Volume 4, Number 4—December 1998
Dispatch
Introduction of HIV-2 and Multiple HIV-1 Subtypes to Lebanon
Abstract
HIV genetic variability, phylogenetic relationships, and transmission dynamics were analyzed in 26 HIV-infected patients from Lebanon. Twenty-five specimens were identified as HIV-1 and one as HIV-2 subtype B. The 25 strains were classified into six env-C2-V3 HIV-1 subtypes: B (n = 10), A (n = 11), C (n = 1), D (n = 1), G (n = 1), and unclassifiable. Potential recombinants combining parts of viral regions from different subtypes Aenv/Dpol/Agag, Genv/Apol and the unclassifiable-subtypeenv/ unclassifiable-subtypepol/Agag were found in three patients. Epidemiologic analysis of travel histories and behavioral risks indicated that HIV-1 and HIV-2 subtypes reflected HIV strains prevalent in countries visited by patients or their sex partners. Spread of complex HIV-subtype distribution patterns to regions where HIV is not endemic may be more common than previously thought. Blood screening for both HIV-1 and HIV-2 in Lebanon is recommended to protect the blood supply. HIV subtype data provide information for vaccine development.
Understanding the global genetic diversity of HIV is important for monitoring the spread of infection and developing effective vaccines. Currently, two principal genetic groups, designated M (main) and O (outlier), have been identified. M, which is highly prevalent, is further classified into 10 established envelope subtypes, A through J (1). Mosaic forms, which combine the genetic material from two distinct subtypes, have also been identified (1,2). HIV-1 subtype B predominates in Europe and the Americas, whereas HIV-1 non-subtype B strains dominate in sub-Saharan Africa. In contrast to HIV-1 infection, which is spread through all continents, HIV-2 is primarily restricted to West Africa and to population movements from or through this region. HIV-2 strains are classified into five subtypes, A through E; only subtypes A and B viruses are predominant (3).
HIV-subtype distribution patterns in regions where HIV is not endemic are not well known (4). Information about these distribution patterns is important for 1) implementing proper diagnostic assays to protect the blood supply; 2) providing basic information on the prevalence of viral sub-types for vaccine development; and 3) monitoring trends of viral spread. Two distribution patterns of HIV subtypes have been recognized in areas where the disease had not been endemic. The first pattern consists of a restricted number of HIV-1 viral subtypes or their combination with HIV-2. Examples include HIV-1 subtypes E and B in Thailand (5) and HIV-1 subtype C and HIV-2 in India (6). In contrast, the second pattern includes a large number of only HIV-1 subtypes as documented in Cyprus (A, B, C, F, and I) and the Philippines (A, B, C, D, and E) (7,8). However, information is limited as to how common the spread of this HIV-subtype pattern is to areas in which the disease is not endemic.
Lebanon was chosen for a genetic variability study for three reasons. First, HIV is not endemic in Lebanon, but its presence is slowly increasing (9,10). From 1984 (when the first case of AIDS was identified) (11) to 1996, 400 cases of HIV infection were reported to the Lebanon National AIDS Control Program (J.E. Mokhbat, pers. comm.). Second, Lebanon has a central geographic location, strong historic links, and frequent population movements to and from other countries. Third, the potential for introduction of multiple HIV subtypes is present given the large number of Lebanese expatriates who return to Lebanon after living and working in HIV-endemic regions such as Africa (mainly West Africa).
With informed consent, interviews and blood samples were obtained from 26 HIV-seropositive Lebanese citizens (17 men and 9 women) living in Lebanon at the time of the study. They were referred (in 1996) from both urban and rural areas to the Medical Center of the American University of Beirut or the Saint George Hospital in Beirut, Lebanon.
HIV-1 antibodies were discriminated from HIV-2 antibodies by a synthetic slot immunoblot composed of recombinant viral proteins and synthetic peptides (Genelavia Mixt, Peptilav 1-2, HIV Blot 2.2-Diagnostic Biotechnology and New LAV Blot II, Sanofi Diagnostic Pasteur, France).
