Volume 26, Number 12—December 2020
Synopsis
Mycoplasma bovis Infections in Free-Ranging Pronghorn, Wyoming, USA
Cite This Article
Citation for Media
Abstract
Mycoplasma bovis is 1 of several bacterial pathogens associated with pneumonia in cattle. Its role in pneumonia of free-ranging ungulates has not been established. Over a 3-month period in early 2019, »60 free-ranging pronghorn with signs of respiratory disease died in northeast Wyoming, USA. A consistent finding in submitted carcasses was severe fibrinosuppurative pleuropneumonia and detection of M. bovis by PCR and immunohistochemical analysis. Multilocus sequence typing of isolates from 4 animals revealed that all have a deletion in 1 of the target genes, adh-1. A retrospective survey by PCR and immunohistochemical analysis of paraffin-embedded lung from 20 pronghorn that died with and without pneumonia during 2007–2018 yielded negative results. These findings indicate that a distinct strain of M. bovis was associated with fatal pneumonia in this group of pronghorn.
The bacterium Mycoplasma bovis is an economically important pathogen of cattle that contributes to the multifactorial bovine respiratory disease complex. In addition to causing respiratory disease, this bacterium can cause polyarthritis, mastitis, otitis media, and a chronic pneumonia–polyarthritis syndrome, impacting beef and dairy cattle worldwide (1). Despite increased recognition of its role in economic loss in the cattle industry, M. bovis remains a clinical challenge because of a common carrier state in clinically healthy animals, variable disease expression, intermittent shedding, and the lack of rapid accurate diagnostic assays (1,2).
Clinical disease is not considered necessary to maintain M. bovis in populations, and M. bovis is commonly detected in asymptomatic adult feedlot cattle (2). Although the upper respiratory tract mucosa is a primary site for M. bovis colonization, presence of the bacterium in the lung is variable in occurrence and clinical manifestation. In 1 study, M. bovis was detected in 46% of cattle with normal lungs, 82% of cattle with acute fibrinous pneumonia, and 98% of cattle with chronic pneumonia (3). Manifestation of M. bovis–associated respiratory disease is particularly common in the wake of stress (e.g., from transportation, comingling, feedlot entry, and harsh temperatures or conditions).
In the early 2000s, M. bovis caused several high-mortality (case-fatality rate 45%) epizootics in bison (Bison bison) in North America (4). These events raised concern about emergent virulent strains, and research began to characterize isolates from different host species (5). An important difference between outbreaks of mycoplasmosis in bison and cattle is that, in the former, few or no co-infecting bacterial or viral pathogens are consistently detected (4,6–8). Although M. bovis virulence factors are poorly defined, evasion of immune response is implicated in maintaining chronic infection (9). One study found that that up to 79% of bison herds in western Canada have >1 M. bovis–seropositive animal and that 8 of 11 herds with no history of M. bovis disease had seropositive animals (10). These findings suggest that host response to M. bovis varies; some exposed bison become subclinical carriers and might also indicate a strain variation in M. bovis that influences the severity of disease.
Despite its recent recognition in bison, documented cases of M. bovis in free-ranging ruminants are rare. M. bovis was reported in farmed white-tailed deer (Odocoileus virginianus) (11) and observed in free-ranging mule deer (Odocoileus hemionus) (P. Wolff, Wildlife Disease Association, pers. comm., August 2019). Pronghorn (Antilocapra americana) are the only extant member of the family Antilocapridae and are native to expansive ranges in the western United States, southern Canada, and northern Mexico (12). Approximately 0.5–1.0 million pronghorn exist in North America (12). Herds are commonly sympatric with range cattle and ranched bison. In this article, we document M. bovis as the cause of a high-mortality outbreak of respiratory disease in a new free-ranging host, using a widely employed multilocus sequence typing (MLST) scheme to characterize associated lesions and the allelic profile of M. bovis.
