Volume 27, Number 12—December 2021
Research
Novel Assay to Measure Seroprevalence of Zika Virus in the Philippines
Cite This Article
Citation for Media
Abstract
Zika virus (ZIKV) is a member of the Flaviviridae family, which includes other clinically notable viruses such as the 4 dengue virus serotypes (DENV-1–4). Distinguishing DENVs from ZIKV using the established serologic assays widely used for monitoring DENV transmission is difficult because of antibody cross-reactivity between these closely related flaviviruses. We describe a modified and improved recombinant envelope domain III–based serologic assay for detecting ZIKV type-specific antibodies in regions with endemic DENV transmission. When the assay was used to measure ZIKV seroprevalence in 2017 among children 9–14 years of age living in a region of the Philippines with endemic DENV transmission, we observed a ZIKV seroprevalence of 18%. Investigators should consider using the ZIKV envelope domain III–based assay, which is simple and readily adaptable for use in standard clinical and public health laboratories, to assess ZIKV seroprevalence in areas with endemic DENV transmission.
Zika virus (ZIKV) is a positive sense RNA virus of the Flaviviridae family, which includes several medically notable arboviruses such as dengue (DENV), yellow fever, Japanese encephalitis virus, and West Nile virus. Before the 2015 epidemic in the Americas that spread to >40 countries and infected >1 million people (1), ZIKV was considered rare and responsible for minor epidemics in East Africa and parts of Asia. Although most ZIKV infections are asymptomatic or clinically mild, the epidemic in the Americas revealed that the virus can cause serious neurologic problems in some persons and severe teratogenic effects in pregnant women (2–4). Because ZIKV and DENV-1–4 share the same Aedes mosquito vectors, areas in the Americas most affected by ZIKV also experience endemic DENV transmission. The ZIKV pandemic in the Americas led to novel observations and questions about its epidemiology and pathogenesis in regions with endemic DENV transmission. Recent studies indicate that cross-reactive immunity between ZIKV and DENV can lead to protection or to more severe disease depending on the context (5–7).
Although sporadic transmission of Asian lineages of ZIKV in Southeast Asia and Pacific islands is well-documented (8,9), its prevalence in the region has been difficult to estimate using current serologic assays because of intense transmission of multiple DENV serotypes and antibody cross-reactivity between DENV and ZIKV. Most serologic assays for flaviviruses measure antibodies binding to viral-envelope glycoprotein (E protein) because this antigen is a major target of human antibodies. The Flavivirus E protein contains immunodominant antibody epitopes that are conserved (cross-reactive) between different flaviviruses or unique to each virus (type-specific) (10–12).
Traditional Flavivirus serologic assays exhibit poor specificity in distinguishing DENV from ZIKV infections because these assays use whole virions or E proteins containing conserved epitopes as antigens (13–15). More recently, the ZIKV epidemic in the Americas spurred the development of recombinant viral antigens and serologic assays for distinguishing ZIKV from DENV (16–18). We previously described a serologic assay using domain III of the ZIKV E protein (EDIII) to detect ZIKV type-specific antibodies among persons in areas with DENV and ZIKV cocirculation (19). Here we describe development of a second-generation ZIKV EDIII–based serologic assay and its use to measure the seroprevalence of ZIKV among children 9–14 years of age in the Cebu Province of the Philippines.
ZIKV EDIII Antigen Production
We expressed a codon-optimized gene encoding for EDIII from ZIKV strain H/PF/2013 in Expi293 cells as a fusion protein containing a human serum albumin signal peptide for secretion, a polyhistidine tag (his-tag) for affinity purification, and a HaloTag (Promega, https://www.promega.com) for biotinylation (20). We deposited the nucleotide sequence of the construct into Genbank (accession no. MZ592925). HaloTag enables single, site-specific biotinylation distant from the folded EDIII protein. We purified recombinant EDIII antigen from the culture supernatant using nickel-nitrilotriacetic acid agarose (QIAGEN, https://www.qiagen.com) and biotinylated it using HaloTag PEG biotin ligand (Promega), according to manufacturer protocol. We analyzed the identity and purity of the biotinylated EDIII antigen using SDS-PAGE (sodium dodecyl-sulfate polyacrylamide gel electrophoresis) mobility-shift analysis.
