Volume 6, Number 6—December 2000
Research
Hantavirus Pulmonary Syndrome Associated with Monongahela Virus, Pennsylvania
Abstract
The first two recognized cases of rapidly fatal hantavirus pulmonary syndrome in Pennsylvania occurred within an 8-month period in 1997. Illness in the two patients was confirmed by immunohistochemical techniques on autopsy material. Reverse transcription-polymerase chain reaction analysis of tissue from one patient and environmentally associated Peromyscus leucopus (white-footed mouse) identified the Monongahela virus variant. Physicians should be vigilant for such Monongahela virus-associated cases in the eastern United States and Canada, particularly in the Appalachian region.
Sin Nombre virus (SNV) was isolated and characterized as the cause of the 1993 cluster of 17 cases of hantavirus pulmonary syndrome (HPS) in the southwestern United States (1,2). HPS cases in other states were predicted because of the widespread distribution of the rodent hosts of hantavirus (2). Subsequent reports described human infection with and rodent carriage of novel genomic variants of hantavirus in New York (3), Florida (4), and Louisiana (5). Each of the four initially characterized strains of hantavirus causing HPS in North America is carried by a primary rodent host species, although spillover to other rodents in the area can occur (6-9).
We describe the first two recognized cases of HPS acquired in Pennsylvania; both were fatal and at least one was caused by infection with the newly characterized Monongahela variant of hantavirus (10). This variant can be present in Peromyscus leucopus mice, in addition to P. maniculatus nubiterrae with which they often share microhabitat.
Case 1. A 40-year-old, previously healthy crossbow hunter was taken by ambulance to a Lehigh County, Pennsylvania, hospital in November 1997, with complaints of back muscle pain, dizziness, diarrhea, fever, and abdominal pain of 3 days' duration. He had been taking oral penicillin and ibuprofen for several weeks for a chronically infected tooth, and initially his symptoms were attributed to severe dental infection or antibiotic-associated diarrhea. A chest X-ray at the time of the first emergency room visit was normal. The patient was given intravenous fluids and was referred for dental extraction, but he was hospitalized the next day for progressively severe respiratory and generalized systemic distress. He was transferred to the intensive care unit, where he was placed on mechanical ventilation 11 hours after hospital admission because of increasing respiratory failure. Sputum, blood, and urine cultures and Gram stains were negative or nondiagnostic; these tests included respiratory viral cultures for respiratory syncytial virus; adenovirus; parainfluenza virus types 1, 2, 3; and influenza virus types A and B (Table 1). Chest X-rays over a 3-day period showed pleural effusions and progressive, refractory bilateral pulmonary infiltrates, initially interstitial then alveolar. Intravenous therapies included fluids, vasopressors, and inotropic agents, including dobutamine, dopamine, and norepinephrine. Antimicrobial drugs administered during the hospital stay included azithromycin, ceftriaxone, doxycycline, clindamycin, levofloxacin, and trimethoprim/sulfamethoxazole. The patient's fever persisted throughout the 3-day hospital stay, his pulmonary and cardiac status deteriorated, and he died 5 days after onset of illness, despite intensive critical care support.
Hantavirus infection was confirmed by both immunohistochemical study of lung and kidney tissue, using a cross-reactive monoclonal antibody (GB04-BF07) as described (11). Enzyme-linked immunosorbent assay for serum antibodies showed an immunoglobulin (Ig) G titer of 1:400 and an IgM titer of 1:1600 for Sin Nombre antibody.
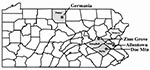
Patient 1 lived in a rural area in Upper Macungie Township near Allentown, Pennsylvania (Figure 1). In the 8 weeks before his death, he hunted deer almost every day and was potentially exposed to mouse habitats in four Pennsylvania counties. The longest exposure was a 3-day, 2-night stay in a small cabin near Germania in Potter County, Pennsylvania (Figure 1), 3 weeks before onset of illness. The patient slept on a mattress in a loft. During an investigation of the cabin in December 1997, rodent droppings were found on the loft floor around the mattress. A dead Peromyscus leucopus (white-footed mouse) was found in a trap on the cabin floor. When unoccupied, the cabin was closed and the downstairs windows were boarded shut. On January 6, 1998, 70 live traps were set in and around the cabin. Eleven small mammals were captured: seven P. leucopus, two P. maniculatus (deer mouse), one Clethrionomys gapperi (red-backed vole), and one Blarina brevicauda (short-tailed shrew). The animals were anesthetized and bled through a capillary tube from the periorbital sinus. Blood was placed on Nobuto blood collection strips, air-dried, and analyzed for hantavirus antibodies. Lungs were collected from the mammals, placed on ice, and kept frozen at -70 C until virus isolation.