Diagnostic Workup
Carcasses, tissue samples, or both were obtained from the site of the outbreak comprising a »20-km2 area northeast of Gillette, Wyoming, USA (Figure 1). The samples were submitted on behalf of the Wyoming Game and Fish Department (WGFD) for a diagnostic workup. For 2 of the 9 cases, an entire carcass was submitted and a detailed postmortem examination was performed by a board-certified pathologist, including histopathologic examination of tissues. For 7 of the 9 cases, limited tissue sampling was performed during field autopsy by WGFD. In all 9 cases, fresh or fresh frozen lung tissue was received, and M. bovis was detected by PCR. Bacteriology (aerobic and anaerobic culture) was performed on fresh or fresh frozen lung from 5 of 9 cases; 4 cases had advanced tissue autolysis and were not cultured. No consistent bacterial co-infections were detected across multiple cases (Table 1). Because Mannheimia spp. and Histophilus spp. were detected by culture in 1 case, we performed PCR for both agents on all 9 cases, yielding negative results. Molecular virology was performed on fresh or fresh frozen lung from all 9 cases; PCR assays included bovine herpesvirus 1, parainfluenza virus 3, bovine viral diarrhea virus, bovine respiratory syncytial virus, epizootic hemorrhagic disease virus, blue tongue virus, and a cervid adenovirus originally identified in mule deer and occasionally detected in pronghorn (13). No viruses were detected in any case. PCR assays were performed according to validated diagnostic protocols at the Wyoming State Veterinary Laboratory (WSVL). Histopathologic and immunohistochemical (IHC) analysis were performed on a total of 5 cases. In all cases, M. bovis was detected by IHC analysis. IHC analysis for Histophilus somni was also performed on these 5 cases; H. somni was not detected.
Histopathology
Tissues collected at autopsy for histopathologic analysis (Appendix) were fixed in 10% buffered formalin and processed conventionally before embedding in paraffin wax. Sections cut at 5 µm were stained with hematoxylin and eosin. IHC analysis of M. bovis and H. somni was performed on lung tissue derived from the same blocks as used in the hematoxylin and eosin assays (Appendix).
Mycoplasma Culture
Approximately 20 mg of lung tissue was placed in a mycoplasma enrichment broth (Hardy Diagnostics’ Mycoplasma Broth; Hardy Diagnostics, https://hardydiagnostics.com) and incubated with a loose lid at 37°C in 10% CO2 for 72 h. Subsequently, 100 μL of broth was inoculated onto a commercial Mycoplasma spp. medium (Hardy Diagnostics’ Mycoplasma Agar with Cefoperazone) and spread evenly over the entire plate with a sterile swab. Plates were incubated at 37°C in 10% CO2 for 72–240 h, depending on appearance of colony growth. Colonies from each isolate were analyzed by matrix-assisted laser desorption/ionization-time of flight mass spectrometry (Bruker’s Biotyper, https://www.bruker.com) according to the manufacturer’s instructions for identification. Additional colonies were used for whole-genome sequencing.
Sequencing
16S
After DNA extraction from fresh lung tissue, a portion of the 16S ribosomal RNA gene was amplified by PCR using universal Mycoplasma primers (13) at the Washington Animal Disease Diagnostic Laboratory. PCR amplicons were directly sequenced, and a GenBank BLAST search was performed (https://blast.ncbi.nlm.nih.gov) on consensus sequence from 2 forward and 2 reverse high-quality reads. This initial confirmation was performed on the first sample only, and isolates from subsequent samples were confirmed by whole-genome sequencing.
Polymerase chain reaction
Diagnostic M. bovis PCR
DNA was extracted from fresh lung tissue and PCR was performed targeting the M. bovis 16S ribosomal RNA gene (Appendix). Confirmation of diagnosis from case 1 (Table 1) by PCR targeting of the uvrC gene was performed at the Washington Animal Disease Diagnostic Laboratory (15).
