Volume 30, Supplement—October 2024
SUPPLEMENT ISSUE
Articles
Using SARS-CoV-2 Sequencing Data to Identify Reinfection Cases in the Global Emerging Infections Surveillance Program, United States
Figure 4
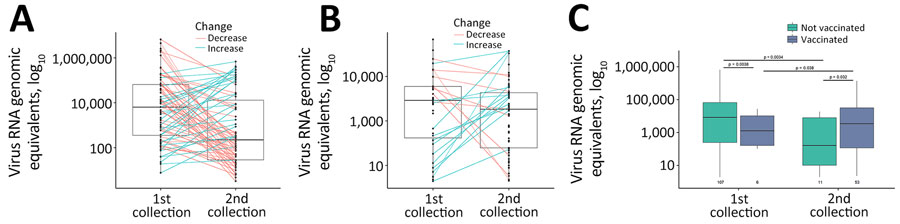
Figure 4. Virus load in patient specimens in study using SARS-CoV-2 sequencing data to identify reinfection cases in Department of Defense Global Respiratory Pathogen Surveillance Program, United States. Virus load was determined in specimens collected during the first and second timepoints by using quantitative PCR of the N1 region of the SARS-CoV-2 nucleocapsid gene for patients who had continuing infections (A) and reinfections (B) or who were vaccinated versus unvaccinated (C). Middle horizontal lines within each box plot are the median virus RNA genomic equivalents, outer horizontal lines indicate the interquartile range, and whiskers (vertical lines) indicate minimum and maximum data points. A) Significant decrease in virus load was observed between the first and second collection timepoints for patients who had continuing infections (p = 0.039 by Student t-test); average number of days between collection dates was 8.7 (range 1–43) days. B) No significant difference in virus RNA load was observed between the first and second collection points for patients who had reinfections (p = 0.290 by Student t-test). C) First collection group shows all first infections for patients who had either continuing or reinfections. Second collection group shows only reinfections. Numbers under box plots indicate the number of cases within each group.