Volume 28, Number 12—December 2022
Dispatch
Natural Mediterranean Spotted Fever Foci, Qingdao, China
Cite This Article
Citation for Media
Abstract
We sequenced DNA from spleens of rodents captured in rural areas of Qingdao, East China, during 2013–2015. We found 1 Apodemus agrarius mouse infected with Rickettsia conorii, indicating a natural Mediterranean spotted fever foci exists in East China and that the range of R. conorii could be expanding.
Mediterranean spotted fever (MSF) is an acute febrile, zoonotic disease caused by the bacterium Rickettsia conorii that is transmitted to humans by the brown dog tick, Rhipicephalus sanguineus (1). MSF was described from the Mediterranean region in 1910 (2); a similar disease, known as Indian tick typhus (ITT), was described in 1925 (3). The causative agent of ITT was later confirmed to be R. conorii (4). Since 1990, the natural foci of MSF has continued to expand into the Middle East, Africa, and central Europe (5). R. conorii ITT has been detected in ticks in Xinjiang Province in West China (6), and an MSF case recently was reported in Shandong Province in East China (7).
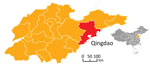
The city of Qingdao, located in the southeast part of Shandong Province, is on the pacific coast of East China (Figure 1). Qingdao has a temperate monsoonal climate that is ideal for rodent propagation, and many natural-focal diseases caused by Rickettsia spp., Orientia tsutsugamushi, and severe fever with thrombocytopenia syndrome virus (8–10). We used PCR amplification to investigate whether rodents in the region are infected with Rickettsia spp. and unexpectedly discovered R. conorii.
We performed a retrospective study by testing rodents captured from Huangdao District, Qingdao, China, during July–October every year from 2013–2015. The rodent collections were previously described (11). We aseptically collected rodent spleens and stored at −80°C. Animal use and sample collection were approved by the ethics committee of Medical School, Shandong University (approval no. 20150501), and performed in accordance with Shandong University Guidelines on the Care and Use of Laboratory Animals.
We extracted DNA from homogenized rodent spleen tissues by using the QIAamp DNA Mini Kit (QIAGEN, https://www.qiagen.com). We initially performed nested PCR on all rodents with 17-kDa antigen (htrA), then we further tested the PCR-positive samples by using primers of 16S ribosomal RNA (rrs) and outer membrane protein A and B (ompA and ompB) genes (12,13) (Table 1). We used nuclease-free water as a negative control in each experiment. We performed DNA extraction, PCR amplification, and PCR product analysis in separate rooms to avoid false-positive results. We visualized PCR products in 1.0%–1.5% agarose gels based on the length of amplified DNA segments. We excised and extracted expected DNA bands by using a gel extraction kit (Tsingke Biotech, https://tsingke.com). We cloned purified PCR products into T-Vector pMD19 (TaKaRa Bio, Inc., https://www.takara-bio.com). Both strands were sequenced by Sangon Biotech (https://www.sangon.com).
We edited DNA sequences by using DNAStar software (https://www.dnastar.com) to remove primers and analyzed sequences in BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to compare with GenBank sequences. We constructed a phylogenetic tree by using the maximum-likelihood method with the Kimura 2-parameter model in MEGA version 7 (https://www.megasoftware.net), and we calculated bootstrap values with 1,000 replicates to determine the relative support for clades in the trees.
We used a total of 121 rodents in this study, including 60 striped field mice (Apodemus agrarius), 19 house mice (Mus musculus), 16 Chinese hamsters (Cricetulus barabensis), 10 brown rats (Rattus norvegicus), 8 greater long-tailed hamsters (Cricetulus triton), and 8 Chinese white-bellied rats (Niviventer confucianus) (Table 2). Among the rodents, 81.82% (99/121) were captured outdoors and 18.18% (22/121) indoors.