Survey of Formalin-Fixed Paraffin-Embedded Archival Lung Tissue
DNA was extracted from formalin-fixed, paraffin embedded lung tissue curls cut at a thickness of 20 μm. In brief, 1–2 curls per sample were dewaxed by using xylene and ethanol according to the DNeasy Blood and Tissue kit’s recommended protocol (QIAGEN, https://www.qiagen.com). The tissue extraction proceeded overnight at 56°C, according to manufacturer instructions. Cases selected for PCR were based on the availability of lung tissue from pronghorn in archived wax blocks, which are retained for 15 years because of limited storage space. A total of 20 cases (13 in pronghorn with previously diagnosed pneumonia) were identified; all cases originated from Wyoming (Table 2). After DNA extraction, PCR was performed as described previously for the diagnostic M. bovis PCR assay.
Diagnostic M. ovipneumoniae PCR
A 2 × 2 cm section of fresh lung tissue was placed into 2 mL modified tryptic soy broth and homogenized for 120 seconds. The homogenous solution was transferred to a snap cap tube and incubated at 37°C with 10% CO2 for 48 h. After centrifugation of 1 mL, the pellet was resuspended and used in a PCR reaction as previously described (16).
adh-1 PCR
DNA was extracted from the Mycoplasma broth of each sample stored at −80°C by using the QIAGEN DNeasy Blood and Tissue Kit fluid protocol. In brief, 200 μL was extracted following manufacturer instructions. Forward and reverse primers targeting the adh-1 gene (0.5 μmol/L of each) were used in a 50 μL reaction containing 22.5 μL GoTaq green master mix (Promega, https://www.promega.com), 1.5 μL of 50 nM MgCl2, and nuclease-free water (5).
MLST Analysis
Paired fastq reads of »250 bp were analyzed as follows: trimming of indexes, primers, low quality (phred <20), and short reads (<50 bp) using Cutadapt (17); mapping of trimmed reads to the genome of M. bovis international reference strain PG45 (GenBank accession no. NC_014760) using Bowtie2 (18); conversion of .sam files to .bam files using Samtools (19); and viewing of sorted .bam files in Geneious Prime 2019.1.3 (https://www.geneious.com). Consensus sequences were generated from mapped reads by using the highest quality parameter in Geneious Prime as a threshold. “N” was assigned to sites with coverage <3 to represent missing data. Consensus sequences were trimmed to loci employed in the MLST scheme described by Register et al. (5) and concatenated in frame. Concatenated sequences were compared for 4 M. bovis isolates recovered from the lung samples of 4 different pronghorn across the following MLST genes: alcohol dehydrogenase (adh-1), glutamate tRNA ligase (gltX), glycerol-3-phosphate dehydrogenase (gpsA), DNA gyrase subunit B (gyrB), phosphate acetyltransferase-2 (pta-2), thymidine kinase (tdk), and transketolase (tkt) (5). Isolates derived in our study were compared with those from the University of Oxford Mycoplasma bovis MLST website (https://pubmlst.org/mbovis) (20). An aligned fasta file was obtained for all publicly available isolates missing the adh-1 gene (i.e., nontypeable isolates). The representative sequence from the 4 identical pronghorn isolates was aligned to the fasta file comprising all nontypeable isolates by using Muscle (21), and model selection was performed in MEGA X (22). Sequence alignments were subjected to maximum-likelihood phylogenetic analyses under the Hasegawa–Kishono–Yano substitution model using PhyML (23) with 10,000 bootstrap replicates for support.
