Volume 21, Number 11—November 2015
Dispatch
RmtC and RmtF 16S rRNA Methyltransferase in NDM-1–Producing Pseudomonas aeruginosa
Cite This Article
Citation for Media
Abstract
We investigated 16S rRNA methyltransferases in 38 blaNDM-1–positive Pseudomonas aeruginosa isolates and found RmtC in 3 isolates, 1 of which also harbored RmtF. The isolates were clonally unrelated; rmtC and rmtF genes were located on a chromosome with the blaNDM-1 gene. Strategies are needed to limit the spread of such isolates.
Pseudomonas aeruginosa causes severe and chronic invasive infections in critically ill patients. Aminoglycosides are used either alone or in combination with β-lactams as effective agents for treating such infections (1). Aminoglycosides block protein synthesis by binding to bacterial 16S rRNA of the 30S ribosomal subunit. Methylation of 16S rRNA makes bacteria highly resistant to aminoglycosides (2). Increasing instances are reported of 16S rRNA methyltransferase (16S RMTase)–producing, Gram-negative bacteria that confer high levels of resistance to aminoglycosides. Eleven types of 16S RMTases (ArmA, RmtA–RmtH, and NpmA) have so far been reported in several nosocomially transmitted pathogens, including P. aeruginosa (2–5). Recently, 16S RMTases have been reported in association with the New Delhi metallo-β lactamase-1 (NDM-1) in Enterobacteriaceae (3). However, such association has not been reported in P. aeruginosa. Therefore, we investigated the presence of 16S RMTases in NDM-1–positive P. aeruginosa isolates recovered from different clinical specimens.
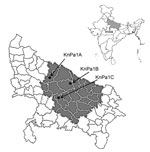
A total of 130 consecutive P. aeruginosa isolates recovered from different clinical specimens at Sanjay Gandhi Postgraduate Institute of Medical Sciences in Lucknow, Uttar Pradesh, India, during November 2013–April 2014 were included in the study; all specimens were collected from within the state (Figure 1). P. aeruginosa isolates were identified by standard microbiological techniques (6) and further confirmed by Phoenix automated identification and sensitivity systems (BD Biosciences, San Jose, CA, USA). The drug susceptibility profile was interpreted by using Clinical and Laboratory Standards Institute breakpoints (7). A total of 38 (29.23%) isolates were resistant to meropenem and imipenem. These isolates were subjected to PCR by using blaNDM–specific primers (8) followed by amplicon sequencing. Sequencing identified blaNDM-1 in all 38 isolates. The isolates were further screened for high-level aminoglycoside resistance by their ability to grow on Muller Hinton agar containing amikacin and gentamicin 256 mg/L each as a marker for 16S RMTase (3). A total of 33 (86.84%) isolates were positive for high-level aminoglycoside resistance. Geographic locations of patients infected with these isolates are provided in Figure 1. Each of these isolates were further subjected to PCR for detection of 16S RMTases (ArmA and RmtA–RmtH) by using primers and conditions described previously (2–5); 17 (51%) isolates were positive for 16S RMTases. Their distributions were as follows: ArmA in 6 (18%); RmtB in 4 (12%); ArmA + RmtB in 4 (12%); RmtC in 2 (KnPa1A and KnPa1C) (6%); and RmtC + RmtF in 1 (KnPa1B) (3%). RmtC and RmtF had not previously been reported in P. aeruginosa. KnPa1A, KnPa1B, and KnPa1C were further characterized; sequence analysis of amplicons confirmed rmtC with 100% nucleotide identity originally described in Proteus mirabilis strain ARS68 from Japan (9) and assigned EMBL/GenBank nucleotide accession nos. KJ476816, KJ476817, and KJ476818. MICs of the 3 isolates for different aminoglycosides, β-lactams, β-lactam/β-lactamase inhibitor combinations, carbapenems, and colistin are provided in the Table.
KnPa1A was isolated from surgical drainage from a woman, 59 years of age, who had hypertension and underwent an abdominal hysterectomy for cervical carcinoma, followed by external beam radiotherapy. Eventually, a vesicovaginal fistula developed, and pelvic fluid was collected. P. aeruginosa (KnPa1A) was isolated from pelvic drainage. Her condition stabilized, and she was discharged with advice for repair of the fistula, but she did not return for further treatment. During hospitalization, she received multiple antimicrobial drugs.