PCR amplification indicated that 1 A. agrarius mouse captured outdoors was positive for a Rickettsia species and the other 120 rodents were negative for Rickettsia by htrA primers. We further amplified the spleen tissue of the PCR-positive mouse by using rrs, ompA, and ompB primers. All 3 pairs of primers generated positive PCR products. Because the PCR fragment was too short and the sequence was conserved, we could not differentiate Rickettsia species by phylogenetic analysis of the htrA gene. However, the rrs gene sequence obtained in this study was 100% identical to R. conorii strains in GenBank (1,185/1,185 bp for accession no. KU364355, 1,267/1,267 bp for accession no. KY069267) that were obtained from Rh. turanicus ticks in Xinjiang, China. The obtained rrs sequence also matched 100% (1,331/1,331 bp) with a GenBank R. conorii strain from India (accession no. L36107), and an R. conorii Malish 7 strain (accession no. NR_041934). The ompA sequence from the mouse was 100% identical to R. conorii strains from Rh. turanicus ticks (494/494 bp for GenBank accession nos. MF002512 and KY069258) and an R. conorii strain from a human blood sample (449/449 bp for accession no. MG190328) from Xinjiang, China. The obtained ompA sequence was also 100% (494/494 bp) identical to GenBank R. conorii strains from India (accession no. U43794) and Italy (accession no. JN944636). The ompB amplified from the mouse was 99.87% (798/799 bp) homologous to the corresponding sequences of R. conorii from Rh. turanicus ticks from Xinjiang, China (GenBank accession nos. MF002514 and KY069249) and an R. conorii strain from India (GenBank accession no. AF123726).
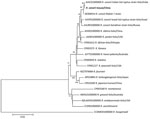
A phylogenetic tree of the concatenated sequences of the 4 genes, rrs (1,331 bp), htrA (169 bp), ompA (494 bp), and ompB (844 bp), showed that the rickettsial sequence from this study clustered with R. conorii ITT from ticks in India (GenBank accession no. AJHC01000000) and within the same clade as the R. conorii Malish 7 strain (GenBank accession no. AE006914), R. conorii Israel tick typhus strain (GenBank accession no. AJVP01000000), and R. conorii Astrakhan strain (GenBank accession no. AJUR01000000) (Figure 2). We deposited nucleotide sequence data from this study into GenBank (accession no. OM230141 for rrs, OM234678 for htrA, OM234679 for ompA, and OM234680 for ompB genes).
We identified Rickettsia spp. in a striped field mouse captured in Qingdao in East China. Phylogenetic analysis showed that the Rickettsia species we detected was identical in multiple gene sequences to R. conorii ITT, indicating the strain we identified is R. conorii. Our results could not be caused by PCR contamination because we do not have an R. conorii strain nor its DNA in our laboratory.
The prevailing vector of R. conorii is the brown dog tick, Rh. sanguineus (2). The widespread distribution of R. conorii might be related to the worldwide spread of its vector tick among dogs (14). R. conorii ITT sequences have been reported in Rh. turanicus ticks from Xinjiang Province in West China (6). We identified R. conorii in 1 rodent in Qingdao, located in the eastern part of Shandong Province. Another recent study reported R. conorii genomic sequences in a patient in the western part of Shandong Province (7). These results demonstrate that the endemic area of R. conorii either recently expanded into Shandong Province in East China or R. conorii has existed in East China but was not detected before.
We speculate that the expansion of MSF foci in China is caused by transportation of dogs from West China to East China (15), contributing to the spread of brown dog ticks and, thus, R. conorii. The tick vector of R. conorii in Shandong Province has not been identified.
In conclusion, we confirmed R. conorii infection in 1 rodent from Qingdao in East China. Further studies are needed to determine the epidemiology of R. conorii in East China. Our study increases our knowledge about the distribution of R. conorii. Identification of MSF foci in East China could indicate that the range of R. conorii and its tick vector are expanding.
Ms. Gu is a PhD candidate at the School of Public Health, Wuhan University, Wuhan, Hubei, China. Her research interests include emerging infectious disease and vector-borne disease.
X.-J.Y. and H.-J.H. organized and designed the study. J.-T.C., X.-L.G., C.-M.Z., R.W., Q.-M.P., Z.-Z.J., B.L., and W.-K.Z. performed the fieldwork. X.-L.G. and R.W. performed laboratory analysis of pathogens. X.-L.G. analyzed the data and drafted the manuscript. X.-J.Y. critically reviewed the manuscript. All authors read and approved the final manuscript.
