Volume 21, Number 3—March 2015
Dispatch
Spillover of Mycobacterium bovis from Wildlife to Livestock, South Africa
Cite This Article
Citation for Media
Abstract
During August 2012–February 2013, bovine tuberculosis was detected in communal livestock bordering the Greater Kruger National Park Complex (GKNPC) in South Africa. Using spacer oligonucleotide and variable number tandem repeat typing, we identified the Mycobacterium bovis strain endemic in GKNPC wildlife. Our findings indicate bovine tuberculosis spillover from GKNPC wildlife to neighboring livestock.
Bovine tuberculosis is an infectious disease caused by Mycobacterium bovis. The wide host range of the pathogen comprises humans and domestic and wild animals. Great strides in controlling bovine tuberculosis have drastically reduced its prevalence in livestock and humans, particularly in industrialized countries. However, in developing countries in southern Africa and elsewhere, bovine tuberculosis remains a challenge to animal health because of a total or partial lack of bovine tuberculosis control, limited by a lack of funds (1,2). The control and/or elimination of bovine tuberculosis in both developing and industrialized countries can be complicated by wildlife reservoirs of the disease, which pose a threat of re-infection in livestock (3). In sub-Saharan Africa, particularly South Africa and Uganda, African buffalos (Syncerus caffer) serve as wildlife reservoirs of bovine tuberculosis; in Zambia, lechwe antelopes (Kobus leche Kafuensis) have been identified as wildlife reservoirs (4,5). New reports have suggested greater kudu (Tragelaphus strepsiceros) and common warthog (Phacochoerus africanus) as potential wildlife reservoirs of bovine tuberculosis (4).
M. bovis is endemic in buffaloes and has spilled into other wildlife species, particularly in the Kruger National Park (KNP) and adjacent game reserves that form part of the Greater Kruger National Park Complex (GKNPC) in South Africa (6,7). Except for data from sporadic regulatory bovine tuberculosis surveillance activities in cattle adjacent to the GKNPC, no data exist on the transmission of bovine tuberculosis from the GKNPC, where it is endemic, into livestock in neighboring communities (3). Because of the potentially negative implications of livestock–wildlife interactions on livestock and human health, the presence and role of zoonotic diseases in these communities needs to be investigated (5). We report on an investigation into the status and genotype of bovine tuberculosis in livestock in rural communities bordering the bovine tuberculosis–endemic GKNPC.
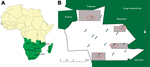
The study was conducted in a rural community under the Mnisi Tribal Authority. The community is situated in Mpumalanga Province, South Africa, and borders the GKNPC in the west and 1 private game reserve (Figure 1). We constructed maps for this study using ArcGIS version 10.2 (http://www.arcgis.com). The KNP and private game reserves are fenced and have buffer zones established by double fencing (8).
Cattle farming, an essential part of the livelihood of the Mnisi community, is practiced primarily on a communal basis. Approximately 12,000 cattle live within the Mnisi area with 15 dip tanks, where the national government provides veterinary extension services and which represent epidemiologic units for sampling (Figure 1). Farmers in the study area are assigned and registered to a particular dip tank by the agricultural authorities by a stock card system. On a stock card, number of cattle, births/deaths, and animal movement are recorded. For this study, a dip tank is considered a whole herd because animals in a particular dip tank interacted extensive during grazing and dip tank inspections, and usually 1 herdsman was in charge of multiple stock cards.
During August 2012–February 2013, a total of 1,166 cattle at the 15 dip tanks in the study area were tested for bovine tuberculosis by using the comparative intradermal tuberculin test (CIDT). Animals selected for testing were chosen randomly from a list of stock cards at each dip tank. From each stock card chosen, we selected 10% of registered animals; however, a minimum of 2–3 animals per stock card were tested. All stock card owners willing to participate were included to reach a target of 10% of cattle assigned to each dip tank. We calculated the bovine tuberculosis status and 95% CI per dip tank assuming a binomial distribution of the data (Table 1).
