Volume 28, Number 12—December 2022
Dispatch
Mass Mortality Caused by Highly Pathogenic Influenza A(H5N1) Virus in Sandwich Terns, the Netherlands, 2022
Cite This Article
Citation for Media
Abstract
We collected data on mass mortality in Sandwich terns (Thalasseus sandvicensis) during the 2022 breeding season in the Netherlands. Mortality was associated with at least 2 variants of highly pathogenic avian influenza A(H5N1) virus clade 2.3.4.4b. We report on carcass removal efforts relative to survival in colonies. Mitigation strategies urgently require structured research.
The 2021–2022 epidemic of highly pathogenic avian influenza (HPAI) A(H5N1) virus clade 2.3.4.4b has been unprecedented in terms of numbers of dead wild birds, species affected, spatial extent, and incidence in spring 2022 (1). Across Europe, multiple colony-breeding seabirds experienced HPAI H5N1–associated mass mortalities during the breeding period, including the Sandwich tern (Thalasseus sandvicensis) (1,2). The Netherlands constitutes a major though vulnerable stronghold of the Sandwich tern in Europe; 15,000–20,000 breeding pairs have been documented across ≈10 colonies (https://stats.sovon.nl/stats/soort/6110). We sought to establish the scale of mortality occurring in Sandwich terns breeding in the Netherlands in 2022, characterize the associated HPAI H5N1 viruses and pathology, report on the carcass removal effort relative to survival, and investigate intracolony transmission dynamics.
We determined breeding colony locations and initial sizes in May 2022 through drone or ground counts (3). To establish breeding success and minimum estimates of mortality, we compiled data from late May through early July 2022 on numbers of live adults, chicks, fledglings, and late clutches in colonies, as well as on numbers of carcasses found in and around colonies, carcasses removed for destruction, and abandoned nests (Appendix 1 Table 1). Sandwich terns lay 1–2 eggs and incubate for 21–29 days. Chicks fledge 25–30 days after hatching; annual fledging success is ≈0.5 per breeding pair (4–6). We used wild bird mortality databases to establish minimum estimates of adult mortality outside the colonies. Finally, we used data from the migration tracking website Trektellen (https://www.trektellen.nl) to compare the hourly averages of Sandwich tern passing rates at coastal observation points per week in 2022 to 2016–2021.
We observed clinical signs and tested 44 carcasses for avian influenza virus by using a quantitative PCR to detect the influenza A virus matrix gene; we then followed up with subtype-specific PCRs on cloacal and tracheal swab specimens (7). We performed necropsy with histopathology and immunohistochemistry on 6 of the carcasses to establish cause of death. To study the relationship between viruses detected in Sandwich terns and other bird species, we determined full-genome sequences directly on swab RNA from 20 birds and submitted them to the GISAID database (https://www.gisaid.org), then compared them to a sample of 57 other birds (Appendix 1 Table 2). We aligned sequences by using MAFFT version 7.475 (8), reconstructed phylogeny by using maximum-likelihood analysis with IQ-TREE software version 2.0.3 (9), and visualized the maximum-likelihood tree by using the R package ggtree (10).
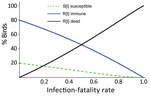
To investigate HPAI H5N1 transmission dynamics within a breeding colony, we developed a susceptible-infectious-recovered model that included infection-fatality rate (IFR) and examined outcome as a function of IFR. IFR is the probability that a bird dies after infection, although in this study IFR also includes birds leaving the colony. The model assumed a naive starting population, an infectious period of 5 days, and frequency-dependent transmission and considered only survivors of infection as recovered and total population size as not constant (Appendix 2).
Mass mortality was seen in 9 of the 10 Sandwich tern breeding colonies in 2022. In those colonies, out of a total of 18,151 breeding pairs, 8,001 adult Sandwich terns were found dead, and only a few chicks fledged (Video 1; Video 2). Only 1 small inland colony of 137 breeding pairs experienced no mass mortality and had a fledgling success rate (0.47 young/pair) consistent with previous years (0.50 young/pair) (Table; Figure 1, panel A). Outside of colonies, another 1,600 adult Sandwich terns were reported dead between late May and end of June. The scale of mortality is reflected in the passage rate of Sandwich terns along the coast in May–June 2022 (Appendix 2 Figure 4).
