Volume 29, Number 2—February 2023
Dispatch
Nipah Virus Exposure in Domestic and Peridomestic Animals Living in Human Outbreak Sites, Bangladesh, 2013–2015
Cite This Article
Citation for Media
Abstract
Spillovers of Nipah virus (NiV) from Pteropus bats to humans occurs frequently in Bangladesh, but the risk for spillover into other animals is poorly understood. We detected NiV antibodies in cattle, dogs, and cats from 6 sites where spillover human NiV infection cases occurred during 2013–2015.
Henipaviruses are batborne zoonoses that have caused fatal neurologic and respiratory disease outbreaks in humans, horses, and pigs. In Bangladesh, the Indian flying fox (Pteropus medius) is the known natural reservoir for Nipah virus (NiV). NiV causes annual outbreaks in humans in Bangladesh, where the primary mode of spillover is through consumption of date palm sap contaminated by P. medius bats (1); NiV infection is a particular concern for public health because of the high case-fatality ratio and the risk for person-to-person transmission (2).
Domestic and peridomestic animals have been important intermediate hosts for zoonotic henipavirus transmission in outbreaks occurring in Australia, Malaysia, and the Philippines (3,4). One cross-sectional study suggested possible exposure of livestock to henipaviruses in Bangladesh (5). Three instances in which animal contact was associated with human NiV infections in Bangladesh have been reported (1,6,7), although little is known about the transmission mechanisms of henipaviruses into livestock and peridomestic animals in Bangladesh. Our study aimed to detect prior NiV infection among livestock and peridomestic animals living in proximity to humans with spillover cases and identify possible exposure pathways in Bangladesh.
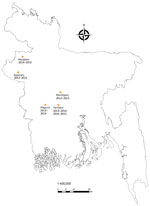
During January 2013–January 2015, a total of 6 confirmed human Nipah outbreaks were identified through the Nipah surveillance system in Bangladesh (Figure) (8). Once an index case-patient was identified, we identified the closest bat roosts to the case-patient’s household and collected urine from underneath the roosts by using plastic tarps. We aliquoted roost urine in cryovials containing lysis buffer, stored them at cryogenic temperatures, and tested them for evidence of NiV RNA. We used extracted RNA from bat roost urine for detecting NiV by using a probe-based real-time reverse transcription PCR assay (9). Roosts were located from 150 m to 2 km from the human spillover index case-patient’s household for all 6 outbreaks (Table 1). Ultimately, we sampled only 5 of the 6 roosts; 1 roost could not be sampled because of political unrest. We identified evidence of NiV RNA shedding in urine collected from 4 roosts. We defined a positive sample as one having >1 aliquot with a cycle threshold value <39.
During October 2013–October 2015, at 4–9 months after onset in human case-patients, we revisited the villages surrounding each bat roost to test for evidence of infection among domestic animals (e.g., cattle and goats) and peridomestic animals (e.g., dogs, cats, rodents, and house shrews) living near the bat roosts; none of the villages had animal cases during outbreaks. Starting from the household closest to each bat roost, we approached each nearby household to determine whether they owned cattle, goats, cats, or dogs that had resided there for the previous month; households owning any of these animals were asked to participate in the study. We continued this process until we reached the target sample size of 100 domestic animals (0.1% prevalence, 3% null, 80% power, 95% CI, and design effect of 2) and 90 peridomestic animals (1.0% prevalence, 3% null, 80% power, 95% CI, and design effect of 2) in each village. In the first 2 sites we could not sample the targeted number of domestic animals because of unavailability according to our inclusion criteria.
