sequencing of the rtxA amplicon were performed directly on the joint fluid sample from the child who had arthritis and on the recovered pharyngeal isolates. All carriage isolates and the arthritis strain belonged to MLST 25 and shared rtxA allele 1, which is among the most common genotypes involved in joint and bone infections in France (26,35). It should be pointed out that, despite the increased sensitivity of NAAAs, when evaluating the efficacy of prophylactic antibacterial drug administration for eradicating K. kingae from colonized children, cultures have the obvious advantage of detecting living bacteria, whereas the viability of K. kingae organisms in PCR-positive/culture-negative specimens is questionable.
In addition detecting invasive K. kingae disease in patients, searching for asymptomatic but colonized children is crucial in the investigation of an outbreak in a daycare facility to assess the full extent of the contagion, identify attendees at risk for clinical disease, and evaluate the effects of prophylactic antibacterial drugs. Because of the high density of the resident bacterial flora and the relatively slow growth of K. kingae, detecting the organism in pharyngeal cultures is difficult. A differential and selective medium consisting of blood agar with 2 mg/mL of vancomycin (BAV medium) added has been developed to improve recovery of K. kingae from respiratory cultures (36). This formulation facilitates recognition of β-hemolytic K. kingae colonies by inhibiting growth of competitive gram-positive bacteria. In a blinded evaluation, the BAV medium detected 43 (97.7%) of 44 pharyngeal cultures positive for the organism; 10 (22.7%) positive cultures were identified on routine blood-agar plates (p<0.001) (36). If the original BAV formulation (36) or a similar medium (23) had not been used, and a chocolate-based agar substituted (in which the faint ring of hemolysis surrounding K. kingae colonies could not be recognized), carriers of the organism might not have been detected among 27 asymptomatic attendees of a daycare facility where a cluster of invasive disease occurred (37).
The K. kingae colonization rate is substantially enhanced among children in daycare centers. In an 11–month longitudinal study, 35 (72.9%) of 48 daycare center attendees carried the organism at least once and an average of 27.5% of the children were colonized at any given time (20). Molecular typing of isolates from asymptomatic colonized attendees showed genotypic similarities, indicating person-to-person transmission of the organism in the facility (21). Two K. kingae strains represented 28.0% and 46.0% of all isolates, demonstrating that some strains are particularly successful in colonizing mucosa. Children harbored the same strain continuously or intermittently for weeks or months, and then it was replaced by a new strain, showing that carriage is a dynamic process in which there is frequent turnover of colonizing organisms, as observed for other respiratory pathogens (21,28). Despite the high prevalence of the organism in the daycare center, an invasive K. kingae infection did not develop in any of the attendees in the course of the follow-up period.
The link between out-of-home child care and K. kingae carriage was recently confirmed in a study conducted among 1,277 children <5 years of age who were referred to a pediatric emergency department (7). Daycare attendance was strongly associated with K. kingae carriage after controlling for other variables (odds ratio 9.66 [95% CI 2.99–31.15], p<0.001) (7). Surveillance studies not only have showed that K. kingae organisms colonizing attendees of a given daycare center are frequently identical, but have also demonstrated that carried strains differ between facilities located close together, indicating that each daycare center is like an independent epidemiologic unit (35,37–39).
Considering these findings, it is not surprising that clusters of proven and presumptive cases of invasive K. kingae disease have been detected in daycare centers in France (35), the United States (37,38), and Israel (39), including 2 recent and still unreported outbreaks (P. Yagupsky, unpub. data) (Table 1). A presumptive case was defined as bacteriologically unconfirmed invasive disease, consistent with the clinical features of K. kingae infections, among daycare center attendees 6 months–3 years of age, within 1 month of an infection confirmed by culture and/or NAAAs. These events have been characterized by the simultaneous or consecutive occurrence of multiple cases of disease that included the entire clinical spectrum of K. kingae infections (septic arthritis, osteomyelitis, bacteremia, spondylodiscitis, cellulitis, and fatal meningitis complicating endocarditis). The 3 patients in the 2005 cluster in Israel (39) and 4 of the 5 patients reported in France (35) had bone infections, supporting the concept that certain K. kingae clones exhibit specific tissue tropism (25).
