Volume 27, Number 12—December 2021
Dispatch
Potential Mosquito Vectors for Shuni Virus, South Africa, 2014–2018
Cite This Article
Citation for Media
Abstract
Shuni virus is associated with neurologic and febrile illness in animals and humans. To determine potential vectors, we collected mosquitoes in South Africa and detected the virus in species of the genera Mansonia, Culex, Aedes, and Anopheles. These mosquitoes may be associated with Shuni virus outbreaks in Africa and emergence in other regions.
The genus Orthobunyavirus (family Peribunyaviridae) includes emerging arthropodborne viruses associated with human and animal disease worldwide (1). In 1966, orthobunyavirus Shuni virus (SHUV) was isolated from a cow, Culicoides midges, and a febrile child in Nigeria (2); SHUV recently emerged in Israel, where it has been associated with birth defects in ruminants (3). SHUV has been associated with neurologic disease in horses and wildlife (4,5) and was recently implicated in human cases of neurologic disease in South Africa (6). SHUV was detected in field-caught Culex theileri mosquitoes in the 1970s (5), and Culicoides midges have been suggested as vectors (7). We investigated mosquitoes collected in northeastern parts of South Africa to identify their potential as vectors of orthobunyaviruses in the Simbu serogroup of arboviruses, including SHUV.
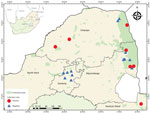
We collected mosquitoes across 5 provinces of South Africa (Figure 1). Site selection was based on historical outbreaks of arboviruses, including SHUV, in animals (4,5) and humans (6). During January 2014–May 2017, we collected mosquitoes monthly; we performed additional collections in 2017 in and around the Kruger National Park (8). In 2018, we performed 1 collection per site during January–May.
We used multiple types of dry ice (carbon dioxide) baited traps: nets, CDC miniature light traps (https://www.johnwhock.com), and BG-Sentinel traps (https://www.bg-sentinel.com). We set traps during 3:30–6:00 pm and emptied them during 5:00–8:00 am. We killed mosquitoes by freezing and then morphologically identified them to the species level. We pooled females (<50 individuals) by species, collection site, and month. We selected mosquitoes for screening from pools collected during January–June, which represents late summer and autumn, when arbovirus infections in animals and humans in South Africa increase. We obtained climate data from the South African Weather Service (http://www.weathersa.co.za).
For the virus assays, we produced homogenate pools by placing 5 sterile glass beads in microcentrifuge tubes containing 2 mL of reconstituted minimum essential medium, which we then vigorously shook and clarified. The resulted supernatant was stored at −80°C. To extract viral RNA from 200 μL homogenate, we used an RNeasy mini kit (QIAGEN, https://www.qiagen.com) according to the manufacturer’s instructions. We screened extracted RNA by using 2 PCRs, each targeting the nucleocapsid (NP) gene on the small (S) segment: a Simbu serogroup/orthobunyavirus–specific one-step TaqMan real-time reverse transcription PCR targeting a 152-bp fragment (4) and an SHUV nested real-time RT-PCR targeting a 460-bp fragment (9). In an attempt to obtain larger fragments, we performed an SHUV conventional PCR with published primers (10).
For mosquito barcoding (species identification), we extracted DNA from 50 μL of the homogenate by using a QIAGEN DNeasy Blood & Tissue Kit according to the manufacturer’s instructions. The subunit I of the cytochrome oxidase gene was amplified by using universal primers (11).
All products of the expected size were sequenced by Sanger sequencing at the Forestry Agriculture Bioinformatics Institute, University of Pretoria (Pretoria, South Africa). We compared the resulting sequences by using BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) with sequences available from GenBank, including SHUV strains from South Africa, Nigeria, and Israel and other representative members of Simbu serogroup. For the cytochrome oxidase gene, we selected representative mosquito sequences from GenBank and BOLD (https://v3.boldsystems.org). We compiled multiple sequence alignments by using MAFFT (https://mafft.cbrc.jp/alignment/software), produced maximum-likelihood trees by using MEGA 7.0 (https://www.megasoftware.net), and calculated maximum-likelihood estimates of mosquito infection rates by using PooledInfRate (https://www.cdc.gov/ncidod/dvbid/westnile/software.htm).
Of the 64,603 adult mosquitoes collected as described (8,12), we tested 39,035 females. A total of 11 pools were positive for SHUV (Table 1). No other orthobunyaviruses were detected. Positive pools for SHUV were detected in conservation areas (6/11, 54.5%) and rural areas (5/11, 45.5%) (Figure 1). Populations of the SHUV-positive mosquito species peaked with the heavy rains and with the highest mean air temperatures (Appendix Figure 1), which promote establishment of breeding sites and favorable habitats for developing stages and subsequent population growth.
