Volume 32, Number 5—May 2026
Dispatch
Tropism and Replication Competence of Cattle Influenza A(H5N1) Genotype B3.13 Virus in Human Bronchus and Lung Tissue
Cite This Article
Citation for Media
Abstract
In 2024, influenza A(H5N1) genotype B3.13 viruses emerged from cattle and caused mild spillover infections in humans. Using human bronchus and lung tissue, we evaluated tropism, replication, and pathogenesis of 2 cattle influenza isolates. Those viruses showed moderate replication competence and induced robust proinflammatory responses, suggesting potential risk for human health.
Highly pathogenic avian influenza (HPAI) H5N1 viruses remain a major global health concern, particularly because of sporadic spillover into mammals (1). HPAI A(H5N1) clade 2.3.4.4b viruses entered the United States through a trans-Atlantic introduction in late 2021, after which extensive reassortment among migratory birds produced the B3.13 and D1.1 genotypes. Those variants have spread widely, driving outbreaks in livestock and causing occasional human infections (2,3). Beginning in 2024, H5N1 clade 2.3.4.4b viruses were detected in dairy cattle across multiple US states, and those infections were linked to mild zoonotic cases in humans (4,5). To assess the health risks of emerging cattle-origin influenza viruses, we examined tropism, replication, receptor use, and innate immune responses of cattle H5N1 viruses in human respiratory tract explants.
We investigated newly emerged cattle influenza A(H5N1) genotype B3.13 virus strains A/dairy_cow/Ohio/B24OSU-439/2024 (H5N1/439) and A/dairy_cow/Texas/98638/2024 (H5N1/98638) in human bronchial and lung tissue cultures (Appendix). In brief, we obtained nonmalignant tissue cultures from patients who underwent elective surgery and consented to tissue use (Appendix). We used the RNeasy Micro Kit (QIAGEN, https://www.qiagen.com) to extract total RNA, according to manufacturer instructions, then reverse transcribed extracted RNA by using the PrimeScript RT Reagent Kit (TaKaRa Bio, Inc., http://www.takara-bio.com). We fixed the explant tissues for immunohistochemical staining of influenza viral proteins.
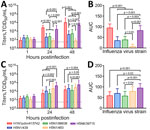
We compared explant cultures with 3 historical human isolates: H5N1/483, H5N6/39715, and H1N1pdm/415742 (Appendix Tables 1, 2). The historical virus strains showed different tropisms and area under the curve (AUC) levels to the explant tissue cultures at 24–48 hours postinfection (hpi). In bronchus tissues, H1N1pdm/415742 replication was highest, followed by H5N6/39715 (Figure 1, panels A, B). The H5N1/439 and H5N1/98638 cattle isolates showed substantially lower replication competence at 48 hpi than H1N1pdm/415742 and H5N6/39715 but much higher competence than the H5N1/483 avian isolate, which replicated poorly in the human bronchial tissues. In human lung tissues, H5N6/39715 replicated to the highest viral titers and AUC values at 24 hpi, followed by H5N1/483 (Figure 1, panels C, D). Both cattle isolates replicated to similar levels as H1N1pdm/415742, but isolate H5N1/439 had a slightly higher replication trend than H5N1/98638. The cattle-origin viruses replicated to lower titers than HPAI H5N1 but similar to H1N1pdm/415742 in human lung tissues, which aligns with other reports in human lung organoids (6). However, cattle-origin H5N1 genotype B3.13 virus has lower replication than genotype D1.1 in human nasal and airway organoids (7). Replication competence in human respiratory tissues might be contributed by the 631L of polymerase basic protein 2 (Appendix Table 2), which promotes polymerase activity in human cells (8). In addition, the cattle-origin H5N1 viruses replicated to higher titers than avian H5N1 viruses in bronchial tissues, implying that the cattle-origin H5N1 viruses might be more transmissible than HPAI H5N1 viruses.
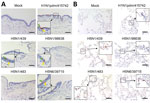
Immunohistochemical staining revealed the tissue and cellular tropism of cattle-origin H5N1 viruses. We found nucleoprotein (NP)–positive cells from H1N1pdm/415742 virus and to a lesser extent H5N6/39715 in the bronchial epithelium, infecting ciliated and nonciliated epithelial cells (Figure 2, panel A). In comparison, we noted moderate levels of NP-positive cells from cattle H5N1/439 and H5N1/98638 viruses in ciliated and nonciliated epithelial cells but identified no NP-positive cells from avian-origin H5N1/483 in the bronchial sections. In the lung sections, H5N1/483 and H5N6/39715 demonstrated the most extensive infections, followed by H5N1/439, H5N1/98638, and H1N1pdm/415742 (Figure 2, panel B). Together with the viral replication data, those results imply that cattle-origin H5N1/439 and H5N1/98638 viruses possess moderate replication capacity in upper and lower airways and are better adapted to human hosts than avian H5N1/483.
