Disclaimer: Early release articles are not considered as final versions. Any changes will be reflected in the online version in the month the article is officially released.
Volume 32, Number 6—June 2026
Dispatch
Concurrent Detection of Swine-Origin Influenza A(H1N1) Virus in Pigs and Farmer, Switzerland
Suggested citation for this article
Abstract
We report zoonotic transmission of Eurasian avian-like swine influenza A(H1N1) virus from pigs to a farmer. The pigs and farmer experienced influenza-like illness. Whole-genome sequencing revealed >99.9% isolate sequence identity between hosts. Our findings highlight the risk posed by enzootic swine influenza A virus and the need for genomic and epidemiologic surveillance.
Since the 2009 swine influenza pandemic, sporadic human infections with swine influenza A viruses (swIAVs) continue to occur, including rare instances of onward human-to-human transmission, highlighting the ongoing pandemic risk (1–4). Pigs are key reservoirs and mixing vessels for influenza A virus (IAV) evolution; transmission between humans and pigs is frequent and bidirectional (5,6). Active epidemiologic and genomic surveillance at the swine–human interface is therefore essential for early detection and risk assessment of emerging strains.
In Switzerland, pig production is less intensive; herds remain relatively isolated from neighboring countries because of strict regulations and minimal import of live pigs (7,8). Infrequent use of swIAV vaccines allows for natural viral evolution without vaccine-driven selective pressures (9); country-specific transmission chains may exist within pig herds in Switzerland. The ongoing national surveillance program for swIAV relies primarily on partial hemagglutinin (HA) and neuraminidase (NA) gene sequences, limiting its ability to identify emerging swIAV lineages in pig herds (10).
To investigate the epidemiology and genetic diversity of swIAV in pig herds, we established a whole-genome sequencing (WGS)–based swIAV surveillance program. We obtained samples from symptomatic and randomly selected pig herds; pig caretakers with respiratory illness voluntarily provided self-collected nasal swab specimens. Here, we report a zoonotic transmission event involving Eurasian avian-like (EA) swine influenza A(H1N1) virus detected concurrently in a farmer and his pig herd through WGS surveillance.
On November 27, 2023, a respiratory disease outbreak was reported in a herd of »180 fattening pigs, 4 weeks after the introduction of 90 growing pigs from another herd from within Switzerland. Upon clinical examination, »80% of pigs showed apathy and fever (<40.3°C) and sporadic respiratory signs. The farmer, who was 40–50 years of age with no reported underlying conditions, experienced a mild influenza-like illness 2 weeks earlier and reported similar symptoms in 2 household members. The farmer was not vaccinated against seasonal influenza and reported no contact with other pig herds.
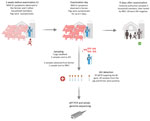
We collected and analyzed nasal swab samples from pigs and the farmer through our WGS-based surveillance program and the national swIAV surveillance system. Within the WGS surveillance framework, we screened 5 pig samples (A0001–A0005) and 1 farmer sample (H0001) for IAV using a pan-IAV matrix gene–specific quantitative reverse transcription PCR assay on extracted viral RNA (Appendix Table 1) (11). All samples tested positive; viral loads were higher in pigs (crossing point [Cp] value 24–26) than in the farmer sample (Cp 32) (Table 1). After confirming variant H1N1 (H1N1v) infection in the farmer, we notified veterinary and public health authorities and reported the case through the World Health Organization National Focal Point in accordance. Nine days after case confirmation, nasal swabs from all 3 household contacts tested negative for IAV at the Swiss National Reference Centre for Influenza (NRCI) (Figure 1) (A.R. Gonçalves Cabecinhas, pers. comm., email, 2024 Dec 2).
We performed WGS as described previously (12), achieving high read depth (>1,000´) overall; 1 segment showed lower coverage (>100´), sufficient for complete genome assembly (Table 1; Appendix Table 1, Figure 1). We subtyped all viruses from human and pig samples as swine H1N1 (swH1N1); nucleotide identity was >99.9% across all genomic segments. Phylogenetic analysis with publicly available swH1N1 sequences from Switzerland and Europe showed that all study viruses were assigned to the EA swH1N1 lineage, clade 1C.2.2, forming a monophyletic cluster for all genomic segments (Figure 2; Appendix Figure 2). Similar to other EA swH1N1 viruses, the Switzerland sequences harbored several amino acid substitutions associated with host specificity and antiviral resistance (Appendix Table 2).