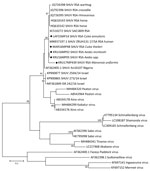
The maximum-likelihood phylogeny based on the genus Orthobunyavirus PCR fragment of 152-bp of the S segment showed that all SHUV viruses from the mosquitoes clustered with the Simbu serogroup (Appendix Figures 1, 2) and were closest to SHUV on the basis of p-distance analyses (data not shown). For 5 samples, a larger region of the S segment could be amplified to confirm the clustering with SHUV strains previously identified in horses and wildlife from South Africa (Figure 2) and p-distances of 94%–100% with strains previously identified in South Africa, Israel, and Nigeria. Mosquito barcodes consisting of 517-bp were used to build a maximum-likelihood tree (Appendix Table 1, Figures 1–3). The barcoding confirmed all morphologic identifications except for a pool of damaged Aedes spp. mosquitoes and for Ae. subargenteus mosquitoes (for which no other sequence was available in the databases).
Of the 11 pools of SHUV-positive mosquitoes, species belonged to the genera Mansonia (5 pools), Aedes (3 pools), Culex (2 pools), and Anopheles (1 pool) (Table 2). Previously, SHUV had been detected in Cx. theileri mosquitoes collected in the 1970s near Johannesburg, South Africa (5). In that study, 2 pools of SHUV-positive Cx. theileri mosquitoes were also identified, although mosquitoes of this species were not abundant in the sites detected.
The highest rate of SHUV detection was in Mansonia uniformis mosquitoes, which were found in high numbers at the Shuni virus–positive pool collection sites. Three other arboviruses have been isolated from M. uniformis mosquitoes in South Africa: Wesselsbron, Ndumu, and Spondeweni (13). M. africana mosquitoes tested positive, but only small numbers of these mosquitoes were collected. Mansonia spp. mosquitoes can feed readily on humans and animals (13) and could have a potential epidemiologic role as bridge species for transmission between animals and humans.
Mosquitoes of other species that tested positive included Aedes mcintoshi and Ae. subargenteus. Positive Ae. mcintoshi mosquitoes were collected from Mnisi, where they were the most abundant Aedes spp. at that site. They are considered nonspecific/opportunistic feeders and have a broad range of mammal hosts (14). Ae. subargenteus mosquitoes are tree hole mosquitoes and are either rare in South Africa (14) or are not attracted to the traps used in our study. Although little information about those mosquitoes is available, they might have a strong preference for biting humans (14).
Although SHUV has been detected in mosquitoes, recent studies have also implicated Culicoides spp. midges as potential competent vectors (15). An investigation of the vector competence of Culicoides midges and laboratory-reared Cx. pipiens and Ae. aegypti mosquitoes for SHUV (7) indicated that neither species of mosquito was susceptible but that Culicoides midges demonstrated the capacity to transmit SHUV. No Ae. aegypti and Cx. pipiens field-caught mosquitoes tested positive for SHUV in this or other studies. Vector competence studies that used SHUV-positive species of mosquitoes identified in our study may define appropriate mosquito vectors and their role in the transmission of SHUV to animals and humans in Africa and the risk to areas where they are found outside the continent.
Entomologic surveillance for orthobunyaviruses revealed a wide range of potential mosquito vectors for SHUV. We identified SHUV in different species of mosquitoes in South Africa, where cases with neurologic signs have been detected in animals (4,5) and humans (6). The identified mosquito species have also been associated with other arboviruses across Africa. SHUV recently emerged in Israel, where it is associated with neurologic disease and birth defects in animals (3). Mosquitoes of the identified species are potential vectors of SHUV and may be associated with SHUV outbreaks in Africa and further emergence in new regions.
Dr. Guarido completed her PhD degree and performs postdoctoral work at the Faculty of Veterinary Sciences, University of Pretoria. Her primarily research interest is the study of mosquitoborne viruses, with a focus on the entomology aspects of arbovirus transmission cycles.
Acknowledgments
We are grateful to Lapalala Wilderness, Marataba Conservation, South African National Park, for logistical assistance and permission to collect mosquitoes. We acknowledge Leo Braack and Danny Govender for their valuable contribution to mosquito collection. We are grateful to the South Africa Weather Service for providing data for this study.