Three of the 6 K. kingae illness outbreaks in daycare centers reported during 2003–2013 were detected in Israel, a small country with a population of 8,000,000 inhabitants (35,37–39; P. Yagupsky, unpub. data). Although Israel’s relatively high annual birthrate compared with Western countries (18.7 per 1,000 population, Israel Central Bureau of Statistics, vs. 10.9 per 1,000 in the European Union, Demography Report 2010, Eurostat), and the widespread and early daycare center attendance could partially account for this observation, it seems plausible that similar events occur worldwide but are frequently overlooked.
The epidemiologic investigation of these outbreaks revealed that the K. kingae colonization rate among asymptomatic attendees to the daycare centers where clinical cases were detected was unusually high (up to 52.9% in a United States cluster, as determined by culture [38], and 68.8% in France, as demonstrated by a sensitive NAAA [35]) (Table 2), and all pharyngeal isolates detected in the classrooms where disease occurred were genotypically identical and indistinguishable from the patients’ clinical isolates. The age of the colonized or infected daycare center attendees coincided with the age of increased susceptibility to K. kingae carriage and disease (7,15,16,28); the organism was detected in the pharyngeal culture of only1 of the caregivers, supporting the hypothesis that healthy adults rarely carry K. kingae (29).
Despite a background carriage rate as high as 5%–12%, the incidence of invasive K. kingae infections reported in Israel was 9.4 per 100,000 children <5 years of age per year only (16), and the calculated annual risk of developing K. kingae osteomyelitis or septic arthritis for young carriers in Switzerland was <1% (40). When data from the 6 clusters of invasive disease detected in daycare centers are pooled, a documented or presumptive K. kingae infection developed in 1 in 7 classmates within a 1-month period (35,37–39), indicating that the outbreak strains combined enhanced colonization fitness, high transmissibility, and remarkable virulence.
The strain responsible for the cluster of osteomyelitis detected in Israel in 2005 (39) belonged to PFGE clone K and MLST-6 that ranked second among strains carried in southern Israel by healthy Jewish children (20), and was responsible for the excess of K. kingae illness observed in the Jewish population of the region over the previous 2 decades, causing 41.7% of all invasive strains isolated in this ethnic group (27). The clone that caused an outbreak in daycare centers in North Carolina, USA (37), represented 11.3% of all organisms carried by healthy children in southern Israel (24) and 11.6% of all isolates from patients in Israel who had invasive infections (27). The international distribution of these clones indicates that the clusters of disease are frequently caused by highly successful strains that exhibit enhanced capability for local and long-range dissemination (26).
To prevent further cases of disease and eradicate the invasive strain, prophylactic antibacterial drugs have been administered to children attending the facilities where clusters of K. kingae were detected. Rifampin was chosen because K. kingae is especially susceptible to that antibacterial drug (35). Rifampin is secreted in saliva and reaches high concentrations in the upper respiratory mucosa and has shown efficacy in the eradication of colonization by N. meningitidis and H. influenzae type b, and in disease prevention in daycare centers (2). However, because only partial success was achieved with this drug in the Minnesota cluster (38), high-dose amoxicillin was added to the regimen in 2 more recent outbreaks (37,39). Following administration of antibacterial drugs, a respiratory carriage of the organism decreased, but complete eradication occurred only in the Durham, North Carolina daycare center (37), and new colonization of several attendees by the original strain was later observed (35,38,39). Persistence of the organism in the facility was not caused by bacterial resistance to the administered antibacterial drugs, however (35,37,39). Similar observations have been made in outbreaks in daycare centers caused by H. influenzae type b and pneumococci (2). Poor compliance or failure to administer prophylactic antibacterial drugs to family contacts could have resulted in incomplete suppression of the reservoir and recurrent dissemination of the strain in the facility (2).