We assessed low-frequency diversity by variant calling across all genomic segments. We detected 26 minor single-nucleotide variants (SNVs) at 6%–50% frequency, resulting in 17 nonsynonymous mutations. We detected 4 human-specific SNVs, including C1771T (H0001), which resulted in a premature stop codon in the polymerase acidic (PA) gene (Q591*). We identified 4 SNVs shared between human and pig samples, including G515A, which resulted in the HA-G172E mutation associated with antigenic drift and potential immune escape (Appendix Table 3, Figure 3) (13). We detected 1 SNV (PB2–1447), leading to a T483A mutation, in all samples. The presence of shared SNVs supports epidemiologic linkage between the farmer and pigs.
Finally, we compared HA antigenic residues of study viruses to the 2023–24 human seasonal vaccine strain and a contemporaneous (2023) human H1N1 sequence (Table 2). We identified multiple substitutions in the swH1N1 sequences across defined antigenic sites Sa/Sb, Ca1, Ca2, and Cb; the contemporaneous human seasonal virus differed from the vaccine strain by 2 antigenic residues. Overall, swH1N1 sequences from pigs and the farmer showed greater divergence at HA antigenic sites relative to the vaccine strain than the contemporaneous human seasonal virus, consistent with their evolutionary divergence from human H1N1 viruses.
This study reports the concurrent detection of EA swH1N1 virus in a farmer and his pigs in Switzerland. Evidence for zoonotic transmission is >99.9% genomic identity between viruses from the farmer and pigs, shared SNVs between hosts, and epidemiologic data consistent with transmission at the swine–human interface. We identified several human-specific SNVs, but their functional significance remains unknown. We observed 1 SNV, PA C1771T (Q591*), at the consensus level in 1 pig sample; however, we cannot infer transmission direction or functional significance on the basis of the available data. The lower viral load we observed in the farmer sample may reflect previous immunity, timing of sampling, host-specific constraints, or the effects of the detected SNVs. We detected no onward transmission to household contacts; we could not exclude the possibility of secondary transmission because of the timing of sample collection.
Genomic analysis revealed that the swH1N1 viruses identified in this study clustered with previously detected Switzerland sequences belonging to the EA lineage, clade 1C.2.2 (10). Despite limited availability of complete swH1N1 genome sequences from Switzerland, our findings are consistent with the presence of a relatively homogeneous swIAV population in pigs, likely shaped by the closed herd system and low annual importation of live pigs. Similarly, in Norway, the circulation of a single lineage under a closed system highlighted the effect of production structure on viral evolution (14). In contrast, greater swIAV genetic diversity has been reported in other countries in Europe (15).
HA sequences of study swH1N1 viruses harbored multiple amino acid substitutions across defined antigenic regions, whereas contemporary human H1N1 strains differed from vaccine strains by only 1 or 2 residues. The detection of 1 SNV associated with HA antigenic drift highlighted hidden evolution; that finding suggests potentially limited cross-protection from human vaccines for exposed workers, underscoring the need for targeted vaccination strategies for high-risk occupational groups. Our investigation was restricted to 1 outbreak and a limited number of samples; we used no serologic data and only partial comparative sequences from Switzerland, which limited our assessment of evolutionary viral dynamics. Despite those constraints, the clinical, epidemiologic, and genomic evidence supports zoonotic transmission.
Concurrent detection of swIAV in pigs and humans is rare globally. The detection of EA swH1N1 in pigs and a farmer in our study demonstrates zoonotic spillover and highlights the value of One Health surveillance. The viruses were genetically homogenous, yet antigenically distinct from contemporary human vaccine strains, emphasizing the need for active WGS surveillance and genomic characterization at the swine–human interface, as well as continued monitoring and targeted preventive strategies for pig-exposed workers.
Dr. Steiner is a veterinarian and PhD student at the Clinic for Swine, Department of Clinical Veterinary Medicine, Vetsuisse Faculty, University of Bern, and the Multidisciplinary Center for Infectious Diseases, University of Bern. His work focuses on the epidemiology of influenza A viruses in pig herds and their caretakers.
Acknowledgments
We thank the Swiss Pig Health Service (Schweinegesundheitsdienst) within SUISAG and the collaborating veterinary practices for referring the case to us, as well as members of the Experimental Virology group and NGS sequencing unit at the Institute for Infectious Diseases, the Swine Clinic of the University of Bern, and the Institute of Virology and Immunology for their support. We thank Ana Rita Goncalves Cabecinhas for kindly sharing the results of samples processed at the Swiss National Reference Centre for Influenza.
This study was supported by the Multidisciplinary Center for Infectious Diseases, University of Bern, and the Swiss National Science Foundation (grant no. IZCOZ0_220329).
We used ChatGPT version 5.2 to assist with structuring the manuscript and adjusting the formatting of the text and the references.
