Volume 28, Number 10—October 2022
Dispatch
Endofungal Mycetohabitans rhizoxinica Bacteremia Associated with Rhizopus microsporus Respiratory Tract Infection
Figure 2
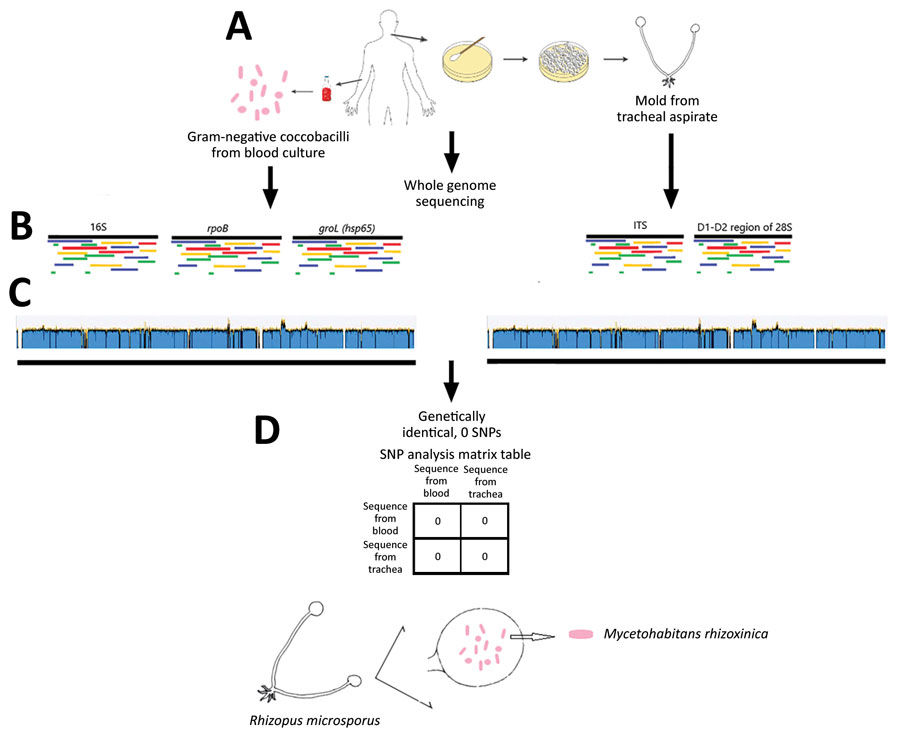
Figure 2. Whole-genome sequencing analysis of bacterial and fungal isolates (A) in a 65-year-old woman with multiple myeloma undergoing chimeric antigen receptor T-cell therapy, California, USA. The bacteria were identified as Mycetohabitans rhizoxinica with >99% identity in all 3 full-length marker genes compared with the reference organism (B, left side). The mold was identified as Rhizopus microsporus with >99% identity in the ITS gene and the D1–D2 region of the 28S gene compared with a reference organism (B, right side). Whole-genome sequences of the bacteria from blood (C, left side) revealed whole-genome coverage 94.0% and pairwise identity 95.5% with sufficient mean coverage of 298×. Whole-genome sequence of the mold from tracheal aspirate (C, right side) aligned to reference bacterial whole-genome sequence showed whole-genome coverage 94.1% and pairwise identity 95.8%, with sufficient mean coverage of 75×. The bacteria inside the mold from the trachea were genetically identical to the bacteria from the blood, as shown by the SNP analysis (D). ITS, internal transcribed spacer; SNP, single nucleotide polymorphism.