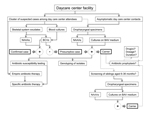
Because antibacterial drugs have been relatively ineffective in eradicating K. kingae carriage, the need for antibacterial drug prophylaxis in the setting of a cluster of invasive disease is being disputed (40). Notably, however, after administration of antibacterial drugs to the asymptomatic children, no further cases of disease were detected in the affected daycare centers (35,37–39), even when a few children continue to carry the invasive strain. Reducing the bacterial density among colonized children by antibacterial drug administration or an effective immune response induced by prolonged mucosal carriage may have been sufficient to prevent new cases of infection. Improved hygiene and institution of other infection control measures could also have limited further transmission of the organism. The Figure depicts an algorithm aimed to guide the investigation and management of clusters of invasive K. kingae in daycare facilities, including areas of controversy.
In recent years, clusters of invasive K. kingae infections among attendees of daycare centers have been reported, although because of the low rate of testing for this pathogen, many events are probably overlooked. Detection of these events requires a high level of clinical suspicion, use of sensitive culture techniques and NAAAs, and familiarity of the clinical microbiology laboratory with the identification of this elusive pathogen. The same improved detection methods should be employed for the thorough investigation of these clusters, and recovered K. kingae isolates should be genotyped and compared. Many issues remain unsettled, including whether antibacterial drugs should be administered prophylactically to daycare center contacts of an index case-patient, to young siblings of clinical case-patients, and to carriers, or whether antibacterial drugs should be offered to confirmed carriers only, or limited to children with disease. If administration of antibacterial drug prophylaxis is decided on, the preferred drug regimen will have to be determined. Whether an epidemiologic investigation should be carried out in daycare centers after detection of a single case of disease also remains to be determined.
Dr Yagupsky is a pediatrician and clinical microbiologist, and the head of the Clinical Microbiology Laboratory of the Soroka University Medical Center, Beer-Sheva, Israel. His main research interests include Kingella kingae infections, human brucellosis, and pediatric skeletal system infections.
References
- Robinson J. Infectious diseases in schools and child care facilities. Pediatr Rev. 2001;22:39–46 . DOIPubMedGoogle Scholar
- Murphy TV, McCracken GH Jr, Moore BS, Gulig PA, Hansen HJ. Haemophilus influenzae type b disease after rifampin prophylaxis in a day care center: possible reasons for its failure. Pediatr Infect Dis. 1983;2:193–8. DOIPubMedGoogle Scholar
- Weintraub A. Immunology of bacterial polysaccharide antigens. Carbohydr Res. 2003;338:2539–47. DOIPubMedGoogle Scholar
- Lu N, Samuels ME, Baker SL, Glover SH, Sanders JM. Child day care risks of common infectious diseases revisited. Child Care Health Dev. 2004;30:361–8 and. DOIPubMedGoogle Scholar
- Huskins WC. Transmission and control of infections in out-of-home child care. Pediatr Infect Dis J. 2000;19:S106–10. DOIPubMedGoogle Scholar
- Osterholm MT. Infectious diseases in child day care: an overview. Pediatrics. 1994;94:987–90 .PubMedGoogle Scholar
- Amit U, Dagan R, Yagupsky P. Prevalence of pharyngeal carriage of Kingella kingae in young children and risk factors for colonization. Pediatr Infect Dis J. 2013;32:191–3. DOIPubMedGoogle Scholar
- Yagupsky P, Dagan R, Howard CW, Einhorn M, Kassis I, Simu A. High prevalence of Kingella kingae in joint fluid from children with septic arthritis revealed by the BACTEC blood culture system. J Clin Microbiol. 1992;30:1278–81 .PubMedGoogle Scholar
- Moumile K, Merckx J, Glorion C, Pouliquen JC, Berche P, Ferroni A. Bacterial aetiology of acute osteoarticular infections in children. Acta Paediatr. 2005;94:419–22. DOIPubMedGoogle Scholar