ZIKV EDIII ELISA
We coated a 96-well high binding microtiter plate (Greiner Bio-One, https://www.gbo.com) with 50 μL of streptavidin at 4 μg/mL in tris-buffered saline (TBS, pH 7.4) for 1 h at 37°C. We captured the biotinylated EDIII at 2 μg/mL in TBS, washed the plate 3 times with wash buffer (TBS containing 0.2% Tween 20), and then blocked it with 100 μL of blocking solution (3% milk in TBS containing 0.05% Tween 20) for 1 h at 37°C. After removing the blocking solution, we added 50 μL of heat-inactivated (56°C for 30 min) serum sample at 1:20 or indicated dilutions in blocking buffer and incubated for 1 hour at 37°C. After washing the plate in the wash buffer, we added 50 μL of alkaline phosphatase-conjugated secondary goat anti-human secondary IgG (Sigma) at 1:2,500 dilution for 1 hour at 37°C. We washed the plate, added 50 μL of p-nitrophenyl phosphate substrate (Sigma, https://www.sigmaaldrich.com), and measured absorbance at 405 nm using an Epoch plate reader (Biotek, https://www.biotek.com). For analyzing the 547 serum samples from the Cebu cohort using ELISA assays performed over several days, we used human monoclonal antibody ZKA190 (21), which binds to highly accessible regions of ZIKV EDIII as a control to standardize EDIII ELISA optical density (OD) values across assays. Each plate was developed until wells with ZKA190 generated an OD value within a 1.0–1.3 range. If the signal was outside this range, we considered the assay invalid and repeated the process. We divided all OD values for human samples by the ZKA190 OD value on the same plate before determining the ZIKV-immune status of each participant (>0.34 cutoff).
Human Serum Panel Used to Validate the EDIII Assay
To validate the EDIII assay as described elsewhere (19), we used a panel of 142 archived convalescent serum samples from 15 participants who received a licensed Flavivirus (yellow fever/Japanese encephalitis virus) vaccine, 27 serum samples from Flavivirus-naive participants, 33 from participants with immunity to ZIKV (including some with both ZIKV and DENV immunity), and 67 from participants with DENV immunity but no immunity to ZIKV. The DENV- and ZIKV-immunity status of convalescent specimens in the panel was based on participants in febrile illness study cohorts in Nicaragua and Sri Lanka with laboratory-confirmed acute DENV or ZIKV infections or tests for the presence of neutralizing DENV or ZIKV antibodies in single serum samples from healthy persons. We designated samples with no detectable neutralizing DENV and ZIKV antibodies or 50% plaque reduction neutralization test (PRNT50) results <10 as naive. Collection, storage, and use of these convalescent serum samples for research was approved by the Institutional Review Board of the University of North Carolina at Chapel Hill (protocol 08–0895).
Centers for Disease Control and Prevention ZIKV Persistence Samples
The Centers for Disease Control and Prevention (CDC) and the Ponce Medical School Foundation monitored DENV and ZIKV patients seeking treatment at 2 hospitals in southern Puerto Rico. During triage, we identified case-patients with >1 signs or symptoms: fever (temperature >38.0°C or >100.5°F) or reporting fever lasting <7 days, rash, arthralgia or arthritis, or conjunctivitis and offered them participation in the study. We identified ZIKV cases through test results from the CDC Dengue Branch laboratory in San Juan, Puerto Rico, where reverse transcription (RT) PCR testing for DENV, ZIKV, CHIKV, and other respiratory infectious diseases is performed and followed ZIKV cases longitudinally as described elsewhere (22). For our study, we selected and prospectively followed 27 ZIKV RT-PCR–positive cases for up to 2.5 years. Specimen collection was approved by institutional review boards at CDC and Ponce Medical School Foundation.
Pediatric Samples from Cebu Province
We published the Cebu study protocol and approvals elsewhere (23). In brief, our study used baseline serum samples that were collected from 2,996 children 9–14 years of age enrolled in a postlicensure DENV vaccine study. Cohort residence was equally split between Bogo and Balamban, both semiurban areas. We collected samples and demographic information before participants were vaccinated at the same visit.
