Volume 21, Number 1—January 2015
Dispatch
Hare-to-Human Transmission of Francisella tularensis subsp. holarctica, Germany
Cite This Article
Citation for Media
Abstract
In November 2012, a group of 7 persons who participated in a hare hunt in North Rhine-Westphalia, Germany, acquired tularemia. Two F. tularensis subsp. holarctica isolates were cultivated from human and hare biopsy material. Both isolates belonged to the FTN002–00 genetic subclade (derived for single nucleotide polymorphisms B.10 and B.18), thus indicating likely hare-to-human transmission.
Tularemia is a zoonotic disease caused by the gram-negative bacterium Francisella tularensis (1). Currently, there are 4 validly published subspecies. F. tularensis subsp. tularensis is the most virulent subspecies and occurs only in North America. F. tularensis subsp. holarctica is less virulent and occurs throughout the Northern hemisphere. F. tularensis subsp. mediasiatica was isolated in central Asia, and F. tularensis subsp. novicida, which has low virulence in humans, seems to be distributed globally (2).
Various PCR-based assays have been established for the detection of F. tularensis or for the diagnosis of tularemia. An accurate population structure has been defined by using single nucleotide polymorphisms (SNPs) and insertion/deletion mutations (INDELs) with potential canonical properties. Currently, this population is divided into 4 major genetic clades: B.4, B.6, B.12, and B.16 (3–6). The taxonomic nomenclature of major clades in F. tularensis subsp. holarctica is based on clade-specific canonical SNP markers (3,4). In Europe, the strains of clades B.12 and B.6 dominate (6). The latter is found particularly in large areas in northern, western, and central Europe, including Germany (5–9).
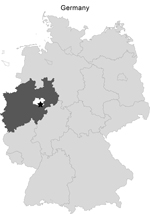
On November 2, 2012, 15 European brown hares (Lepus europaeus) were shot during a hunt in Rüthen-Meiste, district Soest in the federal state of North Rhine-Westphalia, Germany (Figure). The animals seemed healthy and showed normal escape behavior. Upon inspection, the animals that had been shot showed no signs of disease. Consequently, all animals were skinned, eviscerated, and dissected. Portioning of the hares was done 2 days later. Within a few days, 7 healthy persons who had contact with the hare carcasses showed varied symptoms of illness. Tularemia was suspected because of the signs and symptoms in combination with exposure in a tularemia-endemic area. Exposure, clinical symptoms, and time of onset of symptoms of all patients (A to G) are described in the Table. All patients were treated successfully with doxycycline.
Human serum samples were collected about 5 weeks after infection from patients C and E and pleural fluid was obtained from patient E. On day 2 of incubation, the human isolate (12T0062) showed small pale-white to gray colonies on Columbia blood agar and chocolate agar, whereas no growth occurred on MacConkey agar plates. Ten organ specimens (from aorta, back and thigh muscles, lymph nodes, spinal cord) from 4 of the hares handled by the patients were tested for Francisella spp. in the National Reference Laboratory for Tularemia at the Friedrich-Loeffler-Institut in Jena, Germany. From all hare organs, only a single Francisella sp. was isolated from a spinal cord sample (13T0009) on cysteine heart agar Becton Dickinson GmbH, Heidelberg, Germany), which contains antibiotics. The strains were susceptible to erythromycin with inhibition zones between 22 and 24 mm corresponding to biovar I. Details of further methods that have been applied in the study are shown in the Technical Appendix.
Both samples were identified as F. tularensis subsp. holarctica of clade B.6. For B.19, the SNP was C; results of Ftind33 and Ftind38 assays were IN, and the result of the Ftind49 assay was DEL. Both samples had T for SNP B.7, G for B.10 and T for B.18. Therefore, the strains were considered derived from SNPs B.10 and B.18. Blood serum samples of patients C and E were positive for F. tularensis with very high values of the optical density in the ELISA, 2.886 and 3.121, respectively.
The re-emergence of tularemia in Germany has been described in previous studies (10). Infected hares are believed to be the sources of most cases of tularemia in Germany. However, to our knowledge, route of transmission has not been demonstrated by isolation and genotyping of the pathogen from the suspected source and the patient (11–13). In this study, we therefore described not only the clinical and epidemiologic data and the laboratory diagnostic findings for determining tularemia, but also the results of genotyping the Francisella spp. isolated from epidemiologically linked hares and humans.