- Cherkaoui A, Ceroni D, Emonet S, Lefevre Y, Schrenzel J. Molecular diagnosis of Kingella kingae osteoarticular infections by specific real-time PCR assay. J Med Microbiol. 2009;58:65–8. DOIPubMedGoogle Scholar
- Moumile K, Merckx J, Glorion C, Berche P, Ferroni A. Osteoarticular infections caused by Kingella kingae in children: contribution of polymerase chain reaction to the microbiologic diagnosis. Pediatr Infect Dis J. 2003;22:837–9. DOIPubMedGoogle Scholar
- Lehours P, Freydière AM, Richer O, Burucoa C, Boisset S, Lanotte F, The rtxA toxin gene of Kingella kingae: a pertinent target for molecular diagnosis of osteoarticular infections. J Clin Microbiol. 2011;49:1245–50. DOIPubMedGoogle Scholar
- Chometon S, Benito Y, Chaker M, Boisset S, Ploton C, Bérard J, Specific real-time polymerase chain reaction places Kingella kingae as the most common cause of osteoarticular infections in young children. Pediatr Infect Dis J. 2007;26:377–81. DOIPubMedGoogle Scholar
- Ilharreborde B, Bidet P, Lorrot M, Even J, Mariani-Kurkdjian P, Ligouri S, New real-time PCR-based method for Kingella kingae DNA detection: application to samples collected from 89 children with acute arthritis. J Clin Microbiol. 2009;47:1837–41. DOIPubMedGoogle Scholar
- Yagupsky P, Bar-Ziv Y, Howard CB, Dagan R. Epidemiology, etiology, and clinical features of septic arthritis in children younger than 24 months. Arch Pediatr Adolesc Med. 1995;149:537–40. DOIPubMedGoogle Scholar
- Dubnov-Raz G, Ephros M, Garty BZ, Schlesinger Y, Maayan-Metzger A, Hasson J, Invasive pediatric Kingella kingae infections: a nationwide collaborative study. Pediatr Infect Dis J. 2010;29:639–43. DOIPubMedGoogle Scholar
- Porsch EA, Kehl-Fie TE, Geme JW III. Modulation of Kingella kingae adherence to human epithelial cells by type IV pili, capsule, and a novel trimeric autotransporter. MBio. 2012;3:e00372–12 .DOIPubMedGoogle Scholar
- Starr KF, Porsch EA, Heiss C, Black I, Azadi P, St Geme JW III. Characterization of the Kingella kingae polysaccharide capsule and exopolysaccharide. PLoS ONE. 2013;8:e75409. DOIPubMedGoogle Scholar
- Kehl-Fie TE, St Geme JW III. Identification and characterization of an RTX toxin in the emerging pathogen Kingella kingae. J Bacteriol. 2007;189:430–6. DOIPubMedGoogle Scholar
- Yagupsky P, Dagan R, Prajgrod F, Merires M. Respiratory carriage of Kingella kingae among healthy children. Pediatr Infect Dis J. 1995;14:673–7. DOIPubMedGoogle Scholar
- Slonim A, Walker ES, Mishori E, Porat N, Dagan R, Yagupsky P. Person-to-person transmission of Kingella kingae among day care center attendees. J Infect Dis. 1998;178:1843–6. DOIPubMedGoogle Scholar
- Yagupsky P, Porat N, Pinco E. Pharyngeal colonization by Kingella kingae in children with invasive disease. Pediatr Infect Dis J. 2009;28:155–7. DOIPubMedGoogle Scholar
- Basmaci R, Ilharreborde B, Bidet P, Doit C, Lorrot M, Mazda K, Isolation of Kingella kingae in the oropharynx during K. kingae arthritis on children. Clin Microbiol Infect. 2012;18:e134–6. DOIPubMedGoogle Scholar
- Yagupsky P, Weiss-Salz I, Fluss R, Freedman L, Peled N, Trefler R, Dissemination of Kingella kingae in the community and long-term persistence of invasive clones. Pediatr Infect Dis J. 2009;28:707–10. DOIPubMedGoogle Scholar
- Amit U, Porat N, Basmaci R, Bidet P, Bonacorsi S, Dagan R, Genotyping of invasive Kingella kingae isolates reveals predominant clones and association with specific clinical syndromes. Clin Infect Dis. 2012;55:1074–9. DOIPubMedGoogle Scholar
- Basmaci R, Yagupsky P, Ilharreborde B, Guyot K, Porat N, Chomton M, Multilocus sequence typing and rtxA toxin gene sequencing analysis of Kingella kingae isolates demonstrates genetic diversity and international clones. PLoS ONE. 2012;7:e38078. DOIPubMedGoogle Scholar
- Amit U, Dagan R, Porat N, Trefler R, Yagupsky P. Epidemiology of invasive Kingella kingae infections in two distinct pediatric populations cohabiting in one geographic area. Pediatr Infect Dis J. 2012;31:415–7. DOIPubMedGoogle Scholar
- Amit U, Flaishmakher S, Dagan R, Porat N, Yagupsky P. Age-dependent carriage of Kingella kingae in young children and turnover of colonizing strains. J Pediatr Infect Dis Soc [Internet]. In Press [cited 2013 Oct 5].