DENV and ZIKV Focus Reduction Neutralization Test
We determined neutralization titers against DENV and ZIKV by focus-reduction neutralization test (FRNT) in a 96-well format described elsewhere (24). Serially diluted serum was mixed with 50–100 focus-forming units of the virus in Dulbecco modified Eagle medium with 2% fetal bovine serum. We incubated the antibody and virus complexes (1 h, 37°C), then transferred them to a monolayer of Vero-81 cells for infection. After 1 additional hour of incubating antibody and virus complex on Vero-81 monolayer, we overlaid cells with GIBCO Opti-MEM (https://www.thermofisher.com), a modified Eagle medium containing 2% fetal bovine serum and 1% carboxymethylcellulose. After allowing the predetermined time required to form viral foci, we fixed Vero-81 cells and immunostained them with Flavivirus-specific monoclonal antibodies. For neutralization assays, we calculated 50% inhibitory concentration by using the sigmoidal dose-response (variable slope) equation in Prism 6 (GraphPad Software, https://www.graphpad.com). For the study, we included only reported values with an R2 (coefficient of determination) >0.75, hill slope >0.5, and 50% inhibitory concentration within the assay range. We performed FRNT for Flavivirus strains WP74 (DENV-1), S16803 (DENV-2), CH53489 (DENV-3), TVP-376 (DENV-4), and H/PF/2013 (ZIKV).
Depletion of DENV binding Antibodies
We obtained purified viral antigens for antibody depletions by infecting Vero-81 cell cultures in 850 cm2 roller bottles (Greiner Bio-One, https://www.gbo.com) as described elsewhere (25). We conjugated Flavivirus-specific antibody 1M7 to Tosyl-activated Dynabeads (Thermo Fisher) magnetic beads and incubated purified DENV-1–4 viral antigens (1 h at 37°C) with the beads. To deplete DENV-specific antibodies, we incubated serum samples (3 h at 37°C) with DENV-conjugated Dynabeads and incubated serum samples with Dynabeads conjugated to an equivalent bovine serum albumin concentration. We confirmed cross-reactive and DENV antibody depletion using whole virion capture ELISA against DENV-1–4. We measured serotype-specific ZIKV antibodies in serum samples using ZIKV whole-virion capture ELISA after depletion, as described elsewhere (24).
Receiver Operator Characteristic Analysis
We used SPSS Statistics for Macintosh version 27.0 (https://www.ibm.com) to report the performance of the EDIII ELISA based on the receiver operating characteristic (ROC) curve, which presents test performance as true-positive (sensitivity) versus false-positive (1 – specificity). We calculated the optimal assay cutoff value, which maximizes sensitivity and specificity, from the ROC curve using the same software. According to the test, sensitivity is the fraction of total confirmed positive samples with true positives, and specificity is total confirmed negative samples with true negatives.
Development of Immunoassay for Detecting ZIKV Antibodies in Patient Serum
We previously demonstrated that a serologic assay using the ZIKV EDIII fused to maltose-binding protein (MBP) reliably differentiated persons with past ZIKV infections from those with DENV infections (19). However, we observed a high background signal in some specimens, originating from human antibodies binding to MBP or the mouse antigen capture antibody used in the assay. To overcome this problem in the earlier version of the assay, we produced the ZIKV EDIII antigen fused to a HaloTag (Figure 1, panel A), derived from the haloalkane dehalogenase enzyme from Rhodococcus rhodochrous bacteria (26). By using a biotin HaloTag ligand (27), we added a single biotin molecule for each protein molecule at a site distant from the EDIII antigen (Figure 1, panel B). Using streptavidin-coated ELISA plates to capture the antigen, we established an assay for detecting ZIKV EDIII–binding antibodies in human clinical samples (Figure 1, panel C).
Performance of the Second-Generation ZIKV EDIII ELISA
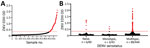
We performed an ROC analysis to determine the diagnostic performance of the new EDIII-capture ELISA in a panel of serum samples collected >12 weeks after symptom onset among persons with laboratory-confirmed ZIKV or DENV infections or who received a licensed Flavivirus vaccine. The panel included serum samples from DENV-naive and DENV-immune participants who received positive ZIKV RT-PCR test results, archived serum samples from persons who had primary (only 1 serotype) or secondary (>2 serotypes) DENV infections, control samples from persons who had not experienced DENV or ZIKV infections, and samples from travelers who had received Japanese encephalitis virus vaccine or yellow fever virus vaccine or both. The EDIII assay was positive for 32 of 33 ZIKV-immune, 7 of 67 DENV-immune, and 2 of 42 DENV- and ZIKV-naive or yellow fever/Japanese encephalitis virus vaccine recipients (Figure 2, panel A). The ROC analysis demonstrated an area under the curve of 0.97 (95% CI of 0.94–0.99) (Figure 2, panel B). At a cutoff value of OD 0.34, the sensitivity of the EDIII-capture ELISA was 97% and the specificity was 92%.