Diseased birds were debilitated, unable to fly, and mostly lethargic, sometimes with wings spread out. At later stages, some displayed opisthotonos and occasionally flipped over backwards (Video 1, Video 3; Appendix 2 Figures 5–10). We confirmed HPAI H5N1 virus infection in 23 of 24 dead Sandwich terns from colonies (the exception was a chick); infection was also confirmed in 20 of 20 birds outside of colonies (Appendix 2 Table 1). In the 4 necropsied PCR-confirmed adult birds, viral antigen expression was detected by immunohistochemistry in the pancreas (n = 3), duodenum (n = 4), or lung and nasal tissue (n = 1), colocalized with necrosis and inflammation (Appendix 2 Figure 18). Necropsy findings and negative immunohistochemistry results in the 2 chicks we examined demonstrated that chick mortality was at least partly caused by starvation, likely after feeding was interrupted because of adult mortality (Appendix 2). This explanation was supported by field observations (Appendix 2 Figure 19).
Phylogenetic analysis demonstrated that the 20 fully sequenced viruses belonged to H5 clade 2.3.4.4b, and clustered with viruses detected in other wild bird species in the Netherlands, including in geese and gulls collected during January–April 2022 (Figure 1, panel B; Appendix 2). The Sandwich tern viruses clustered in 2 groups. Both variants were found in the northern and southern parts of the Netherlands and were even found within a single colony. These results suggest that at least 2 independent virus introductions into Sandwich terns occurred in the Netherlands, followed by transmission of both virus variants within and between breeding colonies.
Carcass removal effort was diverse (Table; Appendix 1 Table 1). In colonies with some survival, effort was overall more regular, frequent, and immediate or included chicks also, although survival might also have been affected by other undetermined factors.
In most colonies, a high proportion of the birds died or left (Table), indicating high IFR. The model shows that, even with a relatively low value of R0 (R0 = 2), at the higher end of the IFR range evidenced here, few of the birds remaining in the colony will have escaped infection at the end of the outbreak. Birds in the colony will have died or recovered and acquired immunity (Figure 2).
Our results substantiate that after Sandwich terns arrived in the Netherlands for breeding, HPAI H5N1 virus was introduced into their population at least twice. The virus then spread widely within and between breeding colonies, causing outbreaks that resulted in high adult and chick mortality in nearly all colonies. Infected birds probably died of systemic HPAI-associated disease, including acute pancreatic necrosis and duodenitis (11,12). Like other seabirds, Sandwich terns have low annual reproductive output but relatively long life-expectancy (2,4,6,13); therefore the effect of high adult mortality on population size could be seen for a long time. The Sandwich tern exemplifies how severely the continued circulation of HPAI H5N1 viruses in spring 2022 affected populations of colony-breeding birds without flock immunity in Europe (1).
Our study also demonstrates how outbreaks in breeding birds boosted virus propagation into the summer of 2022. The future involvement of Sandwich terns in HPAI endemicity can be evaluated once future population size and flock immunity have been analyzed from count data and serosurveillance. On the basis of our model, colony survivors would be mostly immune to HPAI.
Confirming HPAI as a major mortality factor in breeding colonies of Sandwich terns and other seabird species (2,14,15) underlines the paradigm shift to HPAI as a mortality factor of concern to wild species, in addition to poultry and humans. It stresses the importance of close international cooperation and data exchange to better understand and mitigate the global effect of HPAI on nature. More structured research on appropriate strategies to reduce massive propagation is urgently required. Carcass removal takes away a source of infection but might simultaneously enhance spread of infection and thus requires controlled study.
Dr. Rijks is a postdoctoral researcher at the Dutch Wildlife Health Centre in Utrecht, the Netherlands. Her primary research interests are wildlife diseases and epidemiology. Dr. Leopold is a senior seabird ecologist at Wageningen Marine Research. He has studied Sandwich terns on Texel since 2012.
Acknowledgments
We thank Jitske Esselaar, Arie Baas, Jan Veen, Pim Wolf, Ruben Fijn, Rob van Bemmelen, Bob Loos, Eckard Boot, Marc Plomp, Jan van Dijk, Johan Bremer, Toon Pop, Vincent Stork, Irene Oerlemans, Henk van der Jeugd, Jeroen Nienhuis, Natuurmonumenten, Staatsbosbeheer, and Stichting het Zeeuws Landschap for their assistance with access to colonies and bruto data collection and cleaning; Waarneming.nl and trektellen.nl for additional data; Wilma Booij, Marc Plomp, Eckard Boot, Bureau Waardenburg and Het Zeeuws Landschap for additional photos and videos; Reina Sikkema, Oanh Vuong and Sanne Thewessen for technical assistance with virological analysis; and Judith van der Brand and Evelien Germeraard for their contribution to the pathological analyses. We thank the authors and submitting laboratories of the sequences from the GISAID EpiFlu Database (Appendix 1 Table 2).
