Volume 22, Number 8—August 2016
Dispatch
Multilocus Sequence Typing Tool for Cyclospora cayetanensis
Figure 2
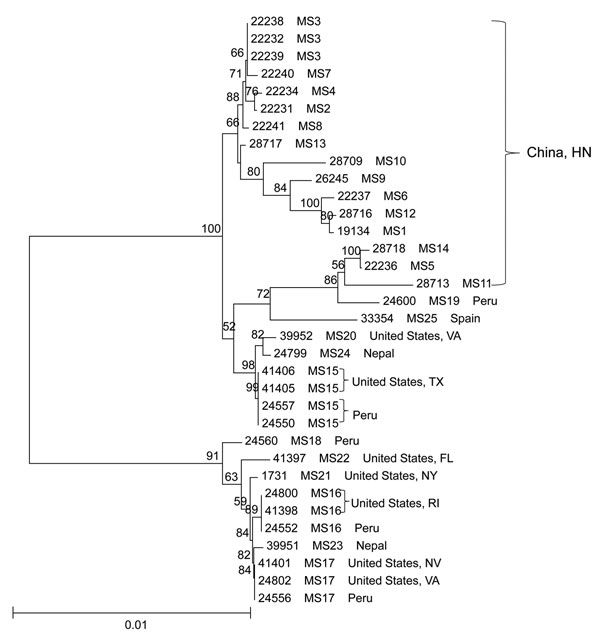
Figure 2. Phylogenetic relationships among concatenated multilocus sequence types of Cyclospora cayetanensis as assessed by a neighbor-joining analysis of the nucleotide sequences, using genetic distances calculated by the Kimura 2-parameter model. Numbers on branches are bootstrap values from 1,000 replicate analyses. Only values >50% are displayed on the left of each node. Scale bar indicates substitution rates per nucleotide. HN, Henan.
Page created: July 15, 2016
Page updated: July 15, 2016
Page reviewed: July 15, 2016
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.