Durability of ZIKV EDIII Antibodies in Persons Exposed to ZIKV Infections
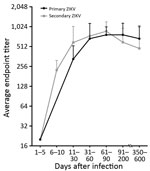
Next, we assessed the durability of EDIII antibodies using a panel of 98 longitudinal samples collected 1–600 days after symptom onset from 27 residents of Puerto Rico with PCR-confirmed ZIKV infections. By testing the acute specimens for DENV-specific antibodies, we stratified the DENV-immune status at the time of ZIKV infection as 12 DENV-naive and 15 DENV-immune cases. We measured endpoint titers at each time point to follow the kinetics of serum ZIKV EDIII antibodies (Figure 3). The ZIKV EDIII antibody titers reached peak levels by 1 month after initial infection and stayed well above detection level for ≥20 months in DENV- naive and DENV-immune participants.
Seroprevalence of ZIKV in Cebu Province
Having established the performance of the ZIKV EDIII ELISA among participants exposed to multiple DENV serotypes, we used the assay to estimate the seroprevalence of ZIKV among children living in Balamban and Bogo City in Cebu Province. In 2017, we collected baseline blood samples from a DENV vaccine study of 2,996 children 9–14 years of age. Elsewhere we reported that 89.3% of the children were DENV-immune at baseline, demonstrating the high endemicity of the virus in this population (23). We selected a representative sample of 547 children on the basis of DENV-immune status from the baseline cohort to determine the seroprevalence of ZIKV (Table). The children selected for the study consisted of 60 (11%) with no immunity to DENV, 43 (8%) with immunity to 1 DENV serotype, and 444 (81%) with immunity to >2 serotypes (Table; Figure 4, panel B) (23). We observed that 98/547 (18%) children were ZIKV antibody positive in the ZIKV EDIII assay (Figure 4, panel A). The number of children who tested positive or negative for ZIKV antibodies did not differ by sex, location, or prior DENV-immune status (Table).
Comparison of ZIKV EDIII ELISA and Virus Neutralizing Antibody Assays
The current standard for estimating the seroprevalence of a particular flavivirus is based on measuring virus neutralizing antibodies in cell culture systems. Although neutralizing antibody assays are more specific than assays that measure antibody binding to flaviviruses or whole recombinant proteins, persons exposed to multiple DENV serotypes can develop ZIKV cross-neutralizing antibodies as reported elsewhere (10,24,28,29). Given the large number of children in our cohort with a history of >2 DENV serotype infections, we compared the performance for estimating ZIKV seroprevalence of the EDIII ELISA using FRNT.
From the 547 baseline samples tested by ZIKV EDIII assay, we tested 495 samples in a single dilution (1:40) ZIKV neutralization assay. We have demonstrated elsewhere that at 1:40 dilution, neutralization of >70% of the input virus is a reliable screening criterion for previous DENV infection (23). When we used the same criterion for the single-dilution ZIKV neutralization assay, we observed a much higher seroprevalence (39%) compared with the estimate (18%) based on the EDIII ELISA (Appendix Table 1). Most of the discordant samples (positive in the ZIKV neutralization assay and negative in the ZIKV EDIII assay) were from children with preexisting multitypic immunity to DENVs (105/110 participants) (Appendix Table 1), raising the possibility that high levels of DENV antibodies cross-neutralize ZIKV.
To evaluate whether the discordant results were because of the poor specificity of the ZIKV neutralization assay or poor sensitivity of the ZIKV EDIII ELISA, we selected 24 samples that had been classified by full curve neutralization testing against the 4 DENV serotypes and ZIKV as primary DENV-immune only (n = 3), primary ZIKV-immune only (n = 4), both primary DENV- and primary ZIKV-immune (n = 6), multitypic DENV-immune only (n = 2), or both multitypic DENV- and ZIKV-immune (n = 9) participants (Appendix Table 2, Figure 2) for further study. By selectively removing all antibodies to DENVs in a sample before testing for ZIKV binding antibodies, it is possible to detect ZIKV type-specific antibodies indicative of a past ZIKV infection (11). We incubated all 24 samples with magnetic beads coated with the 4 DENV serotypes to remove all DENV-binding antibodies. After confirming removal of all DENV-binding antibodies, we tested the samples for ZIKV-binding antibodies. After depleting DENV-binding antibodies, we observed ZIKV-binding antibodies for the participants designated as immune to primary Zika only and primary DENV and primary ZIKV (Appendix Table 2, Figure 1, panel A). In contrast, 6/9 participants designated by neutralization testing as multitypic DENV- and ZIKV-immune showed no ZIKV-binding antibodies (Appendix Figure 1, panel A). All participants that retained ZIKV-binding antibodies after DENV-binding antibody depletion also tested positive in the ZIKV EDIII ELISA and all participants with no detectable ZIKV-specific antibodies tested negative in the EDIII assay (Appendix Table 2, Figure 1, panel B). These results suggest that the ZIKV EDIII ELISA is more reliable than neutralization testing for estimating ZIKV seroprevalence independent of DENV status.