Double-stranded viral DNA of nested polymerase chain reaction (PCR)-amplified env(C2-V3), pol (protease-prot), and gag (p24 fragment) from peripheral blood mononuclear cells were direct sequenced (12,13). Comparative phylogenetic analysis of two or three independent viral regions enables identification of a possible mosaic viral genome (2). The sequences were aligned by use of the CLUSTAL multiple-sequence alignment program (14). After regions containing gaps were eliminated, the aligned sequences were analyzed with the maximum likelihood method by using the fastDNAml program, which uses randomized data input and global rearrangement (15). Additionally, the neighbor-joining method (PHYLIP package version 3.5c [16] or TREECON software [17]) was used, with or without bootstrapping. The stability of the tree's topology was tested by pruning, which consists of removing one species from the alignment and rerunning the phylogenetic analysis. SIV-cpz sequences were used as outgroup. All specimens were also screened for HIV-2 DNA by nested-PCR amplification of viral HIV-2 prot gene. The outer PCR primers were DP20: 5'GACAGAGGACTTGCTGCA; nucleotide position: 2068-2083, HIV-2ROD, and DP21: 5'GGCCATTGTCTCAGTTTTGG; nucleotide position: 2435-2454. The inner primers were DP26a: 5'CACCTCAATTCTCTCTTTGGA; nucleotide position: 2082-2102; and DP27: 5'TAGATTTAATGACATGCCTAA; nucleotide position: 2360-2380. Analysis based on HIV-2 pol region is highly specific and allows classification of HIV-2 sequences into phylogenetic subtypes (3,18,19).
Of 26 patients, 25 were HIV-1 PCR-positive; the remaining patient (LE25) was HIV-2 PCR-positive and was classified as having subtype B infection (Figure 1). No dual infections with HIV-1 and HIV-2 strains were found.
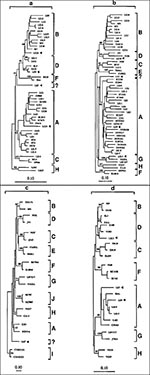
The comparative phylogenetic analysis of 25 HIV-1 specimens identified 23 cases with the same subtype assignment for the env and pol sequences and classified 22 as known subtypes A (n = 10), B (n = 10), C (n = 1), D (n = 1) (Figure 2). In contrast, the remaining specimen (LE7) consisted of distinctively divergent pol and env sequences (Figure 2a, b) as they failed to cluster with known standard HIV-1 sequences, including divergent env subtypes I and J (Figure 2c). Intragene recombination within pol and env sequences of specimen LE7 was not observed when nucleotide signature patterns in HIV-1 subtypes were compared. Thus, this strain might represent a potential novel HIV-1 subtype. However, gag sequence of this virus grouped into HIV-1 subtype A, and this association was supported by the bootstrap value of 90% (Figure 2d). Taken together, these results indicate that LE7 viral genome has a mosaic pattern involving elements of an unclassifiable-divergent strain and subtype A (unclassifiableenv/ unclassifiablepol/Agag). In addition, potential recombinant viruses combining viral env, pol, and gag from different subtypes were identified in two other specimens. Two known recombinant patterns, Genv/Apol and Aenv/Dpol/Agag, have been found in patients LE24 and LE9, respectively. However, a potential recombinant is sometimes difficult to recognize, especially between subtypes B and D viruses, which are relatively closely related; specimen LE22 illustrates such a situation. The protease gene sequence, the most outside branch in this clade, grouped weakly with subtype B (Figure 2a). This association, however, was not stable by analysis that used pruning techniques and not supported by bootstrap analysis (only 48% value as opposed to 72% without this sequence). These data suggest that it is unlikely that LE22 prot is subtype B, and because both env and gag sequences clustered into subtype D, we have classified this sample as subtype D.
To better understand the transmission dynamics and temporal introduction of these HIV variants, we analyzed patients' data (demographics, behavioral risk, modes of transmission, and travel histories) (Table).