Disease Outbreak
At least 60 pronghorn died during February–April 2019 within a total area of »13 km2. WGFD received initial reports of »30 carcasses in early February 2019. An additional 20 pronghorn deaths were identified within 8 km of the site of initial reports within 1 month, and the affected area expanded as wintering herds began to disperse with warmer weather. In March, a herd of »10 pronghorn moved to the same area and began dying within 2 weeks. Landowners reported that affected pronghorn appeared lethargic. Such animals were typically dead within 24 hours. Because of logistics of finding fresh carcasses in remote areas on private land during 2 major winter storms, only a fraction of the dead pronghorn could be sampled. Two carcasses were obtained for autopsy, and tissues were obtained from an additional 3 cases for histopathologic examination and PCR. Samples were collected from 4 additional animals for PCR only. The extent of the die-off could not be estimated until improved weather conditions allowed WGFD biologists to conduct ground and aerial surveys. Bison, cattle, or other free-ranging ungulates (i.e., deer, elk, and moose) deaths associated with pneumonia were not reported in the area during this outbreak. The closest captive bison herd was located »64 km south of the outbreak site. Although pronghorn deaths occur in winter because of starvation, predation, and vehicular collision, 60 deaths in a small area is unusual.
Diagnosis and Characterization of Lesions
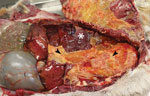
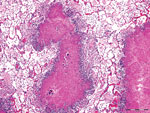
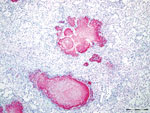
Gross lesions were characterized by severe, regionally extensive to diffuse, bilateral fibrinous pleuropneumonia affecting an estimated 50%–100% of lung parenchyma (Figure 2). Histopathologic examination revealed fibrinosuppurative pneumonia with caseonecrotic foci centered on bronchi and bronchioles. Caseonecrotic foci were characterized by central granular eosinophilic material and necrotic leukocytes surrounded by degenerate and intact neutrophils. Some foci were partly mineralized (Figure 3). Lesions were interpreted as acute to subacute, because features of chronic infection, such as extensive fibrosis, were absent (24). In all 5 cases for which histopathologic examination was performed, pulmonary lesions were strongly immunoreactive for M. bovis antigen, with characteristically strong staining at the margins of necrotic foci as described in affected cattle and bison (3) (Figure 4). The pronghorn in case 9 (Table 1) had pulmonary abscesses up to 1 cm diameter characterized by coagulative necrosis surrounded by a thin band of fibrosis. Features characteristic of mannheimiosis, histophilosis, or both, such as neutrophils with oat cell morphology, were absent. The pronghorn in case 9 also had fibrinosuppurative synovitis and conjunctivitis. M. bovis antigen was detected in the conjunctiva by IHC analysis. In addition, acute centrilobular hepatic necrosis was identified in case 9. We attributed this finding to hypoxia secondary to severe pneumonia.
Lung samples from the 9 pronghorn cases were positive for M. bovis by culture and PCR (Table 1). 16S sequencing revealed that the isolate from case 1 most closely matched that of M. bovis (100% sequence identity [553/553 bp]; GenBank no. KX462388). The next closest match was 99% identity (549/553 bp) to M. agalactiae (GenBank no. AF332750). M. ovipneumoniae PCR was performed on samples of lung; all results were negative (data not shown).
Aside from the consistent detection of M. bovis, aerobic culture results were inconsistent. The pronghorn in case 1 contained Trueperella pyogenes, the pronghorn in case 9 contained both Mannheimia spp. and Histophilus spp., and the pronghorn in cases 3 and 4 had a mixture of bacterial species not typically associated with pneumonia interpreted as incidental (Table 1). The pronghorn in cases 2 and 9 had a mild lungworm infection, including nematode larvae and eggs histologically consistent with Dictyocaulus spp. parasitic infection.
Retrospective Study
We performed a retrospective survey for M. bovis on 20 archived WSVL cases of pronghorn deaths with and without pneumonia (Table 2); embedded lesioned lung from the animals in 2 of the 9 positive cases among the 2019 pronghorn deaths were used as positive controls. All archived pronghorn lung tissues were negative for M. bovis by PCR and IHC analysis, and none had lesions suggestive of mycoplasmosis.