The 2015 ZIKV pandemic in the Americas mostly affected communities with high DENV endemicity because these viruses share the same mosquito vector. Efforts to monitor the spread of ZIKV and the effect of cross-reactive immunity on viral pathogenesis were severely hampered by the inability of conventional serologic assays to accurately distinguish DENV from ZIKV seropositivity. Several groups developed ZIKV recombinant antigen–based assays or antigen-antibody competition assays to improve the specificity of serologic assays (16–18).
In a previous study, we documented development of an ELISA to detect antibodies in persons who had recovered from ZIKV infections, using ZIKV EDIII fused to Escherichia coli MBP as an antigen (19). We observed in some persons high background levels of antibodies to the MBP fusion protein alone, most likely from natural exposure to bacterial proteins, highlighting the need for modifying the fusion protein used for antigen production. We describe a ZIKV EDIII antigen improved by replacing the MBP fusion tag with a HaloTag amenable to site-specific biotinylation. Using streptavidin-biotin chemistry to capture the antigen, we developed an ELISA with 97% sensitivity and 92% specificity for detecting past ZIKV infections.
A strength of our study was the use of serum samples from participants living in Flavivirus-endemic regions with well-defined exposures to DENVs or ZIKV or both to determine the performance of the assay when testing specimens with high levels of cross-reactive antibodies. A potential problem with using modular domains such as EDIII instead of the full-length envelope protein is a loss in assay sensitivity over time. However, our analysis of the longitudinal samples from patients with documented ZIKV infection demonstrated that ZIKV EDIII antibodies robustly developed and remained at high levels even after 2.7 years. This finding suggests that serologic assays based on EDIII will have the necessary sensitivity to characterize past ZIKV circulation at individual and population levels.
The severity of the epidemic in the Americas renewed interest in the epidemiology and pathogenesis of ZIKV in Asia. Asian lineages of the virus have been circulating for decades if not longer. The virus has been detected by molecular methods or virus isolation by cell culture in many countries, including Thailand, the Philippines, Vietnam, Malaysia, Cambodia, and India (30–35). One study of 600 migrant workers in Taiwan (predominantly from Indonesia, the Philippines, Thailand, and Vietnam) reported 37% seroprevalence using a ZIKV IgG assay (36,37). A similar study of Taiwan residents (n = 212) identified 4.2% of all participants with ZIKV IgG but confirmed only 1 participant by neutralization test (36). Investigators tested a cohort of children 1–4 years of age (n = 662) in Indonesia and estimated 9.1% seroprevalence using 90% neutralization as a cutoff value (38). A 2017 study of healthy adults (n = 801) in Vietnam estimated 1.1% seroprevalence using a ZIKV neutralization assay (39). An additional study in Guangxi Province, China, in March 2019 found 6% of 273 participants bound to ZIKV NS1 using IgG ELISA testing and neutralized ZIKV in cell cultures (40). Those past studies, especially those relying on tests of whole Zika virions and full-length antigens, had poor specificity, mainly because of antibodies induced by DENV infections cross-reacting with ZIKV antigens. In our study, we used a recombinant ZIKV antigen with a high specificity for distinguishing ZIKV-induced from DENV-induced antibodies; results showed that 18% of children 9–14 years of age living in Cebu Province had previously experienced a ZIKV infection. Although ZIKV is currently not considered by public health agencies to be a common infection in the Philippines, our results indicate otherwise.
Our results also highlight a problem with using accepted standards for neutralizing antibody testing to identify ZIKV infections in populations heavily exposed to DENVs. We observed ZIKV neutralizing antibodies in 39% of the children in this study, which was more than double the estimate based on the EDIII assay results. By depleting all DENV-binding antibodies from a subset of samples and then measuring ZIKV antibodies, we demonstrated that some children exposed to >2 DENV serotypes have low levels of ZIKV cross-neutralizing antibodies that can lead to false-positive results. In contrast, the ZIKV EDIII assay was not impacted by DENV-immune status because, even after removing all DENV-binding antibodies, the children maintained ZIKV EDIII–binding antibodies.