We asked each animal owner about rearing practices, the health status of animals during the month before the human Nipah outbreak, and whether the animal was fed date palm sap or fruit found on the ground or was ever observed to scavenge bat carcasses or bat placentas. For blood collection, we manually restrained cattle, goats, dogs, and cats and captured rats (Rattus rattus, Bandicota bengalensis, B. indica) and shrews (Suncas murinus) by using baited traps in and around the houses of the study participants. We sent serum samples to the US Centers for Disease Control and Prevention (Atlanta, GA, USA), where a human IgG ELISA (10) was adapted and optimized to test animal serum by using alternative positive and negative control serum and horseradish peroxidase–conjugated Pierce Recombinant Protein A/G (ThermoFisher Scientific, https://www.thermofisher.com). The ELISA was developed by infecting Vero-E6 cells with whole NiV. The study protocol was reviewed and approved by Research Review Committee and Animal Experimentation Ethics Committee of icddr,b.
We sampled 1,156 animals from 369 households at 6 locations (Table 1). No sick animals were reported by animal owners around the time of the human outbreak or during sampling. Previous studies indicate that, except for cats, NiV does not cause severe infection among the animals that we sampled (11). Each study site had >1 animal with evidence of IgG against NiV. Serum samples from 1% of cattle (3/274), 3% of dogs (5/189), and 5% of cats (4/85) had evidence of NiV antibodies (Table 1). Thirteen cattle (5%) and 3 goats (1%) were fed date palm sap, but none had evidence of NiV antibodies. One third of cattle (91/274) and goats (110/330) were fed dropped fruit, including 2 cattle with NiV antibodies (Table 2). No owner of either dogs or cats reported observing their animal feeding on bat carcasses or bat placentas.
No animal owners reported sick animals at the time of the human Nipah outbreak. However, the long gap between the outbreak and the survey may have led to underreporting of clinical signs by owners. Pteropid bats forage on fruit trees near human residences in Bangladesh and drop partially eaten fruits on the ground (12). Those partially eaten fruits could act as a source of infection with NiV (or related henipaviruses) for cattle and goats (13). Serologic evidence of NiV infection in goats was reported from Malaysia during the human Nipah outbreak during 1998–1999 (14). Cattle and goats were very rarely fed date palm sap but frequently fed dropped fruit, and nearly all livestock with NiV antibodies were fed dropped fruit. A previous study from Bangladesh detected NiV antibodies among cattle, goats, and pigs using a Luminex assay (biotechne, https://www.bio-techne.com), but none of the animals with antibodies detected in that study had antibodies detected by CDC’s in-house ELISA used in our study (5). A conjugate that is more general to catching all species was used because controls for each of the species tested, except for pigs, were not available. CDC’s in-house ELISA was previously used to test for Hendra virus and NiV antibodies and showed cross-reaction for Hendra virus IgG (6). The positive animals in the earlier study could have been exposed to NiV or a related henipavirus, given that P. medius bats carry genetically diverse genotypes that might be antigenically similar.
Our study found that a small proportion of domestic animals (e.g., cattle and goats) had evidence of NiV antibodies, whereas evidence of antibodies were more common in peridomestic animals (e.g., cats and dogs). Because peridomestic dogs and cats roam around freely, they may have scavenged bat carcasses or placentas underneath the roost without knowledge of the owners. Previous studies have described henipavirus infection among dogs and cats in other countries (4,11,15). Characterizing the risk factors for infection and the potential role of domestic animals as intermediate or amplifying hosts in Bangladesh would provide a more complete understanding of the ecology of NiV in Bangladesh. In addition, to avoid potential spillover, owners of domestic animals could be advised to not feed them dropped fruit.
Dr. Islam is a veterinarian at icddr,b (formerly known as the International Centre for Diarrhoeal Disease Research, Bangladesh). His primary research interests include zoonotic pathogens.
