Volume 5, Number 2—April 1999
Synopsis
Clonal Differences among Erythromycin-Resistant Streptococcus pyogenes in Spain
Cite This Article
Citation for Media
Abstract
The aim of this study was to determine whether the high levels of erythromycin resistance in Streptococcus pyogenes found in Spain are due to the introduction and spread of one or more clones. Phenotypic and genotypic techniques were used to characterize all erythromycin-resistant S. pyogenes (ErR) isolated in Gipuzkoa, Spain, in the last 10 years and 128 ErR isolated in Vitoria and Madrid during 1996. Of 437 ErR, 97% had the M phenotype; all 283 of the strains studied had the mefA determinant of resistance. After biotyping, T serotyping, emm typing, and genotyping, four major clones were detected. Clones B (biotype I, type T4, emm4, pulsed-field gel electrophoresis [PFGE] II) and D (biotype V, type T8.25, emm75, PFGE IV) comprised 78.8% of all ErR. The resistance of S. pyogenes to erythromycin was mainly due to an efflux mechanism of resistance (M phenotype); few clones were responsible for it.
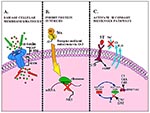
The Lancefield group A streptococci (Streptococcus pyogenes), major causative agents of human disease (1), can produce both mild (e.g., pharyngitis) and severe (e.g., life-threatening "toxic shock-like syndrome," necrotizing fasciitis) infections. During the last few years, erythromycin-resistant S. pyogenes (ErR) has been reported in different parts of the world (2-4). Two distinct mechanisms of erythromycin resistance are described among group A streptococci. One consists of target-site modification by erm methylase (5,6) strains that express the MLSb phenotype of resistance; the other (recently described) consists of an active drug efflux that pumps 14- and 15-membered macrolides out of the cell (7). This novel mechanism of macrolide resistance is encoded by the gene mefA (8), and strains show the M phenotype. In the last few years, increased resistance to erythromycin in S. pyogenes has been detected in Spain (9-11). Therefore, we performed an epidemiologic investigation to determine the biotypes, serotypes (T-agglutination patterns), emm types, and pulsed-field gel electrophoresis (PFGE) patterns of chromosomal DNA and their relationship to macrolide resistance.
Sources of Bacterial Isolates
From 1988 to 1997, 2,561 nonduplicated isolated strains of S. pyogenes were collected from throat swabs and extratonsilar samples at the Nuestra Señora de Aránzazu Hospital and at primary-care centers in two districts of Gipuzkoa (approximately 300,000 residents). In 1996, two other samples were collected and included in this study; 33 ErR strains from Vitoria (Hospital Txagorritxu) and 95 from Madrid (Centro de Especialidades Argüelles). Gipuzkoa Province (San Sebastian is its capital) is located in the northeastern area of the Basque country of Spain, bordered by the Cantabric Sea and France to the north; Madrid is located in the center of Spain (415 km from San Sebastian); and Vitoria (in Alava Province) is located in the north of Spain (110 km from San Sebastian).
Group A streptococci were identified by colony morphology, beta-hemolysis on blood agar, and commercial latex-agglutination techniques (Streptex, Wellcome, Dartford, UK, or Phadebact, Boule Diagnostics AB, Huddinge, Sweden). The confirmation of S. pyogenes and biotyping were done with a commercially available identification system: rapid ID 32 STREP (BioMerieux, La Balme-les-Grottes, France). Biotyping was performed according to Bouvet et al. (12).
All S. pyogenes were tested for susceptibility to erythromycin and other antibiotics by broth microdilution. ErR strains were restudied by agar dilution and by agar diffusion (erythromycin induction of resistance) to determine macrolide and lincosamine resistance phenotypes (11).
T-protein types were determined by slide agglutination of trypsin-digested suspensions of bacteria with rabbit type-specific antiserum (SEVAC, Prague) (13).
The emm gene type of T-serotyped strains was determined by polymerase chain reaction (PCR)-enzyme-linked immunosorbent assay (ELISA), as described by Saunders et al. (14). Capture probes for the emm gene not described by Saunders et al. were selected from the DNA sequences encoding the N-terminal hypervariable region of strains of types emm2, emm9, emm48, and emm75 (GenBank accession nos. X56608, U12002, U11961, and U11993).