The novel assay we describe, which uses a biotinylated EDIII antigen to detect ZIKV type-specific antibodies, is simple and readily adaptable for use in standard clinical and public health laboratories to monitor ZIKV transmission at both the individual and population levels. Given that the 2015 ZIKV epidemic spread to >40 countries and infected more than a million people, many in regions with endemic transmission of DENVs, and that DENVs demonstrate cross-reactive immunity with ZIKV, the improved specificity of this assay provides a necessary resource for accurately monitoring ZIKV transmission in regions with high DENV endemicity.
Mr. Adams is an MD/PhD student at the University of North Carolina School of Medicine. His main research and clinical interests involve global health, infectious diseases, and vaccinology.
Acknowledgments
We thank the children and their families in Balamban and Bogo City who volunteered for this study and the field staff who assisted with the study. We also thank Laura White and Alena Markmann for providing specimens for validating the assay and for supervising sample processing, inventory, and storage and Matt Collins for providing clinical samples for validating the assay.
The field component of the study in the Philippines was supported by the Department of Health (Philippines), the Hanako Foundation, the World Health Organization, and the Swedish International Development Cooperation Agency in partnership with the International Vaccine Institute in Seoul, South Korea. The laboratory studies at the University of North Carolina were supported by grants from the US Centers for Disease Control and Prevention (BAA 2017-N-18041; PI AdeS) and the US National Institutes of Health (U24 AI152170-01 and R21 AI134073 PIs: Ades and LP). C.A. was supported by NIH Predoctoral Training Grant T32 AI007419: “Molecular Biology of Viral Disease.” US patent application no. 20200300855 on the method incorporating the EDIII antigen for serodiagnosis of Zika infection has been filed under the title “Methods and Compositions for Zika Virus Detection.”
References
- Hills SL, Fischer M, Petersen LR. Epidemiology of Zika virus infection. J Infect Dis. 2017;216(suppl_10):S868–74. DOIPubMedGoogle Scholar
- Ventura CV, Maia M, Bravo-Filho V, Góis AL, Belfort R Jr. Zika virus in Brazil and macular atrophy in a child with microcephaly. Lancet. 2016;387:228. DOIPubMedGoogle Scholar
- Johansson MA, Mier-y-Teran-Romero L, Reefhuis J, Gilboa SM, Hills SL. Zika and the risk of microcephaly. N Engl J Med. 2016;375:1–4. DOIPubMedGoogle Scholar
- Hoen B, Schaub B, Funk AL, Ardillon V, Boullard M, Cabié A, et al. Pregnancy outcomes after ZIKV infection in French territories in the Americas. N Engl J Med. 2018;378:985–94. DOIPubMedGoogle Scholar
- Borchering RK, Huang AT, Mier-Y-Teran-Romero L, Rojas DP, Rodriguez-Barraquer I, Katzelnick LC, et al. Impacts of Zika emergence in Latin America on endemic dengue transmission. Nat Commun. 2019;10:5730. DOIPubMedGoogle Scholar
- Gordon A, Gresh L, Ojeda S, Katzelnick LC, Sanchez N, Mercado JC, et al. Prior dengue virus infection and risk of Zika: A pediatric cohort in Nicaragua. PLoS Med. 2019;16:
e1002726 . DOIPubMedGoogle Scholar - Katzelnick LC, Narvaez C, Arguello S, Lopez Mercado B, Collado D, Ampie O, et al. Zika virus infection enhances future risk of severe dengue disease. Science. 2020;369:1123–8. DOIPubMedGoogle Scholar
- Duong V, Dussart P, Buchy P. Zika virus in Asia. Int J Infect Dis. 2017;54:121–8. DOIPubMedGoogle Scholar
- Hu T, Li J, Carr MJ, Duchêne S, Shi W. The Asian lineage of Zika virus: transmission and evolution in Asia and the Americas. Virol Sin. 2019;34:1–8. DOIPubMedGoogle Scholar