To test virus agglutination, we conducted selective desialylation on turkey red blood cells (TRBCs). For controls, we used H1N1pdm/415742, known for binding to α(2–6)-linked sialic acid (SA), and H5N1/483, known for binding to α(2,3)-linked SA. Treating TRBCs with Sialidase S (Agilent, https://www.agilent.com), which preferentially cleaves the α(2,3)-linked SA, prevented hemagglutination of avian H5N1/483 but did not affect H1N1pdm/415742, H5N1/439, or H5N1/98638 isolates (Table). Conversely, treating with Sialidase C (Agilent), which cleaves α(2,3) and α(2–6) linkages, and Sialidase A (Agilent), which cleaves α(2,3), α(2–6), α(2–8), and α(2–9) linkages, prevented hemagglutination of all the influenza strains. Those results suggest that the cattle-origin H5N1/439 and H5N1/98638 isolates resemble H1N1pdm/415742 and acquired at least partial affinity to α(2–6)-linked SA, which differs from the avian H5N1/483 virus, indicating that the 2 emergent cattle viruses could have a higher host adaption to humans.
A/Texas/37/2024 and other bovine-origin H5N1 isolates have shown dual binding affinity to α(2,3)- and α(2–6)-linked SA (7–10). However, multiple studies reported contradictory findings on the α(2–6)-linked receptor-binding specificity for different bovine-origin H5N1 isolates (11–13). One study (13) reported that A/bovine/Ohio/B24OSU-432/2024, which has a hemagglutinin amino acid sequence identical to that of virus examined elsewhere (10), preferentially binds to avian-type α(2,3) sialoside receptors. Genotypic differences cannot explain the binding affinity discrepancies (Appendix Tables 3, 4), but they might be explained by technical differences in the assays and origins of virus propagation in mammalian cells or embryonated eggs, which affect glycosylation of progeny viruses and hence receptor-binding specificity. Moreover, the binding affinity of H5N1/439 and H5N1/98638 aligned with their higher replication competences in bronchial tissues than the avian-origin H5N1/483 virus, which predominantly expresses an α(2–6)-linked SA. Those findings suggest that the 2 cattle-origin H5N1 viruses have a higher potential for human-to-human transmission than avian-origin H5N1.
Because illness severity of H5N1 infection is associated with induction of proinflammatory cytokines (14), we compared innate immune responses among the 5 viruses in human lung tissue. HPAI H5N1/483 infection induced substantially higher mRNA expressions of interferon alpha 1 (IFNA1), C-X-C motif chemokine ligand 10 (CXCL10), interferon-stimulated gene 15 (ISG15), interferon induced with helicase C domain 1 (IFIH1), and retinoic acid–inducible gene I (RIGI) than cattle-origin H5N1/439 and H5N1/98638. However, we observed a trend of elevated C-C motif chemokine ligand 2 (CCL2) and ligand 5 (CCL5) expression (Figure 3). H5N6/39715 infection induced substantially higher levels of ISG15, CCL2, CXCL10, IFIH1, and RIGI than H1N1pdm/415742, but H5N6/39715 had higher CXCL10 induction than H5N1/98638 and elevated ISG15 compared with H5N1/98638 and H5N1/439.
The 2 cattle-origin H5N1 isolates induced similar immune responses and substantially higher levels of IFNB1, ISG15, tumor necrosis factor, CCL2, CXCL10, IFIH1, and RIGI over the mock-infected tissues (Figure 3). Proinflammatory gene induction by H5N1/439 and H5N1/98638 viruses had higher trends than proinflammatory induction by H1N1pdm/415742, and substantial differences in ISG15, CXCL10, IFIH1, and RIGI. Prior animal studies demonstrated that H5N1/439 and H5N1/98638 and a human isolate were lethal to mice and ferrets (8,11). However, we detected lower immune response induction and fewer proinflammatory cytokines in human lung tissues, consistent with other studies (6), suggesting that those factors might contribute to reduced pathogenicity of cattle-origin H5N1 compared with avian H5N1.
Viral titers and influenza NP-positive cells demonstrated that cattle-origin H5N1/439 and H5N1/98638 strains are better adapted to human upper airway tissues than avian H5N1/483 and have similar replication abilities as H1N1pdm/415742 in human lung explants. The ability to bind α(2–6)-linked SA further indicates a shift of receptor affinities that are more compatible with upper respiratory tissues. Innate immune responses of H5N1/439 and H5N1/98638 viruses in human lung tissue fell between those triggered by H1N1pdm/415742 and H5N1/483 viruses, indicating that cattle H5N1 viruses could pose a human health risk. Defining how these strains infect human tissues and shape immune responses is critical for anticipating outbreaks and reducing zoonotic transmission risks. Because influenza viruses continually evolve across diverse avian and mammalian hosts, sustained research and surveillance remain essential to prevent human infections.
Ethics approval of the use of human tissues was granted by the institutional review board of University of Hong Kong (approval no. UW 20–862).