PFGE was done (15) with the following modifications. Cells were resuspended to an optical density (OD)560nm = 1.0, and 4 ml of the adjusted suspension was centrifuged. Pelleted cells were resuspended in 200 µl of Pett IV buffer (10 mM Tris, pH 7.2, 20 mM NaCl, 50 mM EDTA) and mixed with 100 µl of 2% low-melting agarose; incubation of plugs with lysozyme solution (1 mg/ml) was reduced to 3 hours. Slices of plugs were digested at 50ºC for 16 hours with 30 UI of the enzyme SfiI since it has proven satisfactory in differentiating DNA fragments of S. pyogenes (15,16). Digested inserts were electrophoresed by using a CHEF-DR III apparatus (BioRad), along with DNA size standards (BioRad) under the following conditions: 22 hours with an initial switch time of 20 seconds, rising on linear ramp to 75 seconds at 6V/cm, with an included angle of 120ºC. Gels were stained with ethidium bromide and visualized under UV light with Imagestore 5000 ver.7.12 (Ultra-Violet Products Ltd, Cambridge, England). Similarities among PFGE patterns were established using the Dice coefficient and Lane-Manager 2.1 (TDI, Madrid, Spain) commercial computer software.
A clone was defined as a group of strains expressing both the same characteristic phenotype and genotype (PFGE pattern similarity > 90%).
We studied 2,561 strains of S. pyogenes isolated from 1988 to 1997 in Gipuzkoa; 309 (12.1%) were resistant to erythromycin. A report of ErR in Gipuzkoa from 1984 to 1996 (11) showed that until the end of 1990, erythromycin resistance in S. pyogenes was low (1.2%, 13 of 1,060); after 1990, resistance increased, reaching 34.8% (87 of 250) of all S. pyogenes isolated in 1995. In 1997, resistance decreased to 13.7% (57 of 417). In Madrid and Vitoria, resistance to erythromycin in 1996 was 22.4% (126 of 563) and 31.6% (43 of 136), respectively.
Among the ErR strains isolated in Gipuzkoa, two phenotypes of resistance were found: 8 (2.6%) strains showed the classic MLSb phenotype, while the other 301 (97.4%) strains expressed the M phenotype, as shown by susceptibility testing and confirmed by the presence of the mefA gene (11). Of the 128 ErR isolated in Madrid and Vitoria, 5 (3.9%) strains (all from Madrid) displayed the MLSb phenotype, while the remaining 123 (96.1%) showed the M phenotype. The presence of the mefA gene was searched for in 283 ErR with the M phenotype; it was detected in all.
Among the 424 ErR with the M phenotype, only four biotypes (of 10 possible) were identified—biotypes I, II, III, and V. In Gipuzkoa, biotype III was the only biotype found until 1990; between 1991 and 1997, biotypes I and V comprised 275 (93.8%) of the 293 resistant strains. Seven T-agglutination patterns were found in Gipuzkoa, each one correlating with an emm type except for TB3264 (T1 emm1, T2 emm2, T4 emm4, T8.25 emm75, T12 emm12, and T28 emm28). TB3264 biotype III was emm2, but TB3264 biotype I was not typeable with any of the 14 emm types assayed. Until 1990, type T12 emm12 was the only type found. Between 1991 and 1997, type T4 emm4 and T8.25 emm75 comprised 92.2% (270 of 293) of all isolates with the M phenotype of resistance. In Vitoria and Madrid, T4 emm4 and T8.25 emm75 types were also the most frequently found.
Fifteen different PFGE patterns were found among the 424 M phenotype ErR; 92% of these strains belonged to four patterns (clones A-D) (Table, Figure 1). Among ErR with the MLSb phenotype, eight PFGE patterns were found. Each biotype/T-serotype/emm-type combination corresponded with one PFGE pattern except on three occasions. Three different PFGE patterns that could be established among ErR belonged to biotype III/T12/emm12, two PFGE patterns belonged to I/T4/emm4, and another two patterns belonged to V/T8.25/emm75. The types of other clones, their annual distribution, and a dendrogram showing their similarities are given in the Table and Figure 2.
The genetic relationship of ErR and erythromycin-sensitive S. pyogenes with the same biotype and T serotype was analyzed by PFGE. In a sample of 360 erythromycin-sensitive S. pyogenes, biotypes I and III were the most frequent (66.7%); T1 and T28 were the most frequent T serotypes (30.6%). Only 8 (2.2%) erythromycin-sensitive strains of biotype I, serotype T4 were found. None of the 19 T8.25 erythromycin-sensitive strains found were biotype V (18 biotype II, emm 75 and 1 biotype I, emm nontypeable). Infrequent biotype and T-serotype combinations among ErR, such as I/T1 and I/T28, were frequently found among erythromycin-sensitive strains. Similarities between PFGE patterns of most of the erythromycin-sensitive and -resistant strains with the same biotype and T-serotype combination was less than 75%. Exceptions to this were several III/T12, I/T1, and I/T28 erythromycin-sensitive strains that had a close similarity (>90%) with ErR of the same biotype and T-serotype combination.