MLST and Phylogenetics
Genome sequencing of the 4 M. bovis isolates recovered from the pronghorn carcasses was performed at the Wyoming Public Health Laboratory. All isolates from pronghorn had 100% sequence identity at loci used for MLST (5). All assemblies contained an apparent deletion of 1 of the 7 MLST target genes, adh-1 (5). Sequences for the 6 remaining loci are available through GenBank (accession nos. MT782331–6). To confirm the adh-1 deletion, DNA from M. bovis pronghorn isolates was amplified by PCR using the adh-1 primers specified for MLST as described previously (5). DNA extracted from a cattle isolate of M. bovis was strongly positive, whereas DNA from the pronghorn isolates produced no visible band upon gel electrophoresis.
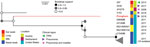
Deletion of adh-1 has been identified in M. bovis isolates derived from cattle (25; https://pubmlst.org/mbovis; K.B. Register, unpub. data). Phylogenetic analysis of these MLST-nontypeable isolates based on DNA sequences of the other 6 MLST targets revealed that the pronghorn isolates we evaluated are divergent from all others typed to date but are most similar to a group of isolates obtained from cattle in the United States since 2011 (Figure 5).
M. bovis is uncommon in free-ranging ungulates. Accounts are limited to cases in farmed white-tailed deer (11), and free-ranging mule deer (26; P. Wolff, Wildlife Disease Association, pers. comm., August 2019). Lesions in lung were compatible with the lesions attributable to M. bovis in cattle and bison (2,6). The distribution of M. bovis antigen in IHC preparations of caseonecrotic foci is typical of fatal mycoplasmosis in cattle (2,3).
To determine whether M. bovis had been previously overlooked in Wyoming pronghorn, we queried the WSVL diagnostic database. We identified 20 cases from different geographic regions of Wyoming that occurred during 2007–2019. This group included 13 cases in pronghorn with previously diagnosed pneumonia and 7 without (Table 2). Although the M. bovis PCR assay used at WSVL has not been validated for formalin-fixed, wax-embedded tissue, positive dewaxed lung samples from pronghorn in the 2019 cases were used as a control. On the basis of these 20 samples, no M. bovis infections in pronghorn before 2019 was evident.
Draft genome sequences were obtained for isolates of M. bovis from 4 pronghorn in the 2019 group. Compared with MLST data available on >700 isolates, only 9 complete genome assemblies from other host species, such as goat, bison, or cattle, were available. Thus, we determined sequence type by extracting regions of 7 genes used in MLST typing (5). The 4 isolates were identical across these loci and contained a deletion encompassing the adh-1 gene. This deletion was confirmed by using adh-1 specific primers as described previously (5). These isolates were compared with others with a deletion at the adh-1 locus (i.e., nontypeable isolates). Although the adh-1 gene deletion has been identified in M. bovis from bison and mule deer (K.B. Register, unpub. data), only sequences from bovine isolates are currently available in the Mycoplasma bovis pubMLST database (https://pubmlst.org/mbovis). The isolates from pronghorn are divergent from other published isolates but are most similar to those from US cattle compared with bovine isolates from Austria or Switzerland (Figure 5).
Although the adh-1 deletion has not yet been thoroughly characterized, the earliest identification of this variant is from 2008. The deletion might be relatively recent and might be associated with expansion of M. bovis host range or emergence in new species. Additional research is needed to investigate the possible association between the adh-1 gene deletion and the recent appearance of M. bovis in pronghorn. It will be of interest to investigate the entire genome for other whole-gene deletions or insertions and to correlate whether genomic changes are associated with certain hosts, levels of virulence, or both.
Surveillance of pronghorn samples submitted to WSVL has not identified M. bovis in other areas of the state at this time. No additional cases have been diagnosed in northeast Wyoming since April 2019. As part of a surveillance effort, we have recently performed M. bovis PCR on lung tissue DNA of any ungulate submitted to WSVL. We have found no evidence of chronically infected pronghorn or other wildlife reservoirs of this bacterium. The host species of origin in this outbreak is unknown. Given the frequency of M. bovis in asymptomatic cattle and bison and its rarity of detection in free-ranging ungulates, transmission to pronghorn from a livestock reservoir seems likely.