A whole-blood interferon-γ (IFN-γ) assay was performed as an ancillary test to the CIDT on all 4 CIDT-positive cattle and all 5 cattle with inconclusive reactions (defined as a difference between the bovine and avian increase in skin-fold thickness of >3 mm) (Table 2). Among the 9 cattle, 4 animals were classified as bovine tuberculosis reactors on the basis of the IFN-γ assay response (9), 1 each in 4 of the 15 dip tanks (Figure 1).
Animals classified as bovine tuberculosis reactors were purchased and slaughtered. These animals included a 1-month-old calf born to a CIDT- and IFN-γ assay–positive cow (animal no. K1). Standard sets of tissue samples were collected and cultured as previously described (10). Pathologic examination and culture results are shown in Table 2.
M. bovis was isolated from the 5 animals, and the isolates were characterized by using spacer oligonucleotide typing (spoligotyping) (11). Spoligotypes were named according to the M. bovis spoligotype database (http://www.mbovis.org). Variable number tandem repeat (VNTR) typing of the isolates was performed as previously described (12). Spoligotyping showed a single M. bovis spoligotype, SB 0121, in all isolates. NTR analysis using a 13-loci panel identified all isolates as the KNP VNTR 1, which constitutes the M. bovis outbreak strain responsible for the bovine tuberculosis epidemic in the KNP and the larger GKNPC (12) (Figure 2).
We detected bovine tuberculosis in livestock directly bordering the GKNPC ecosystem. All 5 animals examined were infected with the same spoligotype and VNTR genotype of M. bovis as wildlife species in the adjacent GKNPC (12). This finding strongly suggests the spillover of M. bovis infection from wildlife to neighboring cattle (Figure 2). Alternatively, the KNP outbreak strain could have persisted in the area from which it entered the wildlife population of the GKNPC (6) and subsequently could have spread outside the KNP and reached the study area, a distance of ≈180 km. However, during 1996–2012, provincial State Veterinary Services of Mpumalanga tested a total of 96,806 head of cattle in this region of interest, which comprises the veterinary districts of Bushbuckridge (where the study area is located), Nsikazi (bordering the GKNPC in the west), and Nkomazi (south of KNP) using the CIDT as part of its regular bovine tuberculosis surveillance (B.J.A. du Plessis, unpub. data). No bovine tuberculosis reactors were detected in Bushbuckridge or in Nsikazi districts. In 3 unrelated outbreaks during 2009, 2010, and 2011 in the Nkomazi district, 1–3 bovine reactor animals were detected (B.J.A. du Plessis, unpub. data.). All outbreak strains were genotyped, and their spoligotypes and VNTR profiles differed from each other and from the M. bovis strain endemic to the GKNPC (results not shown). This information supports the hypothesis that bovine tuberculosis–infected cattle in our current study contracted M. bovis from neighboring wildlife in the GKNPC.
In conclusion, our study provides evidence that infected wildlife in the GKNPC constitute a risk factor for bovine tuberculosis infection of neighboring cattle, despite the separation of livestock and wildlife by a well-maintained disease control fence. These findings are of great concern, not only to livestock health and production in communities bordering the GKNPC but also to public health and to human livelihoods because of the zoonotic potential of bovine tuberculosis.
Ms. Musoke is a microbiologist studying toward a PhD at the University of Pretoria, Department of Veterinary Tropical Diseases, in South Africa. Her research interests include transboundary disease epidemiology and the effect of diseases on the livelihoods of rural communities.
Acknowledgments
We thank the Mpumalanga Veterinary Services, particularly Bjorn Reininghaus, Gypsey Mathumba, Jerry Ndlozi, and Solly Mokoni, for logistical arrangements with the skin test and communicating with farmers. We thank the Hluvukani Animal Health Clinic staff for their assistance in the field. We thank Anne Conan for her assistance in constructing the maps.
