Volume 21, Number 4—April 2015
Letter
Nairobi Sheep Disease Virus RNA in Ixodid Ticks, China, 2013
Figure
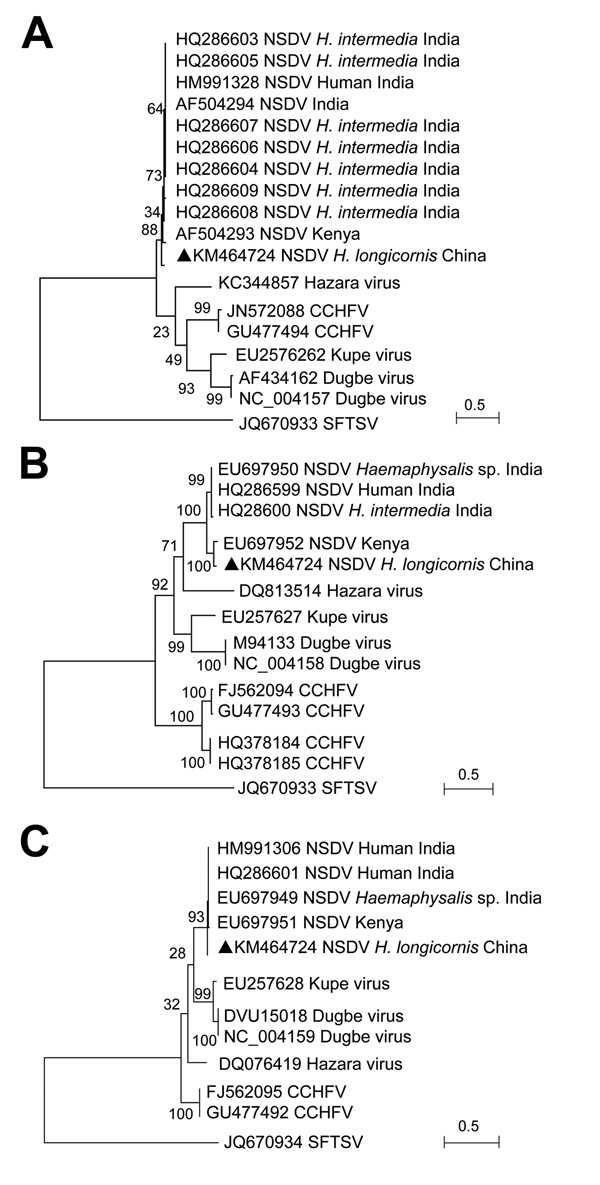
Figure. Phylogenetic analysis of Nairobi sheep disease virus (China) and other nairoviruses. The phylogenetic trees were generated in MEGA5.2 software (http://www.megasoftware.net). The complete coding regions for nucleocapsid protein in the small segment (A), glycoprotein precursor in the medium segment (B), and RNA dependent RNA polymerase in the large segment (C) were analyzed by the maximum-likelihood method. An emergent severe fever thrombocytopenia syndrome virus (SFTSV; family Bunyaviridae, genus Phlebovirus) was used as the outgroup. Bootstrap testing (1,000 replicates) was performed, and the bootstrap values are indicated. Sequences are identified by their GenBank accession numbers, followed by the virus name, host, and country. Black triangles indicate novel strain NSDV (China). Scale bars indicate substitutions per site. CCHFV, Crimean-Congo hemorrhagic fever virus.
1These authors contributed equally to this article.