Four major clones of ErR were detected: clone A (T12, emm12, biotype III, PFGE I), which was present in Gipuzkoa until 1990; clone B (T4, emm4, biotype I, PFGE II), which was introduced in 1991; and clones C (T4, emm4, biotype I, PFGE III) and D (T8,25, emm75, biotype V, PFGE IV), which were introduced in 1995 and persisted during 1996 and 1997. In Madrid and Vitoria, 89.4% (110) of the 123 M-phenotype strains isolated in 1996 belonged to clones B and D.
In 1990, a new phenotype of erythromycin resistance (first designated NR and later M) was found in group A streptococci in Finland (7,17). ErR with the M phenotype had a low level of erythromycin resistance (8 mg/l to 16 mg/l) and showed cross-resistance with the 14- and 15-membered macrolides; however, they showed the same susceptibility to the 16-membered macrolides and to clindamycin as the erythromycin-susceptible strains (11). This M phenotype was prevalent among S. pyogenes in Europe and was the predominant phenotype of resistance among ErR isolated in Finland (18,19), Sweden (20), Austria (21), and Spain (9-11). In Italy, a high prevalence of erythromycin-resistant S. pyogenes was reported, but the prevalence of the M-phenotype strains among ErR varied by geographic area (22-24). Among the 437 ErR in our study in Spain, 424 (97%) showed the M phenotype; the mefA gene was detected and studied in 283 of these strains.
The epidemiologic surveillance of S. pyogenes can be done by using phenotypic or genotypic methods or both (as we did). Biotyping does not need specialized personnel or equipment, and it is useful, in combination with serotyping methods, for a first approximation of the epidemiologic characterization of S. pyogenes. In this study, the most prevalent T serotypes among ErR were infrequent among erythromycin-sensitive strains and vice versa. M serotyping is a classic typing method with at least 74 types recognized, but because it is a very specialized method and reagents are not available commercially, it is restricted to a few reference centers. A rapid PCR-ELISA to determine the emm gene type was an accessible alternative to serology for M-antigen typing.
Although the discriminatory power of biotyping and serotyping is considered poor because different genotypes may express the same phenotypic characteristics (15,25,26), these tests were of great value in our epidemiologic surveillance. The biotype and T-serotype combination was able to discriminate between ErR and erythromycin-sensitive strains and delimited most clones among ErR.
Genomic typing methods have rarely been used in characterizing the epidemiology of noninvasive S. pyogenes. Among these methods, restriction endonuclease analysis of genomic DNA (REA), random amplified polymorphic DNA (RAPD), ribotyping, PFGE of chromosomal DNA, and DNA sequence analysis have been used with varying degrees of success (15,16,19,26-28). PFGE patterns confirmed the results of biotyping and serotyping and further distinguished among isolates within the same biotype and T-serotype combination. However, PFGE is a complex method—results take at least 4 days to obtain—and expensive equipment and specialized personnel are needed.
Although the polyclonal nature of the ErR strains was established in this study and previously (18), most ErR belonged to only a few clones. In Finland, 91% of the ErR isolated in 1994 were serotype T4 M4 and 88% constituted one clone by RAPD and REA (19). In our study, many clones were detected during the 10-year period. Two of the four main clones comprised 78.8% of ErR. The clonal distribution of ErR in Spain could be due to the introduction and spread of ErR from other locations or to mutations in S. pyogenes previously present in our environment. ErR with the M phenotype already existed in Gipuzkoa before 1990, and serotype T12 was the only one found. From 1980 to 1988, serotype T12 was predominant among ErR with the M phenotype in Sweden (20). In Gipuzkoa, the first serotype T4 ErR strain was isolated in 1991; until the end of 1994, 93.8% of ErR isolated belonged to the same clone (clone B: biotype I, type T4, emm4, PFGE II). Only three biotype I serotype T4 erythromycin-sensitive strains were detected before 1991. Clone B probably did not emerge in Gipuzkoa from a mutation of one of these uncommon sensitive strains. Apart from the strains described in Finland, Sweden, and Spain, strains with M phenotype were isolated in Great Britain before 1990 (4,29). Among these British and Finnish strains, type T4 M4 was frequently isolated (4,18,29). In a 1986 outbreak of 10 associated cases in Somerset, Great Britain, isolates were group A, type M4 and resistant to erythromycin (MIC 8 mg/l) but sensitive to clindamycin (29). We do not know whether the strains of clone B isolated in Spain are the same as the ErR type T4 M4 found in Great Britain and Finland before 1991, but it is probable. In Italy, serotypes T4 and T8.25 were found among M-phenotype strains (22), which suggests that a few clones have spread across Europe and caused a regional epidemic. No erythromycin-sensitive strains were detected in Gipuzkoa with the same biotype/T-type/emm type combination as clone D.