KnPa1B was isolated from endoscopic nasobiliary drainage (ENBD) collected from a man, 57 years of age, who had extrahepatic biliary obstruction as a complication of hilar cholangiocarcinoma. Endoscopic retrograde cholangiopancreatography with stenting was performed. Stent block and fever occurred, necessitating a repeat of the procedure and drainage of fluid. He was discharged with the drainage tube in situ and was advised to return for surgery. P. aeruginosa (KnPa1B) was isolated from ENBD. The patient did not report for follow-up treatment.
KnPa1C was isolated from a man, 68 years of age, who had diabetes mellitus and hypertension. He had stricture of the urethra and meatal narrowing after having a transurethral prostate resection. He underwent urethral dilatations and placement of a urinary catheter. A urinary tract infection was diagnosed, and P. aeruginosa (KnPa1C) was recovered from urine. The patient received piperacillin/tazobactam and colistin combination therapy; urine culture was sterile on day 3 posttreatment.
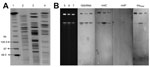
Resistance genes such as metallo-β-lactamases (e.g., IMP, VIM, SIM, GIM, SPM) and extended-spectrum β-lactamases such as TEM, SHV, CTX-M, and AmpC were detected by using PCR (8) (Table). To study genetic relatedness among the 3 isolates, genomic DNA in agarose blocks was separated on 1.0% agarose gels in 0.5 × tris-borate-EDTA buffer with the CHEF II D-Mapper XA pulsed-field gel electrophoresis system (Bio-Rad, Hercules, CA, USA) following standard conditions (10). All 3 isolates had different PFGE patterns (Figure 2). Multilocus sequence typing (MLST) was done according to protocols described in the Pseudomonas aeruginosa MLST Database (http://pubmlst.org/paeruginosa). Seven chromosomal genes were PCR amplified and sequenced; the sequences were compared with those on the MLST database to determine allele numbers and sequence types (STs). KnPa1A, KnPa1B, and KnPa1C belonged to ST764, ST902, and ST880, respectively.
ISEcp1 was previously shown to promote both expression and transposition of rmtC (11); hence, to assess association of ISEcp1 with rmtC, PCR was performed on the 3 isolates with primer pairs ISEcpIR-F and rmtC-down and ISEcp1–5′ and RMTC-R, as described (12). Sequence analysis of amplicons revealed association of an intact ISEcp1 element with rmtC in KnPa1A; however, complete ISEcp1 could not be amplified in KnPa1B and KnPa1C, corroborating earlier observations of either partial deletion of this element or role of a different ISEcp1-like element in the spread of rmtC in gram-negative bacteria (13). Attempts to transfer rmtC from all 3 isolates to rifampin-resistant Escherichia coli 20R764 and ciprofloxacin-resistant P. aeruginosa of strain 105 through conjugation were unsuccessful (1,13). Repeated attempts to obtain amikacin-resistant (MIC ≥16 g/mL) transformants of E. coli DH5α and P. aeruginosa PA01 by electroporation with plasmid preparation by using the Kado and Liu method (14) were also unsuccessful, despite successful transfer of control plasmids. To determine the location of rmtC, rmtF, and blaNDM-1, genomic DNA from the 3 isolates was digested separately with restriction enzyme I-Ceu-1 (New England Biolabs, Beverly, MA, USA), separated by PFGE, and subsequently assayed with 16S rRNA, rmtC, rmtF, and blaNDM-1 probes (13). All these probes were hybridized with chromosomal DNA (Figure 2) and not with plasmid extract. This result shows that rmtC, rmtF, and blaNDM-1 were located and stabilized on the chromosome of P. aeruginosa.
We describe an occurrence of 16S RMTases RmtC and RmtF in clinical isolates of P. aeruginosa co-producing blaNDM-1. The rmtC and rmtF genes might have been acquired from plasmids as part of mobile genetic elements and finally integrated and stabilized on the chromosome, but the underlying mechanism of transmission needs to be elucidated. Further, spread of multidrug-resistant P. aeruginosa strains expressing RmtC with and without an intact ISEcp1 element and NDM-1 is of major clinical concern and calls for further studies to limit the spread of such strains.
Mr. Rahman is a PhD scholar in Department of Microbiology, Sanjay Gandhi Postgraduate Institute of Medical Sciences, Lucknow, India. His research interests are characterization of NDM and 16S RMTases in gram-negative bacteria.