All 7 infected persons (A to G) in this outbreak showed influenza-like symptoms of varying intensity (Table), but symptoms were also related to the route of infection. The 5 patients (A to E) who had fever as well as respiratory and topical symptoms were exposed to aerosols and had direct skin contact when skinning and processing hare carcasses. The 2 patients (F and G) who portioned the meat had lesions on their hands, enlarged lymph nodes, and fever.
The isolated F. tularensis subsp. holarctica strains were susceptible to erythromycin and thus belong to the F. tularensis subsp. holarctica biovar I group. Because of the inability of the duplex PCR assay to distinguish between F. tularensis subsp. holarctica strains (8), we performed a combined SNP and INDEL analysis using real-time PCR. Here, we were able to isolate F. tularensis subsp. holarctica biovar I strains from a hare and a human; both isolates could be assigned to the genetic clade B.6 in the first order of discrimination [B.19(C), Ftind33(IN), Ftind38(IN), and Ftind49(DEL)]. The isolates also showed an identical genotyping profile for B.7(T), B.10(G), B.18(T) in the second order of discrimination, which corresponds with a previously described subclade represented by the strain FTNF002–00 that was isolated from a patient from France who had bacteremia (3,4). Thus, the genetic subtyping results are consistent with the proposed transmission route of the epidemiologically linked (hare–human transmission) F. tularensis subsp., since both belonged to the same genetic subclade.
The current phylogeography of F. tularensis subsp. holarctica revealed that 2 major groups of virulent strains exist in Europe (5). In the western European countries of Spain, France, Switzerland, and Italy, strains of the FTNF002–00 group dominate, whereas strains of clade B.12 seem to predominate in eastern and northern Europe as reported from Austria, Czech Republic, Finland, Georgia, Hungary, Romania, Russia, Slovakia, Sweden, and Ukraine (4,5,8,13). Vogler et al. (4) suggest that it is likely that the spread of strains in subclade FTNF002–00 throughout France and the Iberian Peninsula was a very recent event. In Germany, isolates of both groups have been identified and a sharp dividing line in terms of occurrence of the clades B.12 and B.6 from the northwest to the southeast of the country has been shown (8). The reasons for this are not known; possible causes could be environmental and epidemiologic differences. Alternately, a mixture of both genetic clades and biovars have been reported in Bulgaria, Kazakhstan, Norway, Russia, and Sweden (7,14,15).
The genome of F. tularensis subsp. holarctica is highly conserved and strains can hardly be discriminated. Therefore, the discriminatory power of the applied assays is limited and other field isolates from this area may show identical characteristics (H. Tomaso, unpub. data). For epidemiologic and forensic purposes, whole-genome sequencing of a multitude of strains from well-documented outbreaks and the surrounding areas should be performed to clarify and possibly quantify the genetic changes that can finally confirm or rule out the route of transmission.
Dr. Otto is the head of the working group, Noro- and Rotaviruses, and an employee of the National Reference Laboratory of Tularemia at the Federal Institute of Bacterial Infections and Zoonoses of the Federal Research Institute of Animal Health, Friedrich Loeffler Institut, Jena, Germany. His interests are focused on the development of diagnostic methods, the incidence and epidemiology of Francisella spp., and other bacterial zoonotic agents in wild animals.
Acknowledgment
We thank Kerstin Cerncic, Renate Danner, Anja Hackbart, Byrgit Hofmann, Wolfram Maginot, Petra Sippach, and Karola Zmuda for their excellent technical assistance; and Franz-Josef Röper for accurate documentation of the data of hunting and the disease outbreak.