We believe that clones B and D, the most frequent among ErR in Spain, were of epidemic origin and that clone B probably came from northern or western Europe.
Dr. Perez-Trallero is a clinical microbiologist and infectious disease consultant. He is head of the Microbiology Department at Complejo Hospitalario Donostia and assistant professor of Preventive Medicine and Public Health at the Facultad de Medicina at the Basque Country University. His research focuses on antimicrobial resistance, epidemiology of transmissible diseases, and methods for their prevention and control.
References
- Bisno A. Streptococcus pyogenes. In: Mandell GL, Bennett JE, Dolin R, editors. Principles and practice of infectious diseases. 4th ed. New York: Churchill Livingstone; 1995. p. 1786-99.
- Holmstrom L, Nyam B, Rosengren M, Wallander S, Ripa T. Outbreaks of infection with erythromycin-resistant group A streptococci in child day care centers. Scand J Infect Dis. 1990;22:179–85. DOIPubMedGoogle Scholar
- Stingmore N, Francis GRJ, Toohey M, McGenchie DB. The emergence of erythromycin resistance in Streptococcus pyogenes in Fremantle, Western Australia. Med J Aust. 1989;150:626–31.PubMedGoogle Scholar
- Philips G, Parratt D, Orange GV, Harper I, McEwan H, Young N. Erythromycin-resistant Streptococcus pyogenes. J Antimicrob Chemother. 1990;25:723–4. DOIPubMedGoogle Scholar
- Leclerq R, Courvalin P. Bacterial resistance to macrolide, lincosamide, and streptogramin antibiotics by target modification. Antimicrob Agents Chemother. 1991;35:1267–72.PubMedGoogle Scholar
- Sepälä H, Skurnik M, Soini H, Roberts MC, Huovinen P. A novel erythromycin resistance methylase gene (ermTR) in Streptococcus pyogenes. Antimicrob Agents Chemother. 1998;42:257–62. DOIPubMedGoogle Scholar
- Sutcliffe J, Tait-Kamradt A, Wondrack L. Streptococcus pneumoniae and Streptococcus pyogenes resistant to macrolides but sensitive to clindamycin: a common resistance pattern mediated by an efflux system. Antimicrob Agents Chemother. 1996;40:1817–24.PubMedGoogle Scholar
- Clancy J, Petitpas J, Dib-Hajj F, Yuan W, Cronan M, Kamath AV, Molecular cloning and functional analysis of a novel macrolide-resistance determinant, mefA, from Streptococcus pyogenes. Mol Microbiol. 1996;22:867–79. DOIPubMedGoogle Scholar
- Garcia-Bermejo I, Cacho J, Orden B, Alos JI, Gomez-Garces JL. Emergence of erythromycin-resistant, clindamycin-susceptible Streptococcus pyogenes isolates in Madrid, Spain. Antimicrob Agents Chemother. 1998;42:989–90.PubMedGoogle Scholar
- Orden B, Perez-Trallero E, Montes M, Martínez R. Erythromycin resistant Streptococcus pyogenes in Madrid. Pediatr Infect Dis J. 1998;17:470–3. DOIPubMedGoogle Scholar
- Perez-Trallero E, Urbieta M, Montes M, Ayestaran I, Marimon JM. Emergence of Streptococcus pyogenes resistant to erythromycin in Gipuzkoa, Spain. Eur J Clin Microbiol Infect Dis. 1998;16:25–31. DOIGoogle Scholar
- Bouvet A, Gelsin P, Kriz-Kuzemenska P, Blanc V, Devine C, Grimont F. Restricted association between biotypes and serotypes within Group A Streptococci. J Clin Microbiol. 1994;32:1312–7.PubMedGoogle Scholar
- Johnson DR, Kaplan EL, Sramek J, Bicova R, Havlicek J, Havlickova H, Determination of T-protein agglutination patterns. In: Laboratory diagnosis of group A streptococcal infections. Geneva: World Health Organization; 1996. p. 37-41.