Patient 1 had also cleaned a small trailer near Zion Grove in Schuylkill County, Pennsylvania, approximately 6 weeks before onset of illness, and had noted mouse droppings there. He hunted avidly with a crossbow, often from shelter blinds on Doe Mountain in Berks County, Pennsylvania, in the 8 weeks before his illness.
Case 2. A 39-year-old machine shop worker and mother of two children visited a Monroe County, Pennsylvania, hospital emergency department in March 1997. She had fever, myalgias, profound weakness, cough, and shortness of breath, and had not urinated for 3 days. She received intravenous fluids, and laboratory testing was performed (Table 1). Because of acteriuria, an initial diagnosis of urinary tract infection was made. Treatment with clarithromycin and cefuroxime oral antimicrobial drugs was initiated, and the patient was discharged; 48 hours later, she returned to the hospital, complaining of severe backache, dizziness, abdominal pain, and shortness of breath. She was admitted to the hospital and 12 hours later was placed on mechanical ventilation because of respiratory distress. She rapidly became progressively hypotensive despite fluid and inotropic therapy, and she died on hospital day 2, day 5 after onset of illness. Antimicrobial drugs administered included erythromycin, ceftazidime, and cefotaxime. Blood cultures remained sterile, and sputum cultures grew oral flora. A urine culture taken at admission yielded >100,000 Escherichia coli/mL. A year after the patient's death, immunohistochemical testing of archival tissue from an initially nondiagnostic autopsy confirmed the diagnosis of HPS. No serum was available for testing.
Patient 2 lived in a mobile home in a rural, wooded area near Cresco, Barrett Township, Monroe County, Pennsylvania (Figure 1). The machine shop where she worked was also in an isolated area. Family members stated that she had not left the area for several months before her death. During the environmental studies begun 14 months after the patient's death, rodent droppings were found under the kitchen sink of her home. That evening, 70 rodent live traps were placed in and around the mobile home, and six P. leucopus mice were captured. The rodents were bled, and lung tissue was collected. On June 2, 1998, the machine shop was examined, but no evidence of rodent activity was noted. Seventy traps were set in and around the shop. Six P. leucopus, one Tamias striatus (eastern chipmunk), and one B. brevicauda were captured in the surrounding wooded area; no rodents were captured inside the building.
RNA Extraction, Reverse Transcription-Polymerase Chain Reaction (RT-PCR), and Genetic Analysis
RNA was extracted from Patient 1's lung tissue, and RT-PCR and nucleotide sequence analysis were carried out as described (12). RNA was also extracted from six seropositive rodents, and RT-PCR and nucleotide sequence analysis were performed (13).
Genetic Analysis of Virus from Lung Tissue
The nucleotide sequences of the fragments S, M-G1, and M-G2 were compared with the equivalent regions of other previously characterized Peromyscus-associated hantavirus. The hantavirus in Patient 1's lung was clearly identified as Monongahela virus, which was first detected in P. maniculatus nubiterrae (Cloudland deer mice) captured in West Virginia in 1985 (10). Virus S segment fragment showed 95.7% nucleotide identity and 100% deduced amino acid compared with the Monongahela-1 virus strain. A shorter, overlapping fragment (246 nucleotide) was available to compare with the Monongahela-3 virus strain. Again, a 96.7% nucleotide identity and 100% amino acid identity were found. Other Peromyscus-borne hantaviruses were more distantly related. Comparison of this patient's hantavirus sequence with those of New York virus (strains NY-1 and RI-1) and SNV (strains NMH10, CC74 and CC107) identified a nucleotide difference of 14.8% to 19.6% and an amino acid difference of 2.3% to 3.1% (14-16).