Our findings strongly implicate M. bovis as a primary pathogen in pronghorn, resulting in fatal pneumonia in absence of other respiratory pathogens with changes comparable to those in bison with fatal mycoplasmosis. M. bovis as a primary pathogen in bison is in contrast to M. bovis in adult cattle, where the bacterium tends to occur most commonly as 1 component of chronic, polymicrobial respiratory disease (24,27). The pronghorn M. bovis infection is more analogous to mycoplasmosis in bison, where it is known to be a primary cause of pneumonia, arthritis, pharyngitis, and reproductive disorders (6–8).
Our findings document M. bovis infection in pronghorn and highlight the possible health implications for other wildlife populations and the potential risk for transmission at the wildlife–livestock interface. Furthermore, we document M. bovis genetic variation in association with virulent disease in pronghorn, supporting hypotheses that M. bovis might be expanding in host range and in disease expression. We therefore recommend that M. bovis be considered as a differential diagnosis for pneumonia in wildlife, particularly in outbreak scenarios. Traditionally, pronghorn are not considered a major source of disease threats to either cattle or bison and are therefore widely tolerated on commercial operations. Additional studies are needed to establish transmission potential and direction, which will elucidate the importance of M. bovis at the wildlife–livestock interface.
Dr. Malmberg is a veterinary pathologist at the Wyoming State Veterinary Laboratory and an assistant professor at the University of Wyoming in Laramie. Her research interests include transmission of naturally occurring diseases at wildlife–livestock and urban–rural interfaces.
References
- Pfützner H, Sachse K. Mycoplasma bovis as an agent of mastitis, pneumonia, arthritis and genital disorders in cattle. Rev Sci Tech. 1996;15:1477–94. DOIPubMedGoogle Scholar
- Jeff L. Caswell, Archambault M. Mycoplasma bovis pneumonia in cattle. Anim Health Res Rev. 2008;8:161–86.
- Gagea MI, Bateman KG, Shanahan RA, van Dreumel T, McEwen BJ, Carman S, et al. Naturally occurring Mycoplasma bovis-associated pneumonia and polyarthritis in feedlot beef calves. J Vet Diagn Invest. 2006;18:29–40. DOIPubMedGoogle Scholar
- Kyathanahalli S, Janardhan MH. Neil Dyer, Richard D. Oberst, Brad M. DeBey. Mycoplasma bovis outbreak in a herd of North American bison (Bison bison). J Vet Diagn Invest. 2010;22:797–801. DOIGoogle Scholar
- Karen B. Register, Luke Thole, Ricardo F. Rosenbush, Minion FC. Multilocus sequence typing of Mycoplasma bovis reveals host-specific genotypes in cattle versus bison. Vet Micro. 2015;175:92–8. DOIGoogle Scholar
- Dyer N, Hansen-Lardy L, Krogh D, Schaan L, Schamber E. An outbreak of chronic pneumonia and polyarthritis syndrome caused by Mycoplasma bovis in feedlot bison (Bison bison). J Vet Diagn Invest. 2008;20:369–71. DOIPubMedGoogle Scholar
- Dyer N, Register KB, Miskimins D, Newell T. Necrotic pharyngitis associated with Mycoplasma bovis infections in American bison (Bison bison). J Vet Diagn Invest. 2013;25:301–3. DOIPubMedGoogle Scholar
- Register KB, Woodbury MR, Davies JL, Trujillo JD, Perez-Casal J, Burrage PH, et al. Systemic mycoplasmosis with dystocia and abortion in a North American bison (Bison bison) herd. J Vet Diagn Invest. 2013;25:541–5. DOIPubMedGoogle Scholar