Acknowledgment
This research study was funded by the National Academy of Sciences. The Centers for Disease Control and Prevention conducted the sample testing using their resources. This manuscript was developed with funding from the Defense Advanced Research Projects Agency (administered through cooperative agreement no. D18AC00031-PREEMPT) and from the National Institutes of Health’s National Institute for Allergy and Infectious Diseases (grant no. U01AI15320). icddr,b acknowledges with gratitude the commitment of the National Academy of Sciences, the Centers for Disease Control and Prevention, and the Defense Advanced Research Projects Agency to its research efforts. icddr,b is also grateful to the governments of Bangladesh, Canada, Sweden, and the United Kingdom for providing core and unrestricted support.
References
- Luby SP, Gurley ES, Hossain MJ. Transmission of human infection with Nipah virus. Clin Infect Dis. 2009;49:1743–8. DOIPubMedGoogle Scholar
- Gurley ES, Montgomery JM, Hossain MJ, Bell M, Azad AK, Islam MR, et al. Person-to-person transmission of Nipah virus in a Bangladeshi community. Emerg Infect Dis. 2007;13:1031–7. DOIPubMedGoogle Scholar
- Looi LM, Chua KB. Lessons from the Nipah virus outbreak in Malaysia. Malays J Pathol. 2007;29:63–7.PubMedGoogle Scholar
- Ching PK, de los Reyes VC, Sucaldito MN, Tayag E, Columna-Vingno AB, Malbas FF Jr, et al. Outbreak of henipavirus infection, Philippines, 2014. Emerg Infect Dis. 2015;21:328–31. DOIPubMedGoogle Scholar
- Chowdhury S, Khan SU, Crameri G, Epstein JH, Broder CC, Islam A, et al. Serological evidence of henipavirus exposure in cattle, goats and pigs in Bangladesh. PLoS Negl Trop Dis. 2014;8:
e3302 . DOIPubMedGoogle Scholar - Hsu VP, Hossain MJ, Parashar UD, Ali MM, Ksiazek TG, Kuzmin I, et al. Nipah virus encephalitis reemergence, Bangladesh. Emerg Infect Dis. 2004;10:2082–7. DOIPubMedGoogle Scholar
- icddr,b. Outbreaks of encephalitis due to Nipah/Hendra-like viruses, western Bangladesh. Health Sci Bull. 2003;1:1–6.
- Rahman M, Chakraborty A. Nipah virus outbreaks in Bangladesh: a deadly infectious disease. WHO South-East Asia J Public Health. 2012;1:208–12. DOIPubMedGoogle Scholar
- Lo MK, Lowe L, Hummel KB, Sazzad HMS, Gurley ES, Hossain MJ, et al. Characterization of Nipah virus from outbreaks in Bangladesh, 2008-2010. Emerg Infect Dis. 2012;18:248–55. DOIPubMedGoogle Scholar
- Daniels P, Ksiazek T, Eaton BT. Laboratory diagnosis of Nipah and Hendra virus infections. Microbes Infect. 2001;3:289–95. DOIPubMedGoogle Scholar
- Middleton DJ, Westbury HA, Morrissy CJ, van der Heide BM, Russell GM, Braun MA, et al. Experimental Nipah virus infection in pigs and cats. J Comp Pathol. 2002;126:124–36. DOIPubMedGoogle Scholar
- Sudhakaran MR, Doss PS. Food and foraging preferences of three pteropodid bats in southern India. J Threat Taxa. 2012;4:2295–303. DOIGoogle Scholar
- Chua KB, Koh CL, Hooi PS, Wee KF, Khong JH, Chua BH, et al. Isolation of Nipah virus from Malaysian Island flying-foxes. Microbes Infect. 2002;4:145–51. DOIPubMedGoogle Scholar
- Field H, Young P, Yob JM, Mills J, Hall L, Mackenzie J. The natural history of Hendra and Nipah viruses. Microbes Infect. 2001;3:307–14. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleOriginal Publication Date: January 11, 2023
Table of Contents – Volume 29, Number 2—February 2023
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Ausraful Islam, icddr,b, 68 Shaheed Tajuddin Ahmed Sarani, Mohakhali, Dhaka-1212, Bangladesh
Top