This work was partly funded by the Dutch Ministry of Agriculture, Nature, and Food Quality (project no. WOT-01-003-012).
Field data were generated and compiled by M.F.L., S.K., R.V., F.S., S.J.L., M.Z.B., L.K., J.W.J., W.C., E.K., R.S., and J.M.R. Virological and phylogenetic analyses were performed by N.B., M.E., and R.A.M.F. Pathology and immunohistochemistry were conducted and analyzed by S.V., M.L.K., A.G., and T.K. The epidemiological model was made by M.C.M.J. The study was coordinated and summarized by J.M.R. and M.F.L. All authors provided input on draft versions and read and agreed to the final version of the manuscript.
References
- European Food Safety Authority. European Centre for Disease Prevention and Control, European Reference Laboratory for Avian Influenza, Adlhoch C, Fusaro A, Gonzales JL, Kuiken T, Marangon S, Niqueux É, Staubach C, Terregino C, Aznar I, Muñoz Guajardo I and Baldinelli F. Scientific report: Avian influenza overview March–June 2022. EFSA J. 2022;20:
e07415 . - Camphuysen CJ, Gear SC, Furness RW. Avian influenza leads to mass mortality of adult Great Skuas in Foula in summer 2022. Scottish Birds. 2002;4:312–23
- Vergeer JW, van Dijk AJ, Boele A, van Bruggen J, Hustings F. Sovon breeding bird research manual: breeding bird monitoring project and colony birds [in Dutch]. 2016 [cited 2022 Jul 19]. https://stats.sovon.nl/static/publicaties/Handleiding_Broedvogels_2016.pdf
- Brenninkmeijer A, Stienen EWM. Ecological profile of the Sandwich tern (Sterna sandvicensis) [in Dutch]. Arnhem: DLO-Instituut voor Bos- en Natuuronderzoek; 1992.
- Koffijberg K, de Boer P, Geelhoed SCV, Nienhuis J, Schekkerman H, Oosterbeek K, et al. Breeding success of coastal breeding birds in the Wadden Sea in 2019. WOt-technical report 209. Wageningen: Wettelijke Onderzoekstaken Natuur & Milieu; 2021.
- Schekkerman H, Arts F, Buijs R-J, Courtens W, van Daele T, Fijn R, et al. Population analysis of five coastal breeding bird in the SW Netherlands [in Dutch]. Nijmegen: Sovon Vogelonderzoek Nederland; 2021.
- Beerens N, Heutink R, Bergervoet SA, Harders F, Bossers A, Koch G. Multiple reassorted viruses as cause of highly pathogenic avian influenza A(H5N8) virus epidemic, the Netherlands, Nijmegen. 2016. Emerg Infect Dis. 2017;23:1974–81. DOIPubMedGoogle Scholar
- Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80. DOIPubMedGoogle Scholar
- Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol. 2015;32:268–74. DOIPubMedGoogle Scholar
- Yu G, Smith DK, Zhu H, Guan Y, Tsan-Yuk Lam T. ggtree: an r package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol Evol. 2017;8:28–36. DOIGoogle Scholar
- Lean FZX, Vitores AG, Reid SM, Banyard AC, Brown IH, Núñez A, et al. Gross pathology of high pathogenicity avian influenza virus H5N1 2021-2022 epizootic in naturally infected birds in the United Kingdom. One Health. 2022;14:
100392 . DOIPubMedGoogle Scholar - Garg PK, Singh VP. Organ failure due to systemic injury in acute pancreatitis. Gastroenterology. 2019;156:2008–23. DOIPubMedGoogle Scholar
- Hamer KC, Schreiber E, Burger J. Breeding biology, life histories, and life history-environment interactions in seabirds. In: Schreiber E, Burger J, editors. Biology of marine birds. London/New York: CRC Press; 2001. p. 217–262.
- Banyard AC, Lean FZX, Robinson C, Howie F, Tyler G, Nisbet C, et al. Detection of highly pathogenic avian influenza virus H5N1 clade 2.3.4.4b in great skuas: a species of conservation concern in Great Britain. Viruses. 2022;14:212. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: November 18, 2022
1These authors contributed equally to this article.
Table of Contents – Volume 28, Number 12—December 2022
| EID Search Options |
|---|
|
|
|
|
|
|




Please use the form below to submit correspondence to the authors or contact them at the following address:
Jolianne M. Rijks, Dutch Wildlife Health Centre, Utrecht University, Yalelaan 1, 3584CL, Utrecht, the Netherlands
Top