Volume 25, Number 6—June 2019
Research
Survey of Crimean-Congo Hemorrhagic Fever Enzootic Focus, Spain, 2011–2015
Figure 2
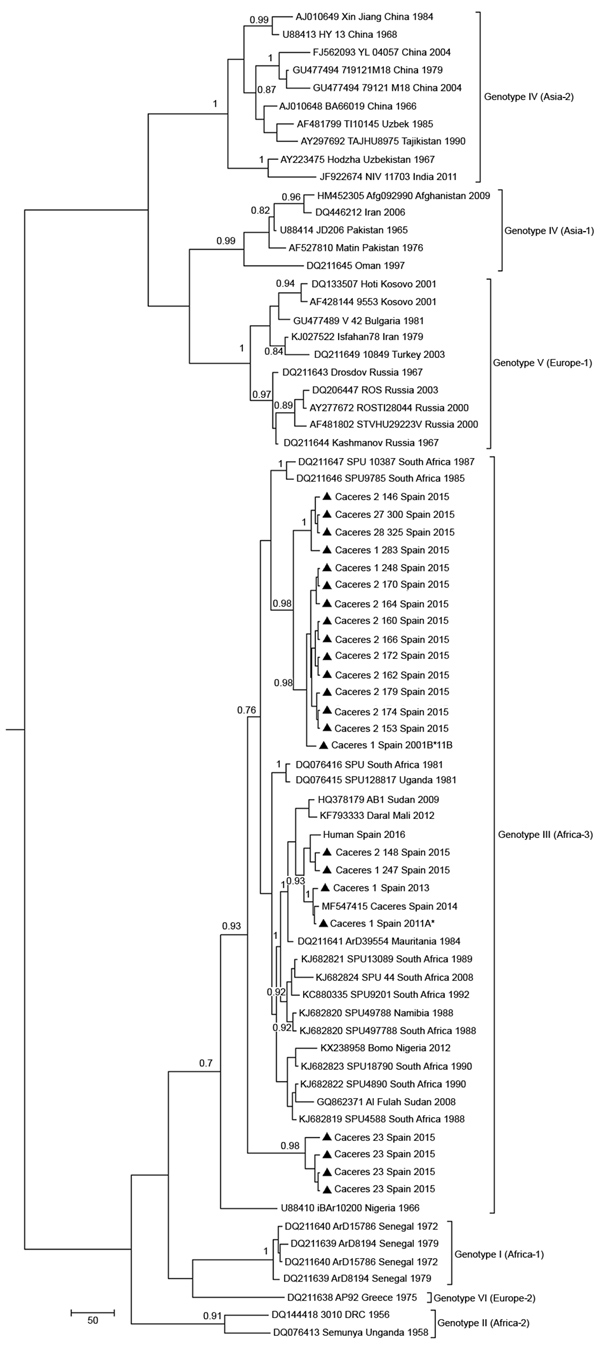
Figure 2. Maximum clade credibility genealogy based on partial (175-nt) sequences of the Crimean-Congo hemorrhagic fever virus small RNA segment of viruses from Spain and reference sequences. Numbers along branches indicate posterior probability values for the groups (values <0.70 are not shown). Triangles indicate newly sequenced strains from Spain reported in this study, which are identified by geographic origin, sampling site, and sampling year. Other sequences are indicated by GenBank accession number, strain, geographic origin, and sampling year. Sequences from this study indicated with an asterisk are included in European Molecular Biology Laboratory/GenBank databases. Genotypes are indicated in roman numerals and named according to Carroll et al. (30). Equivalent clade nomenclature is listed according to Chamberlain et al. (31) and indicated at right. Scale bar indicates nucleotide substitutions per site.