- Saunders NA, Hallas G, Gaworzewska ET, Metherell L, Efstratiou A, Hookey JV, PCR-enzyme-linked immunosorbent assay and sequencing as an alternative to serology for M-antigen typing of Streptococcus pyogenes. J Clin Microbiol. 1997;35:2689–91.PubMedGoogle Scholar
- Single LA, Martin DR. Clonal differences within M-types of the group A Streptococcus revealed by pulsed field gel electrophoresis. FEMS Microbiol Lett. 1992;91:85–90. DOIGoogle Scholar
- Upton M, Carter PE, Morgan M, Edwards GF, Pennington TH. Clonal structure of invasive Streptococcus pyogenes in Northern Scotland. Epidemiol Infect. 1995;115:231–41. DOIPubMedGoogle Scholar
- Seppala H, Nissinen A, Yu Q, Huovinen P. Three different phenotypes of erythromycin-resistant Streptococcus pyogenes in Finland. J Antimicrob Chemother. 1993;32:885–91. DOIPubMedGoogle Scholar
- Seppala H, Nissinen A, Jarvinen H, Houvinen S, Henriksson T, Herva E, Resistance to erythromycin in Group A Streptococci. N Engl J Med. 1992;326:292–7.PubMedGoogle Scholar
- Kataja J, Huovinen P, Muotiala A, Vuopio-Varkila J, Efstratiou A, Hallas G, Clonal spread of group A streptococcus with the new type of erythromycin resistance. Finnish Study Group for Antimicrobial Resistance. J Infect Dis. 1998;177:786–9. DOIPubMedGoogle Scholar
- Jasir A, Schalen C. Survey of macrolide resistance phenotypes in Swedish clinical isolates of Streptococcus pyogenes. J Antimicrob Chemother. 1998;41:135–7. DOIPubMedGoogle Scholar
- Kriebernegg I, Feierl G, Grisold A, Marth E. In vitro susceptibility of group A beta-haemolytic streptococci (GABHS) to penicillin, erythromycin, clarithromycin and azithromycin in Styria, Austria. Zentralbl Bakteriol. 1998;287:33–9.PubMedGoogle Scholar
- Cornaglia G, Ligozzi M, Mazzariol A, Valentini M, Orefici G, the Italian Surveillance Group for Antimicrobial Resistance, . Rapid increase of resistance to erythromycin and clindamycin in Streptococcus pyogenes in Italy, 1993-1995. Emerg Infect Dis. 1996;2:339–42. DOIPubMedGoogle Scholar
- Cocuzza C, Blandino G, Mattina R, Nicoletti F, Nicoletti G. Antibiotic susceptibility of group A streptococci in 2 Italian cities: Milano and Catania. Microb Drug Resist. 1997;3:379–84. DOIPubMedGoogle Scholar
- Borzani M, De Luca M, Varotto F. A survey of susceptibility to erythromycin amongst Streptococcus pyogenes isolates in Italy. J Antimicrob Chemother. 1997;40:457–8. DOIPubMedGoogle Scholar
- Haase AM, Melder A, Mathews JD, Kemp DJ, Adams M. Clonal diversity of Streptococcus pyogenes within some M-types revealed by multilocus enzyme electrophoresis. Epidemiol Infect. 1994;113:455–62. DOIPubMedGoogle Scholar
- Upton M, Carter PE, Orange G, Pennington TH. Genetic heterogeneity of M type 3 group A streptococci causing severe infections in Tayside, Scotland. J Clin Microbiol. 1996;34:196–8.PubMedGoogle Scholar
- Seppala H, Vuopio-Varkila J, Osterblad M, Jahkola M, Rummukaien M, Holm SE, Evaluation of methods for epidemiologic typing of group A streptococci. J Infect Dis. 1994;169:519–25.PubMedGoogle Scholar
- Bert F, Branger C, Lambert-Zechovsky N. Pulsed-field gel electrophoresis is more discriminating than multilocus enzyme electrophoresis and random amplified polymorphic DNA analysis for typing pyogenic streptococci. Curr Microbiol. 1997;34:226–9. DOIPubMedGoogle Scholar
- Scott RJD, Naidoo J, Lightfoot NF, George RC. A community outbreak of group A beta-haemolytic streptococci with transferable resistance to erythromycin. Epidemiol Infect. 1989;102:85–91. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 5, Number 2—April 1999
| EID Search Options |
|---|
|
|
|
|
|
|