Volume 28, Number 2—February 2022
Research Letter
SARS-CoV-2 Circulation, Guinea, March 2020–July 2021
Figure
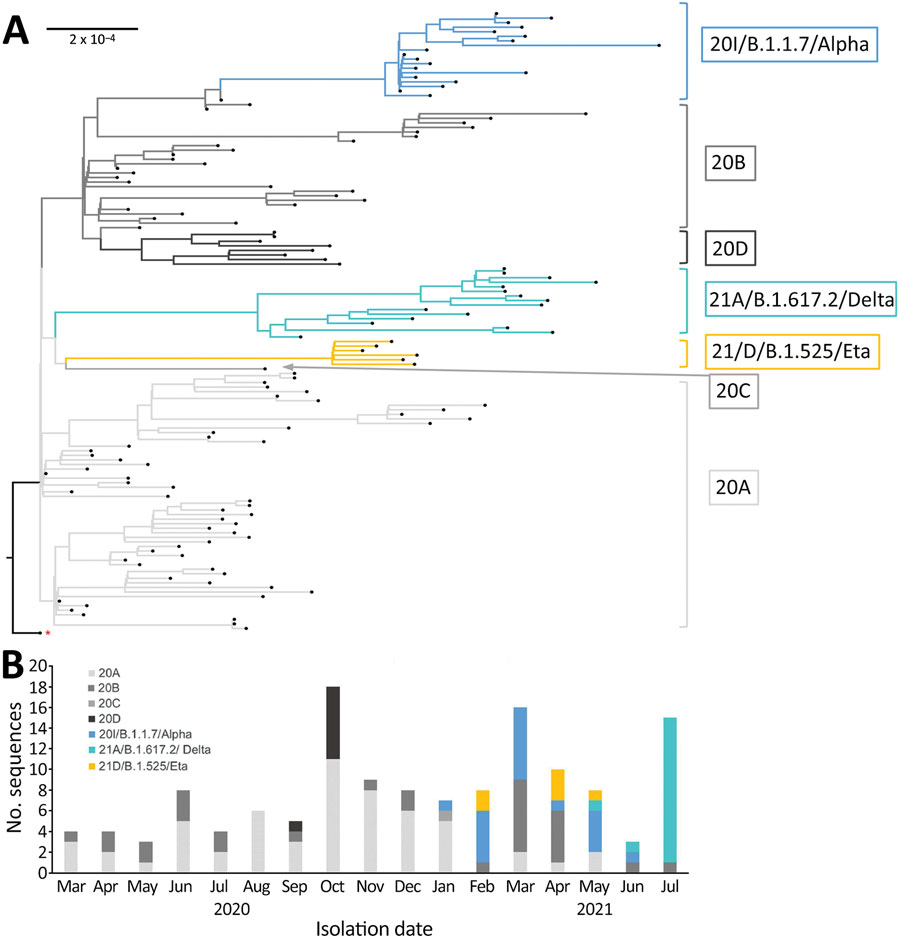
Figure. Phylogenetic and temporal descriptions of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) sequences from Institut Pasteur de Guinée from samples collected in Guinea during March 12, 2020–July 16, 2021. A) Maximum-likelihood phylogenetic tree of 136 SARS-CoV-2 genomic sequences. The tree was constructed with IQ-tree software by using multiple-genome sequence alignment and Wuhan-Hu-1 strain (GenBank accession no. NC 045512) as outgroup reference sequence, indicated by the red asterisk. Branches and the sequence names are colored according to Nextclade assigned clades: 20A, light gray; 20B, medium gray; 20C, dark gray; 20D, black; 20I/B.1.1.7/Alpha, blue; 21A/B.1.617.2/Delta, azure; 21D/B.1.525/Eta, yellow. Each sequence is highlighted by a black tip. Scale bar indicates the distance corresponding to substitution per site. B) Chronologic distribution of SARS-CoV-2 genomic variants over 17 months in Guinea. The 136 selected sequences are assigned by Nextclade and classified according to sampling date from March 31, 2020, to July 16, 2021. Clades are colored as in panel A.
1These authors contributed equally to this article.
2Current affiliation: Institut Pasteur du Cambodge, Phnom Penh, Cambodia.