- Priyamvada L, Quicke KM, Hudson WH, Onlamoon N, Sewatanon J, Edupuganti S, et al. Human antibody responses after dengue virus infection are highly cross-reactive to Zika virus. Proc Natl Acad Sci U S A. 2016;113:7852–7. DOIPubMedGoogle Scholar
- Andrade P, Gimblet-Ochieng C, Modirian F, Collins M, Cárdenas M, Katzelnick LC, et al. Impact of pre-existing dengue immunity on human antibody and memory B cell responses to Zika. Nat Commun. 2019;10:938. DOIPubMedGoogle Scholar
- Rogers TF, Goodwin EC, Briney B, Sok D, Beutler N, Strubel A, et al. Zika virus activates de novo and cross-reactive memory B cell responses in dengue-experienced donors. Sci Immunol. 2017;2:
eaan6809 . DOIPubMedGoogle Scholar - Chua CL, Chan YF, Andu ESGS, Rovie-Ryan JJ, Sitam FT, Verasahib K, et al. Little evidence of Zika virus infection in wild long-tailed Macaques, peninsular Malaysia. Emerg Infect Dis. 2019;25:374–6. DOIPubMedGoogle Scholar
- Beck C, Leparc-Goffart I, Desoutter D, Debergé E, Bichet H, Lowenski S, et al. Serological evidence of infection with dengue and Zika viruses in horses on French Pacific Islands. PLoS Negl Trop Dis. 2019;13:
e0007162 . DOIPubMedGoogle Scholar - Netto EM, Moreira-Soto A, Pedroso C, Höser C, Funk S, Kucharski AJ, et al. High Zika virus seroprevalence in Salvador, Northeastern Brazil limits the potential for further outbreaks. MBio. 2017;8:e01390–17. DOIPubMedGoogle Scholar
- Balmaseda A, Stettler K, Medialdea-Carrera R, Collado D, Jin X, Zambrana JV, et al. Antibody-based assay discriminates Zika virus infection from other flaviviruses. Proc Natl Acad Sci U S A. 2017;114:8384–9. DOIPubMedGoogle Scholar
- Rockstroh A, Moges B, Barzon L, Sinigaglia A, Palù G, Kumbukgolla W, et al. Specific detection of dengue and Zika virus antibodies using envelope proteins with mutations in the conserved fusion loop. Emerg Microbes Infect. 2017;6:
e99 . DOIPubMedGoogle Scholar - Denis J, Attoumani S, Gravier P, Tenebray B, Garnier A, Briolant S, et al. High specificity and sensitivity of Zika EDIII-based ELISA diagnosis highlighted by a large human reference panel. PLoS Negl Trop Dis. 2019;13:
e0007747 . DOIPubMedGoogle Scholar - Premkumar L, Collins M, Graham S, Liou GA, Lopez CA, Jadi R, et al. Development of envelope protein antigens to serologically differentiate Zika virus infection from dengue virus infection. J Clin Microbiol. 2018;56:e01504–17. DOIPubMedGoogle Scholar
- Los GV, Encell LP, McDougall MG, Hartzell DD, Karassina N, Zimprich C, et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem Biol. 2008;3:373–82. DOIPubMedGoogle Scholar
- Wang J, Bardelli M, Espinosa DA, Pedotti M, Ng TS, Bianchi S, et al. A human bi-specific antibody against Zika virus with high therapeutic potential. Cell. 2017;171:229–241.e15. DOIPubMedGoogle Scholar
- Paz-Bailey G, Rosenberg ES, Sharp TM. Persistence of Zika virus in body fluids—final report. N Engl J Med. 2019;380:198–9. DOIPubMedGoogle Scholar
- Lopez AL, Adams C, Ylade M, Jadi R, Daag JV, Molloy CT, et al. Determining dengue virus serostatus by indirect IgG ELISA compared with focus reduction neutralisation test in children in Cebu, Philippines: a prospective population-based study. Lancet Glob Health. 2021;9:e44–51. DOIPubMedGoogle Scholar
- Collins MH, McGowan E, Jadi R, Young E, Lopez CA, Baric RS, et al. Lack of durable cross-neutralizing antibodies against Zika virus from dengue virus infection. Emerg Infect Dis. 2017;23:773–81. DOIPubMedGoogle Scholar
- Putnak R, Barvir DA, Burrous JM, Dubois DR, D’Andrea VM, Hoke CH, et al. Development of a purified, inactivated, dengue-2 virus vaccine prototype in Vero cells: immunogenicity and protection in mice and rhesus monkeys. J Infect Dis. 1996;174:1176–84. DOIPubMedGoogle Scholar
