Volume 32, Number 4—April 2026
Dispatch
Whole-Genome Analysis of Treponema pallidum Subspecies endemicum among Men Who Have Sex with Men, Japan, 2020–2023
Figure
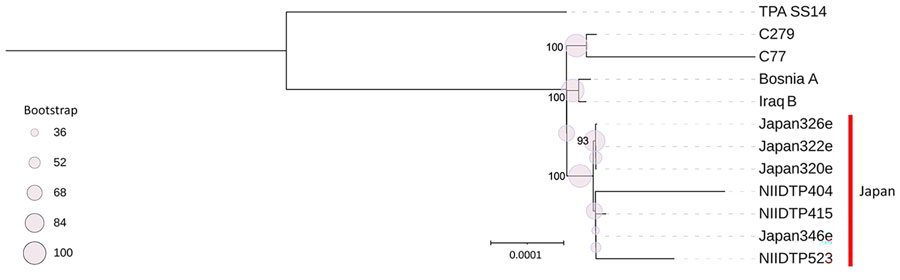
Figure. Phylogenetic analysis of isolates from a study of whole-genome analysis of Treponema pallidum subspecies endemicum among men who have sex with men, Japan, 2020–2023. Open reading frames annotated from de novo assembled contigs using Bakta version 1.7.0 (https://bakta.computational.bio), and corresponding GFF3 annotation files processed with Panaroo version 1.2.3 (https://zenodo.org/records/3945497) to define core genomes across isolates. Multiple alignments of core gene sequences generated by Panaroo were used for phylogenetic reconstruction. T. pallidum subsp. pallidum SS14 (GenBank accession no. NC_021508) was used as a reference strain. IQ-TREE version 2.2.0 (http://www.iqtree.org) was used to infer maximum-likelihood phylogenies, incorporating 1,000 ultrafast bootstrap replicates to assess nodal support. Phylogenetic tree was visualized by using Interactive Tree of Life (iTOL; https://itol.embl.de). The T. pallidum subsp. endemicum strains isolated in Japan formed a clade distinct from classic endemic strains, including Iraq B and strains from Cuba (C77 and C279). Scale bar indicates nucleotide substitutions per site.