- Yagupsky P, Peled N, Katz O. Epidemiological features of invasive Kingella kingae infections and respiratory carriage of the organism. J Clin Microbiol. 2002;40:4180–4. DOIPubMedGoogle Scholar
- Slonim A, Steiner M, Yagupsky P. Immune response to invasive Kingella kingae infections, age-related incidence of disease, and levels of antibody to outer-membrane proteins. Clin Infect Dis. 2003;37:521–7. DOIPubMedGoogle Scholar
- Basmaci R, Lorrot M, Bidet P, Doit C, Vitoux C, Penneçot G, Comparison of clinical and biologic features of Kingella kingae and Staphylococcus aureus arthritis at initial evaluation. Pediatr Infect Dis J. 2011;30:902–4. DOIPubMedGoogle Scholar
- Ceroni D, Cherkaoui A, Combescure C, Francois P, Kaelin A, Schrenzel J. Differentiating osteoarticular infections caused by Kingella kingae from those due to typical pathogens in young children. Pediatr Infect Dis J. 2011;30:906–9. DOIPubMedGoogle Scholar
- Baraff LJ, Bass JW, Fleisher GR, Klein JO, McCracken GH, Powell KR, Practice guideline for the management of infants and children 0 to 36 months of age with fever without source. Agency for Health Policy and Research. Ann Emerg Med. 1993;22:1198–210. DOIPubMedGoogle Scholar
- Infectious Diseases Society of America Emerging Infections Network. Brief Report: Kingella kingae infections in children—United States, June 2001–November 2002. MMWR Morb Mortal Wkly Rep. 2004;53:244 .
- Bidet P, Collin E, Basmaci R, Courroux C, Prisse V, Dufour V, Investigation of an outbreak of osteoarticular infections caused by Kingella kingae in a childcare center using molecular techniques. Pediatr Infect Dis J. 2013;32:558–60. DOIPubMedGoogle Scholar
- Yagupsky P, Merires M, Bahar J, Dagan R. Evaluation of a novel vancomycin-containing medium for primary isolation of Kingella kingae from upper respiratory tract specimens. J Clin Microbiol. 1995;33:1426–7 .PubMedGoogle Scholar
- Seña AC, Seed P, Nicholson B, Joyce M, Cunningham CK. Kingella kingae endocarditis and a cluster investigation among daycare attendees. Pediatr Infect Dis J. 2010;29:86–8 and. DOIPubMedGoogle Scholar
- Kiang KM, Ogunmodede F, Juni BA, Boxrud DJ, Glennen A, Bartkus JM, Outbreak of osteomyelitis/septic arthritis caused by Kingella kingae among child care center attendees. Pediatrics. 2005;116:e206–13. DOIPubMedGoogle Scholar
- Yagupsky P, Erlich Y, Ariela S, Trefler R, Porat N. Outbreak of Kingella kingae skeletal system infections in children in daycare. Pediatr Infect Dis J. 2006;25:526–32. DOIPubMedGoogle Scholar
- Ceroni D, Dubois-Ferrière V, Anderson R, Combescure C. Lamah, Cherkaoui A, et al. Small risk of osteoarticular infections in children with asymptomatic carriage of Kingella kingae. Pediatr Infect Dis J. 2012;31:983–5.