Results were similar, with 95.1% nucleotide and 100% amino acid identity, when the virus M-G2 fragment sequence from Patient 1 was compared with the Monongahela-2 virus strain. Nucleotide and amino acid differences of 15.6% to 21.5% and 1.5 to 7.4%, respectively, were seen when this fragment was compared with the more distantly related Peromyscus-borne hantaviruses, including Blue River virus (strains Indiana and Oklahoma), New York virus (strains RI-1, NY-1, and NY-2), and SNV (strains NM H10, CC74, and CC107) (15-19). No comparable piece of Monongahela virus sequence was available to align with the virus M-G1 fragment from this patient. However, sequence identity differences of 15.1% to 18.1% and 3.5% to 7.0% were seen at the nucleotide and amino acid levels, respectively, when the patient M-G1 sequence was compared with those of the other Peromyscus-associated hantaviruses.
Analysis of Rodent Samples
Three of seven P. leucopus mice (PA1, PA, and PA9) trapped at the suspected exposure site for Patient 1 in Potter County and three of six P. leucopus mice (MC-3, MC-4, and MC-6) trapped at the Monroe County location (suspected exposure site for Patient 2) contained hantavirus-specific antibodies. With the exception of MC-3, Monongahela virus-specific RNA was detected in each of the seropositive rodents by RT-PCR. P. leucopus MC-3 was a juvenile mouse, suggesting that maternal antibody may have been detected. The virus nucleotide sequences obtained from the Potter County mice closely matched those obtained in Case 1, with the M-G1 PCR fragment sequences being identical. The virus nucleotide sequence fragments obtained from the Monroe County mice differed from those from Patient 1 and the Potter County rodent samples by approximately 10%.
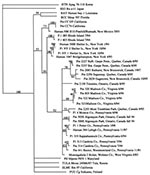
A detailed phylogenetic analysis of the M-G2 fragment sequences of these and other published hantavirus sequences was carried out. A 50% majority rule consensus tree was generated by bootstrap analysis with 1,000 replications (Figure 2). Hantaan and Seoul hantavirus sequences were used as an outgroup. This analysis showed that the viruses detected in Patient 1 (human 584) and rodents (PA1, Potter County; and MC-4, Monroe County) were clustered in the Monongahela virus lineage.
Pathology
Examination of lung tissue from Patient 1 by microscopy showed a mild to moderate interstitial pneumonitis, with mononuclear infiltration and congestion, and edema typical of HPS. Typical immunoblasts were seen within the red pulp in periarteriolar sheaths of the spleen. Immunohistochemical examination showed widespread staining of hantavirus antigens within endothelial and follicular dendritic cells, similar to the pattern seen in previous cases of HPS. However, instead of the typical granular staining, the lung immunostaining was more linear and curvilinear, similar to staining seen in infections with Hantaan virus. The immunostaining in the kidney was also more abundant than with typical HPS cases.
We describe two Pennsylvania cases that add incrementally to knowledge about North American hantavirus. Rodent hosts carrying hantavirus in an asymptomatic but communicable form are found in most areas of North America (21). Clinicians should be vigilant for possible cases of HPS, even in areas where no cases have been previously recognized.
These Pennsylvania cases share with earlier (2) and more recently described cases (22) the clinical picture of a nonspecific and often nonpulmonary flulike prodromal phase. The surprising rapidity with which noncardiogenic pulmonary edema affects often previously healthy victims separates HPS from many other common febrile diseases seen in North America. Some reports have suggested that patients be transferred to facilities with extracorporeal membrane oxygenation capability for patients who do not respond to respiratory and inotropic support (23). However, data may be insufficient to establish the precise role of extracorporeal membrane oxygenation support in the management of these patients (24).
These Pennsylvania cases also reflect the first report of human infection with the Monongahela variant of hantavirus. The initial description of this strain was from archival tissue of P. maniculatus nubiterrae captured in the Monongahela National Forest in West Virginia. Retrospective serologic diagnoses of HPS have been made in nonfatal human cases from Virginia and West Virginia, but none of these cases, to our knowledge, have had RT-PCR confirmation of the Monongahela variant. Recent analysis of hantavirus RNA and rodent mitochondrial DNA has shown genetically distinct clades of virus and rodents useful in understanding suspected cospeciation of virus types within host rodent subspecies (25-27). Monongahela hantavirus appears to be closer in evolutionary distance to the New York/Rhode Island strains than to SNV strains found in the midwestern and southwestern United States (16). Finding Monongahela variant of hantavirus in both P. leucopus and P. maniculatus can be explained by spillover between the two mouse species, given their frequent sympatric and synchronistic existence (16). Allopatric migrations and other biogeographic influences may explain the existence of hantavirus variants in more than one species or subspecies of host rodent (16).