- Boothby JT, Jasper DE, Zinkl JG, Thomas CB, Dellinger JD. Prevalence of mycoplasmas and immune responses to Mycoplasma bovis in feedlot calves. Am J Vet Res. 1983;44:831–8.PubMedGoogle Scholar
- Bras AL, Suleman M, Woodbury M, Register K, Barkema HW, Perez-Casal J, et al. A serologic survey of Mycoplasma spp. in farmed bison ( Bison bison) herds in western Canada. J Vet Diagn Invest. 2017;29:513–21. DOIPubMedGoogle Scholar
- Dyer NW, Krogh DF, Schaan LP. Pulmonary mycoplasmosis in farmed white-tailed deer (Odocoileus virginianus). J Wildl Dis. 2004;40:366–70. DOIPubMedGoogle Scholar
- International Union for Conservation of Nature. IUCN red list of threatened species 2016: e.T1677A115056938. IUCN SSC Antelope Specialist Group 2016 Antilocapra americana [cited 2020 Jul 19]. DOIGoogle Scholar
- Kong F, James G, Gordon S, Zelynski A, Gilbert GL. Species-specific PCR for identification of common contaminant mollicutes in cell culture. Appl Environ Microbiol. 2001;67:3195–200. DOIPubMedGoogle Scholar
- Andrews S. FastQC: a quality control tool for high throughput sequence data. 2010 [cited 2019 Aug 20]. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
- Naikare H, Bruno D, Mahapatra D, Reinisch A, Raleigh R, Sprowls R. Development and evaluation of a novel Taqman real-time PCR assay for rapid detection of Mycoplasma bovis: comparison of assay performance with a conventional PCR assay and another Taqman real-time PCR assay. Vet Sci. 2015;2:32–42. DOIPubMedGoogle Scholar
- Ziegler JC, Lahmers KK, Barrington GM, Parish SM, Kilzer K, Baker K, et al. Safety and Immunogenicity of a Mycoplasma ovipneumoniae bacterin for domestic sheep (Ovis aries). PLoS One. 2014;9:
e95698 . DOIPubMedGoogle Scholar - Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–2. DOIGoogle Scholar
- Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9. DOIPubMedGoogle Scholar
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al.; 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9. DOIPubMedGoogle Scholar
- Jolley KA, Bray JE, Maiden MCJ. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018;3:124. DOIPubMedGoogle Scholar
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7. DOIPubMedGoogle Scholar
- Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol. 2018;35:1547–9. DOIPubMedGoogle Scholar
- Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21. DOIPubMedGoogle Scholar
- Maunsell FP, Woolums AR, Francoz D, Rosenbusch RF, Step DL, Wilson DJ, et al. Mycoplasma bovis infections in cattle. J Vet Intern Med. 2011;25:772–83. DOIPubMedGoogle Scholar
- Josi C, Bürki S, Stojiljkovic A, Wellnitz O, Stoffel MH, Pilo P. Bovine epithelial in vitro infection models for Mycoplasma bovis. Front Cell Infect Microbiol. 2018;8:329. DOIPubMedGoogle Scholar
- Register KB, Jelinski MD, Waldner M, Boatwright WD, Anderson TK, Hunter DL, et al. Comparison of multilocus sequence types found among North American isolates of Mycoplasma bovis from cattle, bison, and deer, 2007-2017. J Vet Diagn Invest. 2019;31:899–904. DOIPubMedGoogle Scholar
- Maunsell FP, Donovan GA. Mycoplasma bovis Infections in young calves. [vii.]. Vet Clin North Am Food Anim Pract. 2009;25:139–77, vii. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: October 29, 2020
Table of Contents – Volume 26, Number 12—December 2020
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Kerry Sondgeroth, Wyoming State Veterinary Laboratory, 1174 Snowy Range Rd, Laramie, WY 82070, USA; emailksondger@uwyo.edu
Top