Acknowledgments
We thank Laixi City Center for Disease Control and Prevention and Qingdao City Center for Disease Control and Prevention for organizing the fieldwork to trap rodents.
This study was supported by National Natural Science Funds of China (grant no. 81971939).
References
- Colomba C, Saporito L, Polara VF, Rubino R, Titone L. Mediterranean spotted fever: clinical and laboratory characteristics of 415 Sicilian children. BMC Infect Dis. 2006;6:60. DOIPubMedGoogle Scholar
- Rovery C, Brouqui P, Raoult D. Questions on Mediterranean spotted fever a century after its discovery. Emerg Infect Dis. 2008;14:1360–7. DOIPubMedGoogle Scholar
- Sentausa E, El Karkouri K, Robert C, Raoult D, Fournier PE. Genome sequence of Rickettsia conorii subsp. indica, the agent of Indian tick typhus. J Bacteriol. 2012;194:3288–9. DOIPubMedGoogle Scholar
- MacConnachie K, Tishkowski K. Boutonneuse fever. Treasure Island (FL): StatPearls Publishing; 2022 [cited 2022 Jul 5]. https://www.ncbi.nlm.nih.gov/books/NBK560914
- Guo LP, Jiang SH, Liu D, Wang SW, Chen CF, Wang YZ. Emerging spotted fever group rickettsiae in ticks, northwestern China. Ticks Tick Borne Dis. 2016;7:1146–50. DOIPubMedGoogle Scholar
- Xu N, Gai W, Zhang Y, Wang W, Wang G, Dasch GA, et al. Confirmation of Rickettsia conorii subspecies indica infection by next-generation sequencing, Shandong, China. Emerg Infect Dis. 2021;27:2691–4. DOIPubMedGoogle Scholar
- Qin XR, Han HJ, Han FJ, Zhao FM, Zhang ZT, Xue ZF, et al. Rickettsia japonica and novel Rickettsia species in ticks, China. Emerg Infect Dis. 2019;25:992–5. DOIPubMedGoogle Scholar
- Li F, Zhang ZT, Fang LZ, Yu H, Qin XR, Yu XJ. Indoor and outdoor rodent hosts of Orientia tsutsugamushi, Shandong Province, China. Emerg Infect Dis. 2021;27:2731–4. DOIPubMedGoogle Scholar
- Liu JW, Wen HL, Fang LZ, Zhang ZT, He ST, Xue ZF, et al. Prevalence of SFTSV among Asian house shrews and rodents, China, January-August 2013. Emerg Infect Dis. 2014;20:2126–8. DOIPubMedGoogle Scholar
- Qin XR, Liu JW, Yu H, Yu XJ. Bartonella species detected in rodents from eastern China. Vector Borne Zoonotic Dis. 2019;19:810–4. DOIPubMedGoogle Scholar
- Huang Y, Zhao L, Zhang Z, Liu M, Xue Z, Ma D, et al. Detection of a novel Rickettsia from Leptotrombidium scutellare mites (Acari: Trombiculidae) from Shandong of China. J Med Entomol. 2017;54:544–9. DOIPubMedGoogle Scholar
- Yuan TT, Du CH, Xia LY, Que TC, von Fricken ME, Jiang BG, et al. Molecular evidence of Candidatus Rickettsia longicornii and a novel Rickettsia strain from ticks in Southern China. Ticks Tick Borne Dis. 2021;12:
101679 . DOIPubMedGoogle Scholar - Gray J, Dantas-Torres F, Estrada-Peña A, Levin M. Systematics and ecology of the brown dog tick, Rhipicephalus sanguineus. Ticks Tick Borne Dis. 2013;4:171–80. DOIPubMedGoogle Scholar
- Chen J, Zou L, Jin Z, Ruan S. Modeling the geographic spread of rabies in China. PLoS Negl Trop Dis. 2015;9:
e0003772 . DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: November 15, 2022
Table of Contents – Volume 28, Number 12—December 2022
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Hui-Ju Han or Xue-Jie Yu, Wuhan University, School of Public Health, Wuhan, 430071, Hubei Province, China
Top