Given the complexity of the genetic relationships between the Peromycine-borne hantaviruses, it is unclear whether the Monongahela virus lineage will be considered a distinct hantavirus species or a subspecies of SNV (18). However, the pathologic findings in Monongahela-associated HPS, together with its primary association with a distinct P. maniculatus subspecies, suggest biologic differences between Monongahela virus and classic SNV lineages.
Clinical differences have been noted for HPS cases due to different hantavirus types in North and South America. Renal disease and myositis are more evident in patients infected with hantavirus carried by Sigmodon sp. and Oryzomys sp. than by Peromyscus sp. (4). The unusual linear immunohistochemical immunostaining and heavy renal staining seen in autopsy material from the two Pennsylvania HPS cases may also have clinical importance. The ultimate goals of effective treatment and prevention of HPS will require continued close cooperation between clinicians, laboratorians, virologists, mammalogists, and epidemiologists.
Dr. Rhodes is chief of the Infectious Diseases Division, Lehigh Valley Hospital, Allentown, Pennsylvania. He is a consultant for clinical Infectious Diseases and maintains special interests in nosocomial infections, hospital epidemiology, and bioterrorism response planning.
Acknowledgment
The authors thank Andre Weltman, Jacquelyn A. Hakim, Samuel Land, and Joni Young for assistance with the investigation; Darla Molnar for photographic assistance; and Pamela Robson and Bernadette Glenn for assistance with manuscript preparation.
References
- Garrett L. The coming plague: newly emerging diseases in a world out of balance. New York: Penguin Books; 1994.
- Duchin JS, Koster FT, Peters CJ, Simpson GL, Tempest B, Zaki SR, Hantavirus pulmonary syndrome: A clinical description of 17 patients with a newly recognized disease. N Engl J Med. 1994;330:949–55. DOIPubMedGoogle Scholar
- White DJ, Means RG, Birkhead GS, Bosler EM, Grady LJ, Chatterjee N, Human and rodent hantavirus infection in New York state. Arch Intern Med. 1996;156:722–6. DOIPubMedGoogle Scholar
- Khan AS, Gaviria M, Rollin PE, Hlady WG, Ksiazek TG, Armstrong LR, Hantavirus pulmonary syndrome in Florida: association with the newly identified Black Creek Canal virus. Am J Med. 1996;100:46–8. DOIPubMedGoogle Scholar
- Morzunov SP, Feldman H, Spiropoulou CF, Semenova VA, Rollin PE, Ksiazek TG, A newly recognized virus associated with a fatal case of hantavirus pulmonary syndrome in Louisiana. J Virol. 1995;69:1980–3.PubMedGoogle Scholar
- Khan AS, Ksiazek TG, Peters CJ. Hantavirus pulmonary syndrome. Lancet. 1996;347:739–41. DOIPubMedGoogle Scholar
- Peters CJ, Khan AS, Zaki SR. Hantavirus in the United States. Arch Intern Med. 1996;156:705–6. DOIPubMedGoogle Scholar
- Mertz GJ, Hjelle BL, Bryan RT. Hantavirus infection. In: Schrier RW, editor. Advances in internal medicine. Chicago: Mosby-Year Book;1997.p. 369-421.
