Volume 23, Number 10—October 2017
Dispatch
Diagnosis of Fatal Human Case of St. Louis Encephalitis Virus Infection by Metagenomic Sequencing, California, 2016
Figure
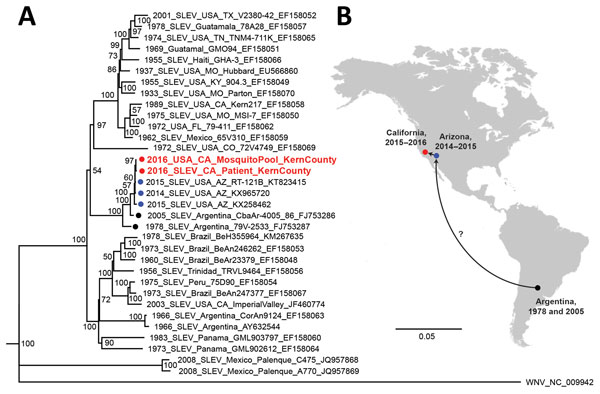
Figure. Phylogeny and spread of St. Louis encephalitis virus. A) Multiple sequence alignment of 32 complete SLEV genomes from GenBank and the 2 SLEV genomes corresponding to the case-patient’s strain and a strain from a mosquito collected in June 2016 from Kern County, California (red circles and text). Alignment was performed using MAFFT (10), followed by tree generation using a neighbor-joining algorithm using Geneious (11). The cluster containing the 2014–2016 California and Arizona SLEV genome, including those from the case-patient and 2016 mosquito pool, is rooted by SLEV strains sequenced from mosquitoes collected in Argentina in 1978 and 2005 (black circles). Isolates are named by location, year of collection, strain name, and GenBank accession number. Bootstrap support values are given for each node. Scale bar indicates nucleotide substitutions per site. B) Geographic spread of SLEV in the Americas, from Argentina in 2005 to California and Arizona during 2014–2016. Because genome sequences from US states reporting SLEV activity are not publicly available and surveillance for SLEV in South and Central America is not routinely performed, the pathway or pathways by which the virus came to the southwestern United States remain unclear (question mark). SLEV, St. Louis encephalitis virus.
References
- Salimi H, Cain MD, Klein RS. Encephalitic arboviruses: emergence, clinical presentation, and neuropathogenesis. Neurotherapeutics. 2016;13:514–34. DOIPubMedGoogle Scholar
- CDC. Saint Louis encephalitis [cited 2016 Dec 8]. http://www.cdc.gov/sle
- Venkat H, Krow-Lucal E, Hennessey M, Jones J, Adams L, Fischer M, et al. Concurrent outbreaks of St. Louis encephalitis virus and West Nile virus disease—Arizona, 2015. MMWR Morb Mortal Wkly Rep. 2015;64:1349–50.PubMedGoogle Scholar
- White GS, Symmes K, Sun P, Fang Y, Garcia S, Steiner C, et al. Reemergence of St. Louis encephalitis virus, California, 2015. Emerg Infect Dis. 2016;22:2185–8. DOIPubMedGoogle Scholar
- Mongkolrattanothai K, Naccache SN, Bender JM, Samayoa E, Pham E, Yu G, et al. Neurobrucellosis: unexpected answer from metagenomic next-generation sequencing. J Pediatric Infect Dis Soc. 2017;•••:piw066; Epub ahead of print. DOIPubMedGoogle Scholar
- Wilson MR, Naccache SN, Samayoa E, Biagtan M, Bashir H, Yu G, et al. Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N Engl J Med. 2014;370:2408–17. DOIPubMedGoogle Scholar
- Naccache SN, Federman S, Veeraraghavan N, Zaharia M, Lee D, Samayoa E, et al. A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res. 2014;24:1180–92. DOIPubMedGoogle Scholar
- Schlaberg R, Chiu CY, Miller S, Procop GW, Weinstock G; Professional Practice Committee and Committee on Laboratory Practices of the American Society for Microbiology; Microbiology Resource Committee of the College of American Pathologists. Microbiology Resource Committee of the College of American Pathologists. Validation of metagenomic next-generation sequencing tests for universal pathogen detection. Arch Pathol Lab Med. 2017;141:776–86. DOIPubMedGoogle Scholar
- Spinsanti LI, Díaz LA, Glatstein N, Arselán S, Morales MA, Farías AA, et al. Human outbreak of St. Louis encephalitis detected in Argentina, 2005. J Clin Virol. 2008;42:27–33. DOIPubMedGoogle Scholar
- Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80. DOIPubMedGoogle Scholar
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9. DOIPubMedGoogle Scholar
- Reisen WK, Lothrop HD, Wheeler SS, Kennsington M, Gutierrez A, Fang Y, et al. Persistent West Nile virus transmission and the apparent displacement St. Louis encephalitis virus in southeastern California, 2003-2006. J Med Entomol. 2008;45:494–508.PubMedGoogle Scholar
- Rahal JJ, Anderson J, Rosenberg C, Reagan T, Thompson LL. Effect of interferon-alpha2b therapy on St. Louis viral meningoencephalitis: clinical and laboratory results of a pilot study. J Infect Dis. 2004;190:1084–7. DOIPubMedGoogle Scholar