Of eight male patients with subtype B HIV-1 infection, six traveled to Europe, the Americas, Asia (Thailand), or different locations in the Middle East. All reported sexual contacts abroad either with commercial sex workers or homosexual or bisexual men. Of the remaining two patients, one traveled selectively to Africa and had sexual contacts in Côte d'Ivoire and Nigeria, whereas the other reported no travel outside Lebanon and had no sexual contacts with commercial sex workers. In contrast, all nine male patients infected with non-subtype B HIV-1, including three mosaic viruses, traveled to Africa or the United Arab Emirates and reported sexual contacts with commercial sex workers.
The behavioral risks of the female patients differed substantially from those of the male patients. All female patients were married, and none reported high-risk activities or contacts with commercial sex workers. Moreover, all female patients (except one [LE12], who had acquired HIV-1 infection from blood transfusion) reported that their spouses were HIV-infected and traveled abroad with them or alone.
To determine if the female patients acquired HIV infection from their spouses, we analyzed available epidemiologic data on the spouses of three female patients LE17, LE19, and LE13. The female patients LE17 and LE19 were spouses of patients LE18 and LE23, respectively. These two couples, LE17/LE18 and LE19/LE23, were infected with HIV-1 subtype A and subtype B viruses, respectively (Figure 2b). In further detailed pairwise analysis of viral DNA isolated from the husband and wife, a small nucleotide divergence of 3.1% within env and of 0.6% for both prot and gag (13,20) showed that in the first couple viral strains were closely related. These data are concordant with epidemiologic findings, which strongly indicate that the husband was the source of infection because of his high-risk sexual activities with commercial sex workers during travels to the United Arab Emirates. His wife did not travel outside Lebanon and did not report activities that would have placed her at risk for HIV-1 infection. In the second couple (LE19/LE23), however, only weak relation between viral strains was observed (nucleotide divergence of 9.8% within env and 2% within prot). This could be due to many factors including 1) the time that elapsed from infection; 2) transmission of different quasispecies to the spouse; 3) detection of only major quasispecies by direct sequencing, which differed in the couple; and 4) a different source of HIV-1 infection in the sex partner. The latter hypothesis was not supported by epidemiologic data, which indicated that only the husband was involved in high-risk sexual activities in and outside Lebanon. The third female patient, LE25, was infected with HIV-2 subtype B strain. Her husband, who died of AIDS-related complications, had traveled to Côte d'Ivoire, a region in which this HIV variant is endemic (21). These data suggest that very likely the female patients acquired infections from their HIV-infected spouses.
The molecular and epidemiologic data indicate multiple HIV-1 introductions to Lebanon. First, most male patients were infected abroad with the HIV-1 strains prevalent in the geographic region they visited. Second, the topology of HIV-1 env subtype A demonstrated that viral sequences of patients who had sexual contacts with persons from West Africa clustered in the branch with marker sequences from Côte d'Ivoire. In contrast, HIV-1 sequences of patients who reported contacts with persons from the United Arab Emirates (LE9 and LE18) clustered in another branch with Ugandan/Kenyan marker sequences (Figure 2b). Independent branching of HIV-1 env sequences of subtype A from West and East Africa had been observed (22). Third, the structures of two mosaic viral genomes and epidemiologic data of patients suggested that they were infected with potential recombinant viruses, which had been identified in two geographic regions. Briefly, patients LE24 and LE9, who reported sexual contacts with commercial sex workers in Nigeria or the United Arab Emirates, respectively, have been infected with mosaic viruses Genv/Apol and Aenv/Dpol/Agag. Such recombination patterns were found in West-Central Africa and East Africa, respectively (23,24). Finally, analysis of env sequences from Lebanese patients revealed a relatively high intra-subtype genetic diversity within subtype B (average 14.9%, range 4.2% to 25.6%) and subtype A (15.5%, range 2.5% to 28.9%. Epidemiologic data and a low prevalence of HIV-1 infection suggest multiple introductions (rather than long existence) of genetically diverse HIV-1 strains to Lebanon.