- Benink HA, Urh M. HaloTag technology for specific and covalent labeling of fusion proteins. Methods Mol Biol. 2015;1266:119–28. DOIPubMedGoogle Scholar
- Grainger DW, Castner DG, Dubey M, Emoto K, Takahashi H. Affinity-based protein surface pattern formation by ligand self-selection from mixed protein solutions. Adv Funct Mater. 2009;19:3046–55. DOIPubMedGoogle Scholar
- Dussupt V, Sankhala RS, Gromowski GD, Donofrio G, De La Barrera RA, Larocca RA, et al. Potent Zika and dengue cross-neutralizing antibodies induced by Zika vaccination in a dengue-experienced donor. Nat Med. 2020;26:228–35. DOIPubMedGoogle Scholar
- Fernandez E, Dejnirattisai W, Cao B, Scheaffer SM, Supasa P, Wongwiwat W, et al. Human antibodies to the dengue virus E-dimer epitope have therapeutic activity against Zika virus infection. [Erratum in Nat Immunol. 2020;21:354.]. Nat Immunol. 2017;18:1261–9. DOIPubMedGoogle Scholar
- Woon YL, Lim MF, Tg Abd Rashid TR, Thayan R, Chidambaram SK, Syed Abdul Rahim SS, et al. Zika virus infection in Malaysia: an epidemiological, clinical and virological analysis. BMC Infect Dis. 2019;19:152. DOIPubMedGoogle Scholar
- Ho ZJM, Hapuarachchi HC, Barkham T, Chow A, Ng LC, Lee JMV, et al.; Singapore Zika Study Group. Outbreak of Zika virus infection in Singapore: an epidemiological, entomological, virological, and clinical analysis. Lancet Infect Dis. 2017;17:813–21. DOIPubMedGoogle Scholar
- Buathong R, Hermann L, Thaisomboonsuk B, Rutvisuttinunt W, Klungthong C, Chinnawirotpisan P, et al. Detection of Zika virus infection in Thailand, 2012–2014. Am J Trop Med Hyg. 2015;93:380–3. DOIPubMedGoogle Scholar
- Ruchusatsawat K, Wongjaroen P, Posanacharoen A, Rodriguez-Barraquer I, Sangkitporn S, Cummings DAT, et al. Long-term circulation of Zika virus in Thailand: an observational study. [Erratum in Lancet Infect Dis. 2019;19:e109.]. Lancet Infect Dis. 2019;19:439–46. DOIPubMedGoogle Scholar
- Alera MT, Hermann L, Tac-An IA, Klungthong C, Rutvisuttinunt W, Manasatienkij W, et al. Zika virus infection, Philippines, 2012. Emerg Infect Dis. 2015;21:722–4. DOIPubMedGoogle Scholar
- Buerano CC, Pangilinan LS, Dimamay MTA, Mapua CA, Dimamay MPS, Matias RR, et al. Zika virus infection, Philippines, 2012. Emerg Infect Dis. 2020;26:2300–1. DOIPubMedGoogle Scholar
- Chien YW, Ho TC, Huang PW, Ko NY, Ko WC, Perng GC. Low seroprevalence of Zika virus infection among adults in Southern Taiwan. BMC Infect Dis. 2019;19:884. DOIPubMedGoogle Scholar
- Perng GC, Ho TC, Shih HI, Lee CH, Huang PW, Chung CH, et al. Seroprevalence of Zika and dengue virus antibodies among migrant workers, Taiwan, 2017. Emerg Infect Dis. 2019;25:814–6. DOIPubMedGoogle Scholar
- Sasmono RT, Dhenni R, Yohan B, Pronyk P, Hadinegoro SR, Soepardi EJ, et al. Zika virus seropositivity in 1–4-year-old children, Indonesia, 2014. Emerg Infect Dis. 2018;24:1740–3. DOIPubMedGoogle Scholar
- Nguyen CT, Moi ML, Le TQM, Nguyen TTT, Vu TBH, Nguyen HT, et al. Prevalence of Zika virus neutralizing antibodies in healthy adults in Vietnam during and after the Zika virus epidemic season: a longitudinal population-based survey. BMC Infect Dis. 2020;20:332. DOIPubMedGoogle Scholar
- Zhou CM, Liu JW, Qi R, Fang LZ, Qin XR, Han HJ, et al. Emergence of Zika virus infection in China. PLoS Negl Trop Dis. 2020;14:
e0008300 . DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: November 10, 2021
1Deceased.
Table of Contents – Volume 27, Number 12—December 2021
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Aravinda de Silva or Lakshmanane Premkumar, Department of Microbiology and Immunology, University of North Carolina School of Medicine, Chapel Hill, NC 27599-7292, USA
Top