The Institute of Tropical Medicine funded this project through the framework agreement with the Belgian Development Cooperation.
References
- Michel AL, Muller B, van Helden PD. Mycobacterium bovis at the animal–human interface: a problem, or not? Vet Microbiol. 2010;140:371–81 . DOIPubMedGoogle Scholar
- Grange JM. Mycobacterium bovis infection in human beings. Tuberculosis (Edinb). 2001;81:71–7. DOIPubMedGoogle Scholar
- De Garine-Wichatitsky M, Caron A, Kock R, Tschopp R, Munyeme M, Hofmeyr M, A review of bovine tuberculosis at the wildlife–livestock–human interface in sub-Saharan Africa. Epidemiol Infect. 2013;141:1342–56. DOIPubMedGoogle Scholar
- Michel AL, Bengis RG, Keet DF, Hofmeyr M, Klerk LM, Cross PC, Wildlife tuberculosis in South African conservation areas: implications and challenges. Vet Microbiol. 2006;112:91–100. DOIPubMedGoogle Scholar
- Munyeme M, Muma JB, Samui KL, Skjerve E, Nambota AM, Phiri IG, Prevalence of bovine tuberculosis and animal level risk factors for indigenous cattle under different grazing strategies in the livestock/wildlife interface areas of Zambia. Trop Anim Health Prod. 2009;41:345–52. DOIPubMedGoogle Scholar
- Michel AL, Coetzee ML, Keet DF, Mare L, Warren R, Cooper D, Molecular epidemiology of Mycobacterium bovis isolates from free-ranging wildlife in South African game reserves. Vet Microbiol. 2009;133:335–43. DOIPubMedGoogle Scholar
- Kukumi S. Africa walking safaris [cited 2013 Dec 10]. http://www.africawalkingsafaris.co.za/Greater%20Kruger%20National%20Park.html
- Brahmbhatt DP, Fosgate GT, Dyason E, Budke CM, Gummow B, Jori F, Contacts between domestic livestock and wildlife at the Kruger National Park Interface of the Republic of South Africa. Prev Vet Med. 2012;103:16–21 . DOIPubMedGoogle Scholar
- Michel AL, Cooper D, Jooste J, de Klerk LM, Jolles A. Approaches towards optimising the gamma interferon assay for diagnosing Mycobacterium bovis infection in African buffalo (Syncerus caffer). Prev Vet Med. 2011;98:142–51 . DOIPubMedGoogle Scholar
- Hlokwe TM, van Helden P, Michel AL. Evidence of increasing intra and inter-species transmission of Mycobacterium bovis in South Africa: are we losing the battle? Prev Vet Med. 2014;115:10–7. DOIPubMedGoogle Scholar
- Kamerbeek J, Schouls L, Kolk A, van Agterveld M, van Soolingen D, Kuijper S, Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol. 1997;35:907–14 .PubMedGoogle Scholar
- Hlokwe TM, van Helden P, Michel A. Evaluation of the discriminatory power of variable number of tandem repeat typing of Mycobacterium bovis isolates from southern Africa. Transbound Emerg Dis. 2013;60(Suppl 1):111–20. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 21, Number 3—March 2015
| EID Search Options |
|---|
|
|
|
|
|
|
![Thumbnail of Dendogram depicting the genetic homology between isolates obtained in study of bovine tuberculosis transmission in the Greater Kruger National Park Complex during August 2012–February 2013 and from other outbreaks in South Africa. Colors differentiate the isolates. EC, Eastern Cape; FS, Free State; HiP, Hluhluwe iMofolozi Game Reserve; KNP, Kruger National Park (current study area [Mnisi]); KZN, Kwa-Zulu Natal; LP, Limpopo; MP, Mpumalanga; VNTR, variable number tandem repeat typing.](/eid/images/13-1690-F2-tn.jpg)
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jolly Musoke, University of Pretoria, Private Bag X04, Old Soutpan Rd, Soutpan M35, Onderstepoort, South Africa
Top