Figure
Tables
Follow Up
Earning CME Credit
To obtain credit, you should first read the journal article. After reading the article, you should be able to answer the following, related, multiple-choice questions. To complete the questions (with a minimum 75% passing score) and earn continuing medical education (CME) credit, please go to www.medscape.org/journal/eid. Credit cannot be obtained for tests completed on paper, although you may use the worksheet below to keep a record of your answers. You must be a registered user on Medscape.org. If you are not registered on Medscape.org, please click on the New Users: Free Registration link on the left hand side of the website to register. Only one answer is correct for each question. Once you successfully answer all post-test questions you will be able to view and/or print your certificate. For questions regarding the content of this activity, contact the accredited provider, CME@medscape.net. For technical assistance, contact CME@webmd.net. American Medical Association’s Physician’s Recognition Award (AMA PRA) credits are accepted in the US as evidence of participation in CME activities. For further information on this award, please refer to http://www.ama-assn.org/ama/pub/category/2922.html. The AMA has determined that physicians not licensed in the US who participate in this CME activity are eligible for AMA PRA Category 1 Credits™. Through agreements that the AMA has made with agencies in some countries, AMA PRA credit may be acceptable as evidence of participation in CME activities. If you are not licensed in the US, please complete the questions online, print the certificate and present it to your national medical association for review.
Article Title: Outbreaks of Kingella kingae Infections in Daycare Facilities
CME Questions
1. You are seeing a 3-year-old boy who was just diagnosed with osteomyelitis of the left tibia. The child attends day care 5 times per week. You consider whether this patient might have a hematogenous infection with Kingella kingae. Besides the bones, this organism displays a tropism for what other anatomic site?
A. Esophagus
B. Kidneys
C. Brain
D. Heart
2. What should you consider regarding the clinical presentation of invasive infection with K. kingae?
A. Most children with invasive infection have multiple chronic medical conditions
B. Nearly all children have fever
C. A normal serum C-reactive protein level rules out the possibility of K. kingae osteomyelitis
D. The typical presentation of K. kingae bacteremia makes the diagnosis easy to miss
3. You initiate laboratory work to identify possible K. kingae infection for this patient. Which of the following statements regarding the diagnostic process for K. kingae infections is most accurate?
A. Less than 1% of children between 12 and 24 months old harbor the K. kingae bacterium
B. Bone exudates in particular grow well on routine solid culture media
C. Medium consisting of blood-agar with added 2 mg/mL of vancomycin should be avoided in respiratory cultures
D. Nucleic acid amplification assays have improved the sensitivity of detection of K. kingae and reduce the time needed for diagnosis
4. The patient is diagnosed with K. kingae osteomyelitis. What should you consider regarding a possible outbreak of infection in his day care class?
A. Previous data suggest that the risk for another K. kingae infection within 1 month exceeds 10%
B. Isoniazid is the drug of choice as prophylaxis
C. Azithromycin is the drug of choice as prophylaxis
D. Multiple antibiotic prescriptions are associated with 100% eradication of K. kingae
Activity Evaluation
|
1. The activity supported the learning objectives. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
2. The material was organized clearly for learning to occur. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
3. The content learned from this activity will impact my practice. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
4. The activity was presented objectively and free of commercial bias. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
Related Links
Table of Contents – Volume 20, Number 5—May 2014
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Pablo Yagupsky, Clinical Microbiology Laboratory, Soroka University Medical Center, Ben-Gurion University of the Negev, Beer-Sheva 84101, IsraelPablo Yagupsky, Clinical Microbiology Laboratory, Soroka University Medical Center, Ben-Gurion University of the Negev, Beer-Sheva 84101, Israel
Top