References
- World Health Organization. Crimean-Congo hemorrhagic fever. Fact sheet no. 208; 2013 [cited 2019 Mar 6]. http://www.who.int/mediacentre/factsheets/fs208/en
- Whitehouse CA. Crimean-Congo hemorrhagic fever. Antiviral Res. 2004;64:145–60. DOIPubMedGoogle Scholar
- Ergonul O. Crimean-Congo hemorrhagic fever virus: new outbreaks, new discoveries. Curr Opin Virol. 2012;2:215–20. DOIPubMedGoogle Scholar
- Estrada-Peña A, Palomar AM, Santibáñez P, Sánchez N, Habela MA, Portillo A, et al. Crimean-Congo hemorrhagic fever virus in ticks, Southwestern Europe, 2010. Emerg Infect Dis. 2012;18:179–80. DOIPubMedGoogle Scholar
- Negredo A, de la Calle-Prieto F, Palencia-Herrejón E, Mora-Rillo M, Astray-Mochales J, Sánchez-Seco MP, et al.; Crimean Congo Hemorrhagic Fever@Madrid Working Group. Autochthonous Crimean-Congo Hemorrhagic Fever in Spain. N Engl J Med. 2017;377:154–61. DOIPubMedGoogle Scholar
- Hoogstraal H. The epidemiology of tick-borne Crimean-Congo hemorrhagic fever in Asia, Europe, and Africa. J Med Entomol. 1979;15:307–417. DOIPubMedGoogle Scholar
- Gonzalez JP, Camicas JL, Cornet JP, Wilson ML. Biological and clinical responses of west African sheep to Crimean-Congo haemorrhagic fever virus experimental infection. Res Virol. 1998;149:445–55. DOIPubMedGoogle Scholar
- Spengler JR, Estrada-Peña A, Garrison AR, Schmaljohn C, Spiropoulou CF, Bergeron É, et al. A chronological review of experimental infection studies of the role of wild animals and livestock in the maintenance and transmission of Crimean-Congo hemorrhagic fever virus. Antiviral Res. 2016;135:31–47. DOIPubMedGoogle Scholar
- Wasfi F, Dowall S, Ghabbari T, Bosworth A, Chakroun M, Varghese A, et al. Sero-epidemiological survey of Crimean-Congo hemorrhagic fever virus in Tunisia. Parasite. 2016;23:10. DOIPubMedGoogle Scholar
- Gargili A, Estrada-Peña A, Spengler JR, Lukashev A, Nuttall PA, Bente DA. The role of ticks in the maintenance and transmission of Crimean-Congo hemorrhagic fever virus: A review of published field and laboratory studies. Antiviral Res. 2017;144:93–119. DOIPubMedGoogle Scholar
- Gunes T, Poyraz O, Vatansever Z. Crimean-Congo hemorrhagic fever virus in ticks collected from humans, livestock, and picnic sites in the hyperendemic region of Turkey. Vector Borne Zoonotic Dis. 2011;11:1411–6. DOIPubMedGoogle Scholar
- Sherifi K, Cadar D, Muji S, Robaj A, Ahmeti S, Jakupi X, et al. Crimean-Congo hemorrhagic fever virus clades V and VI (Europe 1 and 2) in ticks in Kosovo, 2012. PLoS Negl Trop Dis. 2014;8:
e3168 . DOIPubMedGoogle Scholar - Bente DA, Forrester NL, Watts DM, McAuley AJ, Whitehouse CA, Bray M. Crimean-Congo hemorrhagic fever: history, epidemiology, pathogenesis, clinical syndrome and genetic diversity. Antiviral Res. 2013;100:159–89. DOIPubMedGoogle Scholar
- Spengler JR, Estrada-Peña A. Host preferences support the prominent role of Hyalomma ticks in the ecology of Crimean-Congo hemorrhagic fever. PLoS Negl Trop Dis. 2018;12:
e0006248 . DOIPubMedGoogle Scholar - Papa A, Ma B, Kouidou S, Tang Q, Hang C, Antoniadis A. Genetic characterization of the M RNA segment of Crimean Congo hemorrhagic fever virus strains, China. Emerg Infect Dis. 2002;8:50–3. DOIPubMedGoogle Scholar
- Burt FJ, Swanepoel R. Molecular epidemiology of African and Asian Crimean-Congo haemorrhagic fever isolates. Epidemiol Infect. 2005;133:659–66. DOIPubMedGoogle Scholar
- Christova I, Di Caro A, Papa A, Castilletti C, Andonova L, Kalvatchev N, et al. Crimean-Congo hemorrhagic fever, southwestern Bulgaria. Emerg Infect Dis. 2009;15:983–5. DOIPubMedGoogle Scholar
- Maltezou HC, Maltezos E, Papa A. Contact tracing and serosurvey among healthcare workers exposed to Crimean-Congo haemorrhagic fever in Greece. Scand J Infect Dis. 2009;41:877–80. DOIPubMedGoogle Scholar
- Mishra AC, Mehta M, Mourya DT, Gandhi S. Crimean-Congo haemorrhagic fever in India. Lancet. 2011;378:372. DOIPubMedGoogle Scholar
