Volume 9, Number 11—November 2003
Dispatch
Levofloxacin Treatment Failure in Haemophilus influenzae Pneumonia
Cite This Article
Citation for Media
Abstract
We describe the first case of failure of oral levofloxacin treatment of community-acquired pneumonia caused by Haemophilus influenzae. The strain showed cross-resistance to fluoroquinolones and carried four mutations in quinolone resistance–determining regions of DNA gyrase and topoisomerase IV genes.
Fluoroquinolones active against Streptococcus pneumoniae and Haemophilus influenzae, such as levofloxacin, moxifloxacin, and gatifloxacin, are currently recommended for the treatment of patients with community-acquired pneumonia whose infection may be due to strains resistant to ß-lactam and macrolide antimicrobial drugs (1–3). In Spain, both pathogens often display multiple resistance (4,5). The failure of oral levofloxacin treatment in four patients with pneumococcal pneumonia has recently been reported (6).
We describe failure of treatment of H. influenzae pneumonia in a patient who died after several days of systemic treatment with levofloxacin. The H. influenzae strain, isolated from the blood and the respiratory tract of the patient, was resistant to levofloxacin, ciprofloxacin, moxifloxacin, and gatifloxacin and carried several mutations in the quinolone resistance–determining regions (QRDR) of DNA gyrase and topoisomerase IV genes.
On February 15, 2002, a 71-year-old woman was admitted to Hospital de l’Esperit Sant, Santa Coloma de Gramenet, Barcelona, with cough, low-grade fever, and reported fatigue. A radiograph of her chest showed infiltrates covering the right upper, middle, and lower lobes and the left basal lobe. A systemic, antimicrobial treatment of levofloxacin (500 mg/day) and teicoplanin (400 mg/day) was administered for severe community-acquired pneumonia. (The patient was allergic to penicillin.) Three months earlier, she had received erythromycin. In a second visit, she was treated with inhaled salbutamol and glucocorticoids. In a third visit, 2 months before she became ill, she received oral moxifloxacin for 5 days.
During her hospital stay, the patient’s clinical condition worsened. Ventilation and perfusion scintiphotographs with Tc99m, an echocardiograph, and a high resolution thoracic scan were performed in an effort to identify another illness or a possible nondrained focus; results were negative. After 7 hospital days of systemic levofloxacin therapy, her clinical condition had not improved. Two blood cultures taken after an episode of fever with a temperature of 38.5°C showed that H. influenzae was resistant to fluoroquinolones (strain 32602). Treatment was switched to chloramphenicol (1 g/6 h), to which the strain was susceptible. However, on the same day as the treatment change, the patient was intubated during a severe episode of respiratory impairment and transferred to the intensive care unit of Hospital de l’Hospitalet, L’Hospitalet de Llobregat, Barcelona. Fluoroquinolone resistance was confirmed with blood cultures (strain 35102) and a bronchial aspirate (strain 35202). Antimicrobial treatment was then switched to aztreonam (1 g/8 h). The patient died 48 hours later.
Species identification, biotyping, and serotyping followed recommended protocols (7), including capsule genotyping of six capsular types (types a–f). Susceptibility testing was performed by microdilution methods, following the guidelines of the National Committee for Clinical Laboratory Standards (8). ß-lactamase activity was determined by the chromogenic cephalosporin test with nitrocephin as substrate. Quality control strains used were H. influenzae ATCC 49247, H. influenzae ATCC 51907, and Escherichia coli ATCC 25922. Determination of an eventual active efflux mechanism of fluoroquinolone resistance was carried out by MIC determination in the presence and absence of carbonyl cyanide 3-chlorophenyl (CCCP) and reserpine in all three clinical strains (9).
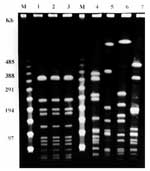
The three test strains and four additional H. influenzae biotype II isolates (obtained previously from blood and respiratory specimens from the geographic area where the patient lived) were examined by pulsed-field gel electrophoresis (PFGE) after digestion of bacterial DNA with SmaI (MBI Fermentas, Vilnius, Lithuania) and separation of the fragments with the CHEF Mapper apparatus (BioRad Laboratories, Hercules, CA) and were visually compared (10).
Amplification and sequence analysis of the QRDRs of gyrA, gyrB, parC, and parE was performed in the three clinical test isolates and in the reference strain H. influenzae ATCC 51907 (11). These specific primers were used: gyrA-f (5´-ccgccgcgtactattctcaat-3´), gyrA-r (5´-gttgccatccccaccgcaatacca-3´), gyrB-f (5´-cctgctctttctgaaactttac-3´), gyrB-r (5´-ccatctaacgcaagggttaatc-3´), parC-f (5´-tctgaacttggcttaattgcc-3´), parC-r (5´-gccacgaccttgctcataaat-3´), parE-f (5´-tcgttagtggccctgcattac-3´), and parE-r (5´-gaacagggcacagagtagggt-3´). Sequencing was done on both DNA strands with the Big Dye Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City, CA), according to the manufacturer’s instructions. The products were resolved and analyzed with an ABI PRISM 377 DNA sequencer (Applied Biosystems). Nucleotide sequences were analyzed by using sequence analysis software (DNAstar, Inc., Madison, WI).