The results presented in this report provide new information on HIV genetic variation in Lebanon. The identification of HIV-2 and 6 env HIV-1 subtypes, including three distinct HIV-1 mosaics, confirms high heterogeneity of HIV variants in Lebanon. Given the limited number of samples tested, finding both HIV-2 and multiple HIV-1 subtypes is remarkable and indicates that the distribution pattern of HIV variants resembled that of Cyprus or the Philippines rather than that of Thailand or India. These data indicate that the spread of complex HIV-subtype distribution patterns involving a large number of distinct HIV-1 and HIV-2 strains to countries without endemic HIV may be more common than previously thought.
The HIV-2 infection detected in this study, the first documented in Lebanon, reiterates the importance of appropriate screening methods for detecting both HIV-1 and HIV-2 to protect the blood supply. Our findings provide preliminary information for monitoring the spread of HIV variants in Lebanon and developing subtype-specific subunit vaccines.
Danuta Pieniazek is a molecular biologist in the HIV/AIDS and Retroviral Branch, CDC. Her research interests include implications of mixed HIV infections for the evolution of the HIV/AIDS epidemic and for vaccine development; her areas of expertise include developing molecular assays for detection of novel HIV strains and the molecular epidemiology of HIV.
Acknowledgments
We thank Priscilla Swanson for her help in sequencing, Eve Lackritz for critical review of the manuscript, and Mr. Issam Khneisser, American University of Beirut, for processing blood specimens .
Part of this work was done at the Institute of Tropical Medicine, Antwerp, Belgium, and was sponsored by grant numbers G 3301.96 and G 30134.97 of the Belgian Nationaal Fonds voor Wetenschappelijk Onderzoek.
References
- Leitner T. Genetic subtypes of HIV-1. In: Human Retroviruses and AIDS. Edited by Myers G, Foley B, Mellors JW, Korber B, Jeang KT, Wain-Hobson S. Theoretical Biology and Biophysics, Los Alamos National Laboratory, Los Alamos (NM): 1996:III28-III40.
- Robertson DL, Sharp PM, McCutchan FE, Hahn BH. Recombination in HIV-1. Nature. 1995;374:124–6. DOIPubMedGoogle Scholar
- Gao F, Yue L, Robertson DL, Hill SC, Hui H, Biggar RJ. at al. Genetic diversity of human immunodeficiency virus type 2: evidence for distinct sequence subtypes with differences in virus biology. J Virol. 1994;68:7433–47.PubMedGoogle Scholar
- Hu DJ, Dondero TJ, Rayfield MA, George R, Schochetman G, Jaffe HW, The emerging diversity of HIV: the importance of global surveillance for diagnostics, research, and prevention. JAMA. 1996;275:210–6. DOIPubMedGoogle Scholar
- Ou C-Y, Takebe Y, Weniger BG, Luo CC, Kalish ML, Auwanit W, Independent introduction of two major HIV-1 genotypes into distinct high-risk populations in Thailand. [Erratum in Lancet 1993;342:250]. Lancet. 1993;341:1171–4. DOIPubMedGoogle Scholar
- Dietrich U, Maniar JK, Rubsamen-Waigmann H. The epidemiology of HIV in India. Trends Microbiol. 1995;3:17–21. DOIPubMedGoogle Scholar
- Kostrakis LG, Bagdades E, Cao Y, Zhang L, Dimitriou D, Ho DD. Genetic analysis of human immunodeficiency virus type 1 strains from patients in Cyprus: identification of a new subtype designated subtype I. J Virol. 1995;69:6122–30.PubMedGoogle Scholar
- Paladin FJ, Monzon OT, Tsuchie H, Aplaska MRA, Learn GH Jr, Kurimura T. Genetic subtypes in the Philippines. AIDS. 1998;12:291–300. DOIPubMedGoogle Scholar
- Naman RE, Mokhbat JE, Abdelnoor AM, Irani-Hakimeh N, Alami SY, Nassif RE. Seroepidemiology of the human immunodeficiency virus-1 infections in Lebanon. Leb Science Bull. 1989;5:9–14.