Acknowledgment
This study was supported by Indo-Spanish research grant (DST/INT/SPAIN/P-28/2011 C), Department of Science and Technology, New Delhi, India, and the Spanish Ministry of Science and Innovation (PRI-PIBIN-2011–0915).
References
- Yokoyama K, Do Y, Yamane K, Kurokawa H, Shibata N, Shibayama K, Acquisition of 16S rRNA methylase gene in Pseudomonas aeruginosa. Lancet. 2003;362:1888–93. DOIPubMedGoogle Scholar
- Wachino J, Arakawa Y. Exogenously acquired 16S rRNA methyltransferases found in aminoglycoside-resistant pathogenic Gram-negative bacteria: an update. Drug Updat 2012;15:133–48.
- Hidalgo L, Hopkins KL, Gutierrez B, Ovejero CM, Shukla S, Douthwaite S, Association of the novel aminoglycoside resistance determinant RmtF with NDM carbapenemase in Enterobacteriaceae isolated in India and the UK. J Antimicrob Chemother. 2013;68:1543–50. DOIPubMedGoogle Scholar
- Bueno MF, Francisco GR, O’Hara JA, de Oliveira Garcia D, Doi Y. Coproduction of 16S rRNA methyltransferase RmtD or RmtG with KPC-2 and CTX-M group extended-spectrum β-lactamases in Klebsiella pneumoniae. Antimicrob Agents Chemother. 2013;57:2397–400. DOIPubMedGoogle Scholar
- O’Hara JA, McGann P, Snesrud EC, Clifford RJ, Waterman PE, Lesho EP, Novel 16S rRNA methyltransferase RmtH produced by Klebsiella pneumoniae associated with war-related trauma. Antimicrob Agents Chemother. 2013;57:2413–6. DOIPubMedGoogle Scholar
- Collee JG, Miles RS, Wan B. Tests for the identification of bacteria. In: Collee JG, Fraser AG, Marmion BP, Simmons A, editors. Mackie and McCartney practical medical microbiology. 14th ed. Edinburgh (Scotland): Churchill Livingstone; 1996. p. 131–50.
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Twenty-fourth informational supplement. M100-S24. Wayne (PA): The Institute; 2014.
- Rahman M, Shukla SK, Prasad KN, Ovejero CM, Pati BK, Tripathi A, Prevalence and molecular characterization of New Delhi metallo-β-lactamases NDM-1, NDM-5, NDM-6 and NDM-7 in multidrug resistant Enterobacteriaceae from India. Int J Antimicrob Agents. 2014;44:30–7. DOIPubMedGoogle Scholar
- Wachino J, Yamane K, Shibayama K, Kurokawa H, Shibata N, Suzuki S, Novel plasmid-mediated 16S rRNA methylase, RmtC, found in a Proteus mirabilis isolate demonstrating extraordinary high-level resistance against various aminoglycosides. Antimicrob Agents Chemother. 2006;50:178–84. DOIPubMedGoogle Scholar
- Grundmann H, Schneider C, Hartung D, Daschner FD, Pitt TL. Discriminatory power of three DNA-based typing techniques for Pseudomonas aeruginosa. J Clin Microbiol. 1995;33:528–34 .PubMedGoogle Scholar
- Wachino J, Yamane K, Kimura K, Shibata N, Suzuki S, Ike Y, Mode of transposition and expression of 16S rRNA methyltransferases gene rmtC accompanied by ISEcp1. Antimicrob Agents Chemother. 2006;50:3212–5. DOIPubMedGoogle Scholar
- Zong Z, Partridge S, Iredell J. RmtC 16S rRNA methyltransferases in Australia. Antimicrob Agents Chemother. 2008;52:794–5. DOIPubMedGoogle Scholar
- Hopkins KL, Escudero JA, Hidalgo L, Zorn BG. 16S rRNA methyltransferases RmtC in Salmonella enterica serovar Virchow. Emerg Infect Dis. 2010;16:712–5. DOIPubMedGoogle Scholar
- Kado CI, Liu ST. Rapid procedure for detection and isolation of large and small plasmids. J Bacteriol. 1981;145:1365–73 .PubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 21, Number 11—November 2015
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Kashi Nath Prasad, Department of Microbiology, Sanjay Gandhi Postgraduate Institute of Medical Sciences, Lucknow-226014, India
Top