Volume 21, Number 4—April 2015
Letter
Avian Influenza A(H10N7) Virus–Associated Mass Deaths among Harbor Seals
Cite This Article
Citation for Media
To the Editor: Avian influenza A viruses occasionally cross the species barrier; influenza A(H5N1) virus and the recently emerged influenza A(H7N9) virus are prime examples of bird-to-human transmission (1,2). In addition, avian influenza A viruses can cross to various other mammalian species, including pinnipeds (e.g., seals) (3,4).
Recently, mass deaths have occurred among harbor seals (Phoca vitulina); hundreds of carcasses washed up the shores of Sweden (March 2014), Denmark (July 2014), and Germany (October 2014). Approximately 1,400 dead harbor seals were seen in the coastal waters of Schleswig-Holstein in Germany alone, where the population is ≈12,000 animals.
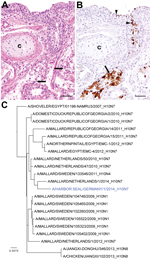
We report the investigation of the deaths of 17 seals from different age groups that were found dead on the islands of Helgoland and Sylt, Germany, during the second week of October 2014. Complete necropsies were performed on the carcasses, which were in variable nutritional conditions, ranging from very poor to good. Necropsy results showed consistently poorly retracted lungs with severe congestion, occasional diffuse consolidation, and multifocal firm nodular areas of gray-yellow discoloration with varying numbers of metazoic parasites. Histologic examinations showed acute necrotizing bronchitis and adenitis of bronchial glands with sloughing of epithelial cells (Figure, panel A). Occasionally, mild interstitial pneumonia was found. Multifocal pyogranulomatous to necrotizing pneumonia was associated with parasite infestation. A few animals had suppurative to necrotizing or nonsuppurative rhinitis and tracheitis.
Because mass deaths among seals were caused by phocine distemper virus in the same area in 1988 and 2002, we tested lung and throat swab samples for morbillivirus using reverse transcription PCR (RT-PCR) and immunohistochemical analysis (5). In addition, real-time RT-PCR targeting the influenza A virus matrix gene was performed (6). No indications for the morbillivirus were detected by RT-PCR and immunohistochemistry; however, in lung lesions and throat swab samples of 11 animals, a positive signal was observed by the influenza A matrix gene real-time RT-PCR (cycle threshold values 15.0–33.9). Influenza A virus (A/harbor seal/Germany/1/2014) was subsequently isolated from lung and throat swab samples; the virus replicated to high titers in 11-day-old embryonated chicken eggs and on MDCK cells. By PCR using specific primers and subsequent Sanger sequencing of the hemagglutinin and neuraminidase genes, this virus was characterized as an influenza A virus of the H10N7 subtype, commonly found in migratory waterfowl (6). In addition, genetic analyses of all other gene segments indicated that the influenza virus A/harbor seal/Germany/1/2014 was most closely related to various influenza A viruses detected in wild birds. Specifically, the hemagglutinin and neuraminidase genes were genetically most closely related to subtype H10N7 viruses recently found in migratory ducks in Georgia, Egypt, and the Netherlands (Figure, panel C) (7). Genetic analyses were based on BLAST analyses using public databases available as of October 17, 2014 (http://www.ncbi.nlm.nih.gov, http://www.gisaid.com) and supplemented with H10 and N7 sequences from the international wild bird surveillance program of Erasmus Medical Center (Technical Appendix Table). A maximum-likelihood phylogenetic tree of the hemagglutinin gene was generated by using PhyML version 3.1 (8) with the general time reversible +I+Γ model of nucleotide substitution; a full heuristic search and subtree pruning and regrafting searches were performed. The tree was visualized by using Figtree version 1.4.0 (http://tree.bio.ed.ac.uk/software/figtree).
To further elucidate the role of influenza A(H10N7) virus in the pathogenesis of the disease causing deaths among the seals, we conducted immunohistochemical analysis on the lungs using an influenza A virus nucleoprotein-specific monoclonal antibody (9). Evaluation of the lung tissues of the dead seals showed viral antigen in cytoplasm and nuclei of epithelial cells of bronchi and bronchial glands of affected lung areas (Figure, panel B), which suggests that this virus played a major role in the deaths. Immunohistochemical analysis performed on various organs (including brain and olfactory bulb) indicated that viral antigen was restricted to the respiratory tract.
Although avian influenza A virus infections previously have caused mass deaths in seals (3,4,10), subtype H10N7 has not been associated with such events. We can speculate that the ongoing deaths could eventually affect all harbor seal populations of northwestern Europe and have consequences for wildlife management and seal rehabilitation activities. In addition, preliminary analysis of the hemagglutinin sequence of the influenza A(H10N7) virus suggests the presence of molecular determinants that indicate mammalian adaptation. Various analyses are ongoing to answer questions about the route of transmission among seals and possible transmissibility to humans.
Note added in proof: Zohari et al. also recently reported the involvement of avian influenza A(H10N7) virus in mass deaths of harbor seals in Sweden (Euro Surveill. 2014;19:pii: 20967).
Acknowledgment
This work was supported in part by a grant from the Niedersachsen-Research Network on Neuroinfectiology of the Ministry of Science and Culture of Lower Saxony, Germany, and the European Research Council project FLUPLAN (2503136).
References
- Claas EC, Osterhaus AD, van Beek R, De Jong JC, Rimmelzwaan GF, Senne DA, Human influenza A H5N1 virus related to a highly pathogenic avian influenza virus. Lancet. 1998;351:472–7 and. DOIPubMedGoogle Scholar
- Gao R, Cao B, Hu Y, Feng Z, Wang D, Hu W, Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med. 2013;368:1888–97 and. DOIPubMedGoogle Scholar
- Reperant LA, Rimmelzwaan GF, Kuiken T. Avian influenza viruses in mammals. Rev Sci Tech. 2009;28:137–59 .PubMedGoogle Scholar
- Anthony SJ, St Leger JA, Pugliares K, Ip HS, Chan JM, Carpenter ZW, Emergence of fatal avian influenza in New England harbor seals. MBio. 2012;3:e00166–12 . DOIPubMedGoogle Scholar
- Jensen T, van de Bildt M, Dietz HH, Andersen TH, Hammer AS, Kuiken T, Another phocine distemper outbreak in Europe. Science. 2002;297:209 and. DOIPubMedGoogle Scholar
- Munster VJ, Baas C, Lexmond P, Waldenstrom J, Wallensten A, Fransson T, Spatial, temporal, and species variation in prevalence of influenza A viruses in wild migratory birds. PLoS Pathog. 2007;3:e61 and. DOIPubMedGoogle Scholar
- Lewis NS, Javakhishvili Z, Russell CA, Machablishvili A, Lexmond P, Verhagen JH, Avian influenza virus surveillance in wild birds in Georgia: 2009–2011. PLoS ONE. 2013;8:e58534 . DOIPubMedGoogle Scholar
- Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21 and. DOIPubMedGoogle Scholar
- de Boer GF, Back W, Osterhaus AD. An ELISA for detection of antibodies against influenza A nucleoprotein in humans and various animal species. Arch Virol. 1990;115:47–61 and. DOIPubMedGoogle Scholar
- Hinshaw VS, Bean WJ, Webster RG, Rehg JE, Fiorelli P, Early G, Are seals frequently infected with avian influenza viruses? J Virol. 1984;51:863–5 .PubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 21, Number 4—April 2015
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Albert D.M.E. Osterhaus, University of Veterinary Medicine—Department of Pathology, Buenteweg 17, Lower Saxony, Hannover D-30559, Germany
Top