Dr. Hui is an assistant professor in the School of Public Health at the University of Hong Kong, China. Her research interests include risk assessment, understanding the pathogenesis of emerging respiratory viruses, and the development of therapeutic options for severe influenza diseases and coronavirus infections.
Acknowledgments
We thank Michael Kuan Yew Hsin and Lucius Kwok Fai Lee for providing human tissues. We also thank Rachel H.H. Ching and Kevin Fung for their technical support.
This work was supported by grants from the InnoHK initiative of the Innovation and Technology Commission of the Hong Kong Special Administrative Region Government and Theme-Based Research Scheme under the University Grants Committee of Hong Kong Special Administrative Region (project no. T11-712/19-N).
References
- Peacock TP, Moncla L, Dudas G, VanInsberghe D, Sukhova K, Lloyd-Smith JO, et al. The global H5N1 influenza panzootic in mammals. Nature. 2025;637:304–13. DOIPubMedGoogle Scholar
- Mostafa A, Nogales A, Martinez-Sobrido L. Highly pathogenic avian influenza H5N1 in the United States: recent incursions and spillover to cattle. Npj Viruses. 2025;3:54. DOIPubMedGoogle Scholar
- Nguyen TQ, Hutter CR, Markin A, Thomas M, Lantz K, Killian ML, et al. Emergence and interstate spread of highly pathogenic avian influenza A(H5N1) in dairy cattle in the United States. Science. 2025;388:
eadq0900 . DOIPubMedGoogle Scholar - Garg S, Reinhart K, Couture A, Kniss K, Davis CT, Kirby MK, et al. Highly pathogenic avian influenza A(H5N1) virus infections in humans. N Engl J Med. 2025;392:843–54. DOIPubMedGoogle Scholar
- Uyeki TM, Milton S, Abdul Hamid C, Reinoso Webb C, Presley SM, Shetty V, et al. Highly pathogenic avian influenza A(H5N1) virus infection in a dairy farm worker. N Engl J Med. 2024;390:2028–9. DOIPubMedGoogle Scholar
- Flagg M, Williamson BN, Ortiz-Morales JA, Lutterman TR, de Wit E. Comparison of contemporary and historic highly pathogenic avian influenza A(H5N1) virus replication in human lung organoids. Emerg Infect Dis. 2025;31:318–22. DOIPubMedGoogle Scholar
- Zhang X, Lam SJ, Chen LL, Fong CH, Chan WM, Ip JD, et al. Avian influenza virus A(H5N1) genotype D1.1 is better adapted to human nasal and airway organoids than genotype B3.13. J Infect Dis. 2026;233:e662–6. DOIPubMedGoogle Scholar
- Gu C, Maemura T, Guan L, Eisfeld AJ, Biswas A, Kiso M, et al. A human isolate of bovine H5N1 is transmissible and lethal in animal models. Nature. 2024;636:711–8. DOIPubMedGoogle Scholar
- Song H, Hao T, Han P, Wang H, Zhang X, Li X, et al. Receptor binding, structure, and tissue tropism of cattle-infecting H5N1 avian influenza virus hemagglutinin. Cell. 2025;188:919–929.e9. DOIPubMedGoogle Scholar
- Eisfeld AJ, Biswas A, Guan L, Gu C, Maemura T, Trifkovic S, et al. Pathogenicity and transmissibility of bovine H5N1 influenza virus. Nature. 2024;633:426–32. DOIPubMedGoogle Scholar
- Fabrizio TP, Kandeil A, Harrington WN, Jones JC, Jeevan T, Andreev K, et al. Genotype B3.13 influenza A(H5N1) viruses isolated from dairy cattle demonstrate high virulence in laboratory models, but retain avian virus-like properties. Nat Commun. 2025;16:6771. DOIPubMedGoogle Scholar
- Santos JJS, Wang S, McBride R, Adams L, Harvey R, Zhao Y, et al. Bovine H5N1 binds poorly to human-type sialic acid receptors. Nature. 2025;640:E18–20. DOIPubMedGoogle Scholar
- Chopra P, Ray SD, Page CK, Shepard JD, Kandeil A, Jeevan T, et al. Receptor-binding specificity of a bovine influenza A virus. Nature. 2025;640:E21–7. DOIPubMedGoogle Scholar
- Cheung CY, Poon LL, Lau AS, Luk W, Lau YL, Shortridge KF, et al. Induction of proinflammatory cytokines in human macrophages by influenza A (H5N1) viruses: a mechanism for the unusual severity of human disease? Lancet. 2002;360:1831–7. DOIPubMedGoogle Scholar
Figures
Cite This ArticleOriginal Publication Date: April 01, 2026
Table of Contents – Volume 32, Number 5—May 2026
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Michael C.W. Chan, School of Public Health, Li Ka Shing Faculty of Medicine, University of Hong Kong, 6/F William MW Mong Block, 21 Sasson Rd, Pokfulam, Hong Kong, China
Top