References
- Petersen JM, Schriefer ME. Tularemia: emergence/re-emergence. Vet Res. 2005;36:455–67. DOIPubMedGoogle Scholar
- Keim P, Johansson A, Wagner DM. Molecular epidemiology, evolution, and ecology of Francisella. Ann N Y Acad Sci. 2007;1105:30–66 . DOIPubMedGoogle Scholar
- Svensson K, Granberg M, Karlsson L, Neubauerova V, Forsman M, Johannson A. A real-time PCR assay for hierarchical identification of Francisella isolates. PLoS ONE. 2009;4:e8360 . DOIPubMedGoogle Scholar
- Vogler AJ, Birdsell D, Price LB, Bowers JR, Beckstrom-Sternberg SM, Auerbach RK, Phylogeography of Francisella tularensis: global expansion of a highly fit clone. J Bacteriol. 2009b;191:2474–84. DOIPubMedGoogle Scholar
- Karlsson E, Svensson K, Lindgren P, Byström M, Sjödin A, Forsman M, The phylogeographic pattern of Francisella tularensis in Sweden indicates a Scandinavian origin of Eurosiberian tularaemia. Environ Microbiol. 2013;15:634–45. DOIPubMedGoogle Scholar
- Gyuranecz M, Birdsell DN, Splettstoesser M, Seibold E, Beckstrom-Sternberg SM, Makrai L, Phylogeography of Francisella tularensis subsp. holarctica, Europe. Emerg Infect Dis. 2012;18:290–3. DOIPubMedGoogle Scholar
- Dempsey MP, Dobson M, Zhang C, Zhang M, Lion C, Gutiérrez CB, Genomic deletion marking an emerging subclone of Francisella tularensis subsp. holarctica in France and the Iberian Peninsula. Appl Environ Microbiol. 2007;73:7465–70. DOIPubMedGoogle Scholar
- Müller W, Hotzel H, Otto P, Karger A, Bettin B, Bocklisch H, German Francisella tularensis isolates from European brown hares (Lepus europaeus) reveal genetic and phenotypic diversity. BMC Microbiol. 2013;13:61. DOIPubMedGoogle Scholar
- Vogler AJ, Birdsell DN, Lee J, Vaissaire J, Doujet CL, Lapalus M, Phylogeography of Francisella tularensis ssp. holarctica in France. Lett Appl Microbiol. 2011;52:177–80. DOIPubMedGoogle Scholar
- Grunow R, Priebe HS. Tularemia— occurrence in Germany, the analysis based on the data reporting 1949 to 2006 [in German]. Epidemiologisches Bulletin. 2007;7:51–6. https://www.rki.de/DE/Content/Infekt/EpidBull/Archiv/2007/Ausgabenlinks/07_07.pdf?__blob=publicationFile [cited 2014 Nov 18]
- Robert-Koch-Institut. Tularemia—two cases of disease after processing and consumption of a hare [in German]. Epidemiologisches Bulletin. 2002;9:71–2. https://www.rki.de/DE/Content/Infekt/EpidBull/Archiv/2002/Ausgabenlinks/09_02.pdf?__blob=publicationFile [cited 2014 Nov 18]
- Hofstetter I, Eckert J, Splettstoesser W, Hauri A. Tularaemia outbreak in hare hunters in the Darmstadt-Dieburg district, Germany. Euro Surveill. 2006;11:E060119.3 .PubMedGoogle Scholar
- Robert-Koch-Institut. Annual report on zoonoses in 2002 [in German]. Epidemiologisches Bulletin. 2003;46:379. https://www.rki.de/DE/Content/Infekt/EpidBull/Archiv/2003/Ausgabenlinks/46_03.pdf?__blob=publicationFile [cited 2014 Nov 18]
- Forsman M, Sandström G, Sjöstedt A. Analysis of 16S ribosomal DNA sequences of Francisella strains and utilization for determination of the phylogeny of the genus and for identification of strains by PCR. Int J Syst Bacteriol. 1994;44:38–46. DOIPubMedGoogle Scholar
- Petersen JM, Molins CR. Subpopulations of Francisella tularensis ssp. tularensis and holarctica: identification and associated epidemiology. Future Microbiol. 2010;5:649–61 . DOIPubMedGoogle Scholar
Figure
Table
Cite This ArticleTable of Contents – Volume 21, Number 1—January 2015
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Herbert Tomaso, Friedrich-Loeffler-Institut, Federal Research Institute for Animal Health, Institute of Bacterial Infections and Zoonoses, Naumburger Str. 96A, 07743 Jena, Germany
Top