- Tawilah J, Adib SM. Epidemiological characteristic of HIV/AIDS infection of Beirut, Lebanon. X International Conference on AIDS. Tokyo, August 1994 [abstract PC0055].
- Mokhbat JE, Ibrahim N, Abdelkarim F, Kuleilat-Shatila M, Salem Z. The acquired immunodeficiency syndrome: report of the first case in Lebanon and a review of the literature. Leb Med J. 1985;35:295–311.
- Janini LM, Pieniazek D, Peralta JM, Schechter M, Tanuri A, Vicente AC, Identification of single and dual infections with distinct subtypes of human immunodeficiency virus type 1 by using restriction fragment length polymorphism analysis. Virus Genes. 1996;13:69–81. DOIPubMedGoogle Scholar
- Janini LM, Tanuri A, Schechter M, Peralta M, Vicente ACP, Dela Torre N, Horizontal and vertical transmission of human immunodeficiency virus type 1 dual infections caused by viruses of subtypes B and C. J Infect Dis. 1998;177:227–31. DOIPubMedGoogle Scholar
- Higgings DG, Sharp PM. Fast and sensitive multiple sequence alignment on a microcomputer. Comput Appl Biosci. 1989;5:151–3.PubMedGoogle Scholar
- Larsen N, Olsen GJ, Maidak BN, McCaughey MJ, Oberbeek R, Macke TJ, The ribosomal data base project. Nucleic Acids Res. 1993;21:3021–3. DOIPubMedGoogle Scholar
- Felsenstein J. PHYLIP-phylogeny interference package (version 3.2). Cladistics. 1989;5:164–6.
- Van de Peer Y, De Wachter R. TREECON for Windows: a software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment. Comput Appl Biosci. 1994;10:569–70.PubMedGoogle Scholar
- Pieniazek D, Peralta JM, Ferreira JA, Krebs JW, Owen SM, Sion FS, Identification of mixed HIV-1/HIV-2 infections in Brazil by polymerase chain reaction. AIDS. 1991;5:1293–9. DOIPubMedGoogle Scholar
- Takehisa J, Osei-Kwasi M, Ayisi NK, Hishida O, Mura T, Igarashi T, Phylogenetic analysis of HIV type 2 in Ghana and intrasubtype recombination in HIV type 2. AIDS Res Hum Retroviruses. 1997;13:621–3. DOIPubMedGoogle Scholar
- Ou CY, Ciesielski C, Myers G, Bandea CI, Luo CC, Mullins JI, Molecular epidemiology of HIV transmission in a dental practice. Science. 1992;256:1167–71. DOIGoogle Scholar
- Pieniazek D, Victor S, Ellenberger D, Janini LM, Ramos A, Greenberg A, Genetic analysis of HIV type 2 strains in Cote d'Ivoire: evidence for endemicity of HIV-2 subtype B. 4th Conference on Retroviruses and Opportunistic Infections. Washington D.C., 1997 Jan [abstract 171].
- Rayfield M, Downing RG, Baggs J, Hu DJ, Pieniazek D, Luo C-C, A molecular epidemiologic survey of HIV in Uganda. AIDS. 1998;12:521–7. DOIPubMedGoogle Scholar
- Abimicu AG, Stern TL, Zwandor A. Subgroup G HIV-type 1 isolates from Nigeria. AIDS Res Hum Retroviruses. 1994;10:1581–3. DOIPubMedGoogle Scholar
- Desselberger U, Garnegie G, Baker C. Phylogenetic analysis of partial gag and env sequences of HIV-1 isolates collected in England and Uganda, 1993-1995: evidence for A/D recombinants in Uganda. 16th Annual Meeting of the American Society for Virology. Bozeman (MT): July 1997 [abstract P2-1].
Figures
Table
Cite This ArticleTable of Contents – Volume 4, Number 4—December 1998
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Danuta Pieniazek, HIV/Retrovirus Diseases Branch, Division of AIDS, STD, and TB Laboratory Research, CDC, 1600 Clifton Road, Mail Stop G19, Atlanta, GA 30333, USA; fax: 404-639-1010
Top