This study was supported by Cooperative Agreement no. 5 NU2GGH001874-02-00, funded by the Centers for Disease Control and Prevention. It was also funded in part through scholarships from the National Research Foundation and The World Academy of Sciences. A.P.G.A. has been a recipient of the Visiting Professor Programme by the University of Pretoria, South Africa, and acknowledges the Global Health and Tropical Medicine unit. This study was also funded by the Gratama Fund (grant no. 2016.08), the Uyttenboogaart-Eliasen foundation (SUB.2016.12.08), and the RCN-IDEAS travel grant to M.S. and E.E.G.
References
- Hughes HR, Adkins S, Alkhovskiy S, Beer M, Blair C, Calisher CH, et al.; Ictv Report Consortium. ICTV virus taxonomy profile: Peribunyaviridae. J Gen Virol. 2020;101:1–2. DOIPubMedGoogle Scholar
- Causey OR, Kemp GE, Causey CE, Lee VH. Isolations of Simbu-group viruses in Ibadan, Nigeria 1964-69, including the new types Sango, Shamonda, Sabo and Shuni. Ann Trop Med Parasitol. 1972;66:357–62. DOIPubMedGoogle Scholar
- Golender N, Brenner J, Valdman M, Khinich Y, Bumbarov V, Panshin A, et al. Malformations caused by Shuni virus in ruminants, Israel, 2014–2015. Emerg Infect Dis. 2015;21:2267–8. DOIPubMedGoogle Scholar
- Steyn J, Motlou P, van Eeden C, Pretorius M, Stivaktas VI, Williams J, et al. Shuni virus in wildlife and non-equine domestic animals in South Africa. Emerg Infect Dis. 2020;26:1521–5. DOIPubMedGoogle Scholar
- van Eeden C, Williams JH, Gerdes TG, van Wilpe E, Viljoen A, Swanepoel R, et al. Shuni virus as cause of neurologic disease in horses. Emerg Infect Dis. 2012;18:318–21. DOIPubMedGoogle Scholar
- Motlou T, Venter M. Shuni virus in cases of neurological diseases in humans, South Africa. Emerg Infect Dis. 2020;27:567–9.
- Möhlmann TWR, Oymans J, Wichgers Schreur PJ, Koenraadt CJM, Kortekaas J, Vogels CBF. Vector competence of biting midges and mosquitoes for Shuni virus. PLoS Negl Trop Dis. 2018;12:
e0006993 . DOIPubMedGoogle Scholar - Gorsich EE, Beechler BR, van Bodegom PM, Govender D, Guarido MM, Venter M, et al. A comparative assessment of adult mosquito trapping methods to estimate spatial patterns of abundance and community composition in southern Africa. Parasit Vectors. 2019;12:462. DOIPubMedGoogle Scholar
- Van Eeden C, Zaayman D, Venter M. A sensitive nested real-time RT-PCR for the detection of Shuni virus. J Virol Methods. 2014;195:100–5. DOIPubMedGoogle Scholar
- Bowen MD, Trappier SG, Sanchez AJ, Meyer RF, Goldsmith CS, Zaki SR, et al.; RVF Task Force. A reassortant bunyavirus isolated from acute hemorrhagic fever cases in Kenya and Somalia. Virology. 2001;291:185–90. DOIPubMedGoogle Scholar
- Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol. 1994;3:294–9.PubMedGoogle Scholar
- Guarido MM, Riddin MA, Johnson T, Braack LEO, Schrama M, Gorsich EE, et al. Aedes species (Diptera: Culicidae) ecological and host feeding patterns in the north-eastern parts of South Africa, 2014-2018. Parasit Vectors. 2021;14:339. DOIPubMedGoogle Scholar
- Sharp BL, Appleton CC, Thompson DL, Meenehan G. Anthropophilic mosquitoes at Richards Bay, Natal, and arbovirus antibodies in human residents. Trans R Soc Trop Med Hyg. 1987;81:197–201. DOIPubMedGoogle Scholar
- Muspratt J. The Stegomyia mosquitoes of South Africa and some neighbouring territories. Memoirs of the Entomological Society of Southern Africa. 1956;4:1–138.
- Snyman J, Venter GJ, Venter M. An investigation of Culicoides (Diptera: Ceratopogonidae) as potential vectors of medically and veterinary important arboviruses in South Africa. Viruses. 2021;13:1978. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: November 04, 2021
Table of Contents – Volume 27, Number 12—December 2021
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Marietjie Venter, Zoonotic Arbo- and Respiratory Virus Program, Centre for Viral Zoonosis, Department Medical Virology, Pathology Building, Prinshof Campus South, University of Pretoria, Private Bag X323, Gezina 0031, South Africa
Top