- Aradaib IE, Erickson BR, Karsany MS, Khristova ML, Elageb RM, Mohamed ME, et al. Multiple Crimean-Congo hemorrhagic fever virus strains are associated with disease outbreaks in Sudan, 2008-2009. PLoS Negl Trop Dis. 2011;5:
e1159 . DOIPubMedGoogle Scholar - Mustafa ML, Ayazi E, Mohareb E, Yingst S, Zayed A, Rossi CA, et al. Crimean-Congo hemorrhagic fever, Afghanistan, 2009. Emerg Infect Dis. 2011;17:1940–1. DOIPubMedGoogle Scholar
- Nabeth P, Cheikh DO, Lo B, Faye O, Vall IO, Niang M, et al. Crimean-Congo hemorrhagic fever, Mauritania. Emerg Infect Dis. 2004;10:2143–9. DOIPubMedGoogle Scholar
- Papa A, Velo E, Papadimitriou E, Cahani G, Kota M, Bino S. Ecology of the Crimean-Congo hemorrhagic fever endemic area in Albania. Vector Borne Zoonotic Dis. 2009;9:713–6. DOIPubMedGoogle Scholar
- Ramírez de Arellano E, Hernández L, Goyanes MJ, Arsuaga M, Cruz AF, Negredo A, et al. Phylogenetic Characterization of Crimean-Congo Hemorrhagic Fever Virus, Spain. Emerg Infect Dis. 2017;23:2078–80. DOIPubMedGoogle Scholar
- Cajimat MNB, Rodriguez SE, Schuster IUE, Swetnam DM, Ksiazek TG, Habela MA, et al. Genomic characterization of Crimean-Congo hemorrhagic fever virus in Hyalomma tick from Spain, 2014. Vector Borne Zoonotic Dis. 2017;17:714–9. DOIPubMedGoogle Scholar
- Deyde VM, Khristova ML, Rollin PE, Ksiazek TG, Nichol ST. Crimean-Congo hemorrhagic fever virus genomics and global diversity. J Virol. 2006;80:8834–42. DOIPubMedGoogle Scholar
- Zehender G, Ebranati E, Shkjezi R, Papa A, Luzzago C, Gabanelli E, et al. Bayesian phylogeography of Crimean-Congo hemorrhagic fever virus in Europe. PLoS One. 2013;8:
e79663 . DOIPubMedGoogle Scholar - de Sanidad M, Sociales e Igualdad S. Status report and assessment of the risk of transmission of Crimean-Congo hemorrhagic fever (FHCC) in Spain, April 2017 [in Spanish] [cited 2017 May 1]. http://www.msssi.gob.es/profesionales/saludPublica/enfermedadesEmergentes/Crimea_Congo/docs/ACTUALIZACION_ER_FHCC_20.04.2017.pdf
- Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29:1969–73. DOIPubMedGoogle Scholar
- Carroll SA, Bird BH, Rollin PE, Nichol ST. Ancient common ancestry of Crimean-Congo hemorrhagic fever virus. Mol Phylogenet Evol. 2010;55:1103–10. DOIPubMedGoogle Scholar
- Chamberlain J, Cook N, Lloyd G, Mioulet V, Tolley H, Hewson R. Co-evolutionary patterns of variation in small and large RNA segments of Crimean-Congo hemorrhagic fever virus. J Gen Virol. 2005;86:3337–41. DOIPubMedGoogle Scholar
- Gergova I, Kunchev M, Kamarinchev B. Crimean-Congo hemorrhagic fever virus-tick survey in endemic areas in Bulgaria. J Med Virol. 2012;84:608–14. DOIPubMedGoogle Scholar
- Estrada-Peña A, Vatansever Z, Gargili A, Ergönul O. The trend towards habitat fragmentation is the key factor driving the spread of Crimean-Congo haemorrhagic fever. Epidemiol Infect. 2010;138:1194–203. DOIPubMedGoogle Scholar
- Spengler JR, Bente DA. Crimean-Congo hemorrhagic fever in Spain: new arrival or silent resident? N Engl J Med. 2017;377:106–8. DOIPubMedGoogle Scholar
- Filipe AR, Calisher CH, Lazuick J. Antibodies to Congo-Crimean haemorrhagic fever, Dhori, Thogoto and Bhanja viruses in southern Portugal. Acta Virol. 1985;29:324–8.PubMedGoogle Scholar
- Palomar AM, Portillo A, Santibáñez S, García-Álvarez L, Muñoz-Sanz A, Márquez FJ, et al. Molecular (ticks) and serological (humans) study of Crimean-Congo hemorrhagic fever virus in the Iberian Peninsula, 2013-2015. Enferm Infecc Microbiol Clin. 2017;35:344–7. DOIPubMedGoogle Scholar
- Lindeborg M, Barboutis C, Ehrenborg C, Fransson T, Jaenson TG, Lindgren PE, et al. Migratory birds, ticks, and crimean-congo hemorrhagic fever virus. Emerg Infect Dis. 2012;18:2095–7. DOIPubMedGoogle Scholar
- Palomar AM, Portillo A, Santibáñez P, Mazuelas D, Arizaga J, Crespo A, et al. Crimean-Congo hemorrhagic fever virus in ticks from migratory birds, Morocco. Emerg Infect Dis. 2013;19:260–3. DOIPubMedGoogle Scholar
- Vanhomwegen J, Alves MJ, Zupanc TA, Bino S, Chinikar S, Karlberg H, et al. Diagnostic assays for Crimean-Congo hemorrhagic fever. Emerg Infect Dis. 2012;18:1958–65. DOIPubMedGoogle Scholar