- Doyle TJ, Bryan RT, Peters CJ. Viral hemorrhagic fevers and hantavirus infection in the Americas. Infect Dis Clin North Am. 1998;12:95–110. DOIPubMedGoogle Scholar
- Song JW, Baek LJ, Nagle JW, Schlitter D, Yanagihara R. Genetic and phylogenetic analysis of hantaviral sequences amplified from archival tissues of deer mice (Peromyscus maniculatus nubiterrae) captured in the eastern United States. Arch Virol. 1996;141:959–67. DOIPubMedGoogle Scholar
- Zaki SR, Greer PW, Coffield LM, Goldsmith CS, Nolte KB, Foucar K, Hantavirus pulmonary syndrome: pathogenesis of an emerging infectious disease. Am J Pathol. 1995;146:552–79.PubMedGoogle Scholar
- Johnson AM, Bowen MD, Ksiazek TG, Williams RJ, Bryan RT, Mills JN, Laguna negra virus associated with HPS in western Paraguay and Bolivia. Virology. 1997;238:115–27. DOIPubMedGoogle Scholar
- Campbell WP, Huang C. Detection of California serogroup viruses using universal primers and reverse transcription-polymerase chain reaction. J Virol Methods. 1995;53:55–61. DOIPubMedGoogle Scholar
- Hjelle B, Krolikowski J, Torrez-Martinez N, Chavez-Giles F, Vanner C, Laposata E. Phylogenetically distinct hantavirus implicated in a case of hantavirus pulmonary syndrome in the northeastern United States. J Med Virol. 1995;46:21–7. DOIPubMedGoogle Scholar
- Li D, Schmaljohn AL, Anderson K, Schmaljohn CS. Complete nucleotide sequences of the M and S segments of two hantavirus isolates from California: evidence for re-assortment in nature among viruses related to hantavirus pulmonary syndrome. Virology. 1995;206:973–83. DOIPubMedGoogle Scholar
- Spiropoulou CF, Morzunov S, Feldmann H, Sanchez A, Peters CJ, Nichol ST. Genome structure and variability of a virus causing hantavirus pulmonary syndrome. Virology. 1994;200:715–23. DOIPubMedGoogle Scholar
- Hjelle B, Lee SW, Song W, Torrez-Martinez N, Song JW, Yanagihara R, Molecular linkage of hantavirus pulmonary syndrome to the white-footed mouse, Peromyscus leucopus: genetic characterization of the M genome of New York virus. J Virol. 1995;69:8137–41.PubMedGoogle Scholar
- Morzunov SP, Rowe JE, Ksiazek TG, Peters CJ, St. Jeor SC, Nichol ST, Genetic analysis of the diversity and origin of hantaviruses in Peromyscus leucopus mice in North America. J Virol. 1998;72:57–64.PubMedGoogle Scholar
- Monroe MC, Morzunov SP, Johnson AM, Bowen MD, Artsob H, Yates T, Genetic diversity and distribution of peromyscus-borne hantaviruses in North America and comparison with other hantaviruses. Emerg Infect Dis. 1999;5:75–86. DOIPubMedGoogle Scholar
- Swofford DL. PAUP*: Phylogenetic analysis using parsimony (*and other methods) (computer program). Version 4.0. Sinauer, Sunderland MA. 1998.
- Mills JN, Johnson JM, Ksiazek TG, Ellis BA, Rollin PE, Yates TL, A survey of hantavirus antibody in small-mammal populations in selected United States national parks. Am J Trop Med Hyg. 1998;58:525–32.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Hantavirus pulmonary syndrome--Colorado and New Mexico, 1998. MMWR Morb Mortal Wkly Rep. 1998;47:449–52.PubMedGoogle Scholar
- Crowley MR, Katz RW, Kessler R, Simpson SQ, Levy H, Hallin GW, Successful treatment of adults with severe hantavirus pulmonary syndrome with extracorporeal membrane oxygenation. Crit Care Med. 1998;26:806. DOIPubMedGoogle Scholar
- Serna D, Brenner M, Chen JC. Severe hantavirus pulmonary syndrome: a new indication for extracorporeal life support? Crit Care Med. 1998;26:217–8. DOIPubMedGoogle Scholar
- Schmaljohn C, Hjelle B. Hantaviruses: a global disease problem. Emerg Infect Dis. 1997;3:95–104. DOIPubMedGoogle Scholar
- Hjelle B, Jenison SA, Goade DE, Green WB, Fedderson RM, Scott AA. Hantaviruses: clinical, microbiologic and epidemiologic aspects. Crit Rev Clin Lab Sci. 1995;32:469–508. DOIPubMedGoogle Scholar
Figures
Table
Cite This Article1Three fragments of the virus genome were amplified by nested polymerase chain reaction to allow generation of nucleotide (nt) sequences 393 nt in length for an S segment region encoding the N protein, 259 nt in length for an M segment region (M-G1) encoding the G1 protein, and 205 nt in length for an M segment region (M-G2) encoding the G2 protein (12). Sequence data were analyzed by using the Wisconsin GCG Package (version 9.1) software. Initial multiple sequence alignment was done with the GCG Pileup program, and phylogenetic analysis was performed by using the Phylogenetic Analysis Using Parsimony (and other methods) program (version 4.0) (20).
Table of Contents – Volume 6, Number 6—December 2000
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Luther V. Rhodes, 1210 South Cedar Crest Boulevard, Suite #2700, Allentown, Pennsylvania, 18103; Fax: (610) 402-1676
Top