References
- Andersen KM, Vestergaard LS, Nissen JN, George SJ, Ryt-Hansen P, Hjulsager CK, et al. Severe human case of zoonotic infection with swine-origin influenza A virus, Denmark, 2021. Emerg Infect Dis. 2022;28:2561–4. DOIPubMedGoogle Scholar
- Dürrwald R, Wedde M, Biere B, Oh DY, Heßler-Klee M, Geidel C, et al. Zoonotic infection with swine A/H1avN1 influenza virus in a child, Germany, June 2020. Euro Surveill. 2020;25:
2001638 . DOIPubMedGoogle Scholar - Parys A, Vandoorn E, King J, Graaf A, Pohlmann A, Beer M, et al. Human infection with Eurasian avian-like swine influenza A(H1N1) virus, the Netherlands, September 2019. Emerg Infect Dis. 2021;27:939–43. DOIPubMedGoogle Scholar
- Jhung MA, Epperson S, Biggerstaff M, Allen D, Balish A, Barnes N, et al. Outbreak of variant influenza A(H3N2) virus in the United States. Clin Infect Dis. 2013;57:1703–12. DOIPubMedGoogle Scholar
- Vincent Baker AL, Van Reeth K. Influenza viruses. In: Burrough ER, Karriker LA, Schwartz KJ, Zhang J, editors. Diseases of swine. 12th ed. Hoboken (NJ): John Wiley & Sons; 2026. p. 645–64.
- Gray GC, McCarthy T, Capuano AW, Setterquist SF, Olsen CW, Alavanja MC, et al. Swine workers and swine influenza virus infections. Emerg Infect Dis. 2007;13:1871–8. DOIPubMedGoogle Scholar
- Federal Office for Customs and Border Security. Swiss-Impex—public import/export database. 2025 [cited 2025 Jan 27]. https://www.gate.ezv.admin.ch/swissimpex/index.xhtml
- Federal Food Safety and Veterinary Office. Pig husbandry [cited 2025 Jan 31]. https://www.blv.admin.ch/blv/de/home/tiere/tierschutz/nutztierhaltung/schweine.html
- Li C, Culhane MR, Schroeder DC, Cheeran MCJ, Galina Pantoja L, Jansen ML, et al. Quantifying the impact of vaccination on transmission and diversity of influenza A variants in pigs. J Virol. 2024;98:
e0124524 . DOIPubMedGoogle Scholar - Lechmann J, Szelecsenyi A, Bruhn S, Harisberger M, Wyler M, Bachofen C, et al. The Swiss national program for the surveillance of influenza A viruses in pigs and humans: genetic variability and zoonotic transmissions from 2010–2022. Schweiz Arch Tierheilkd. 2025;167:600–16. DOIPubMedGoogle Scholar
- Nagy A, Černíková L, Kunteová K, Dirbáková Z, Thomas SS, Slomka MJ, et al. A universal RT-qPCR assay for “One Health” detection of influenza A viruses. PLoS One. 2021;16:
e0244669 . DOIPubMedGoogle Scholar - Licheri M, Mwanga M, Licheri MF, Graaf-Rau A, Sägesser C, Bittel P, et al. Optimized high-throughput whole-genome sequencing workflow for surveillance of influenza A virus. Genome Med. 2025;17:103. DOIPubMedGoogle Scholar
- Yoshida R, Igarashi M, Ozaki H, Kishida N, Tomabechi D, Kida H, et al. Cross-protective potential of a novel monoclonal antibody directed against antigenic site B of the hemagglutinin of influenza A viruses. PLoS Pathog. 2009;5:
e1000350 . DOIPubMedGoogle Scholar - Forberg H, Hauge AG, Gjerset B, Hungnes O, Kilander A. Swine influenza in Norway: a distinct lineage of influenza A(H1N1)pdm09 virus. Influenza Other Respir Viruses. 2013;7(Suppl 4):21–6. DOIPubMedGoogle Scholar
- Richard G, Ryt-Hansen P, Byrne A; European Swine Influenza Network. European Swine Influenza Network report #2 on swine influenza A viruses evolution and diversity of in Europe from October 2022–September 2024. 2025 [cited 2026 May 7]. DOIGoogle Scholar
Figures
Tables
Suggested citation for this article: Steiner J, Mwanga M, Oberholster L, Licheri M, Licheri MF, Nathues H, et al. Concurrent detection of swine-origin influenza A(H1N1) virus in pigs and farmer, Switzerland. Emerg Infect Dis. 2026 Jun [date cited]. https://doi.org/10.3201/eid3206.251487
Original Publication Date: May 13, 2026
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
Table of Contents – Volume 32, Number 6—June 2026
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Jenna N. Kelly, Institute of Virology and Immunology (IVI), Länggassstrasse 122, CH-3012 Bern, Switzerland
Top