The three clinical isolates were identified as noncapsulated H. influenzae biotype II. The PFGE yielded the same pattern for each, unlike the marked diversity obtained with the four other clinical isolates from the same geographic area (Figure), and the same antimicrobial drug susceptibility testing results (Table). The results of DNA sequencing of the QRDRs of the gyrA, parC, parE, and gyrB genes are shown in the Table. All three strains presented identical amino acid substitutions. They did not produce ß-lactamase and were fully susceptible to several ß-lactam antimicrobial drugs and chloramphenicol. MICs of additional antimicrobial drugs were penicillin, 0.25 µg/mL; ampicillin, 0.12 µg/mL; amoxicillin-clavulanic acid, 0.12/0.06 µg/mL; cefuroxime, 0.12 µg/mL; cefotaxime, 0.03 µg/mL; aztreonam, 0.03 µg/mL; chloramphenicol, 0.12 µg/mL; erythromycin, 16 µg/mL; clarithromycin, 16 µg/mL; and azithromycin, 8.0 µg/mL. Quinolones, other than those depicted in the Table, had the following activity levels: ofloxacin, 32.0 µg/mL; grepafloxacin, 32 µg/mL; clinafloxacin, 2.0 µg/mL. No changes in the MICs of fluoroquinolones were observed either in the presence or absence of CCCP or reserpine, suggesting that active efflux did not contribute to fluoroquinolone resistance in any of the isolates.
This study is the first description of a clinical case of a fluoroquinolone-resistant H. influenzae pneumonia and bacteremia, resulting in therapeutic failure and death after systemic levofloxacin treatment. The causative strain was present in cultures from the patient’s blood and bronchial aspirate. This finding is consistent with the interpretation of the initial chest x-rays, which indicated a clinical case of severe community-acquired pneumonia. Similar cases of community-acquired pneumococcal pneumonia treated with levofloxacin have recently been reported (6).
We have previously described the emergence of fluoroquinolone-resistant H. influenzae in the upper respiratory tract of patients with chronic respiratory infections treated with oral fluoroquinolones (7,12). However, in this patient the only documented exposure to a fluoroquinolone was a 5-day treatment with moxifloxacin 2 months before the pneumonia appeared. This case suggests that exposure to a fluoroquinolone may contraindicate the use of another fluoroquinolone; however, this proposal may not be practical, especially in a patient who is allergic to penicillin.
We have already established that amino acid substitutions in the QRDRs of DNA gyrase and topoisomerase IV accounted for the mechanism of fluoroquinolone resistance in H. influenzae (12).The positions and the amino acid substitutions found in GyrA and ParC in this study were also present in the fluoroquinolone-resistant strains previously studied by our group (12). We have also described the Asp420Asn substitution (ParE) in another strain that exhibited high resistance levels to ciprofloxacin and other fluoroquinolones (13). However, the pattern of QRDR mutations found in the clinical strains described in this study is novel.
Use of the “mutant prevention concentration” has been proposed to discourage the selection of fluoroquinolone-resistant mutants, since resistant mutants are selected exclusively within a concentration range known as the “mutant selection window” (14). This “window” may be more easily attained with therapeutic regimes based on oral and once-daily administered doses, as is the case for moxifloxacin and levofloxacin. An important difference between S. pneumoniae and H. influenzae is that the MIC90’s for levofloxacin are 1.0-2.0 µg/mL and 0.03 µg/mL, respectively (3), a 30-fold difference. This suggests that the events necessary to reach a similar level of resistance in H. influenzae are more complex than they are in S. pneumoniae.
The peak level of levofloxacin, 4 hours after a 500-mg dose, is approximately 4.0 µg/mL (serum) and 11.0 µg/mL (epithelial fluid) (3,15), which is clearly below the MIC of 16.0 µg/mL shown by these resistant strains. Where these clinical H. influenzae strains originated is unknown. Following the patient’s oral moxifloxacin treatment 2 months before hospital admission, the H. influenzae may have become resistant to fluoroquinolones through a typical, step-by-step mutation process in the primary targets, DNA gyrase and topoisomerase IV, by means of four mutations in gyrA, parC, and parE genes. The initial treatment also may have resulted in the mutations in gyrA only, with additional mutations occurring during the subsequent treatment with levofloxacin.
The H. influenzae strain acquired several mutations in the QRDRs and was cross-resistant to all members of a group of assayed fluoroquinolones. We believe that this case may represent an emerging clinical and microbiologic challenge, which may threaten the management of pneumonia and other infections. We recommend that blood cultures taken from patients with community-acquired pneumonia be tested for fluoroquinolone resistance, particularly when the patient has already been given these antimicrobial drugs.
Dr. Bastida is a clinical microbiologist in charge of a diagnostic microbiology laboratory in a 165-bed hospital. Her research interests are focused in clinical microbiology.
Acknowledgments
This study was supported by grants from the Spanish Ministry of Health, Fondo de Investigaciones Sanitarias (J.C., 304-99 and A.G.C., 00/0258). M.P.-V. is a recipient of a fellowship (02/0056) from the Instituto de Salud Carlos III, Ministry of Health, Spain.
The DNA sequences obtained in this study have been submitted to GenBank under accession nos. AJ508043, AJ508044, AJ508045, and AJ508046.
References
- Ball P, Baquero F, Cars O, File T, Garau J, Klugman K, Antibiotic therapy of community respiratory tract infections: strategies for optimal outcomes and minimized resistance emergence. J Antimicrob Chemother. 2002;49:31–40. DOIPubMedGoogle Scholar
- Niederman MS, Mandell LA, Anzueto A, Bass JB, Broughton WA, Campbell GD, Guidelines for the management of adults with community-acquired pneumonia: diagnosis, assessment of severity, antimicrobial therapy, and prevention. Am J Respir Crit Care Med. 2001;163:1730–54.PubMedGoogle Scholar
- Lode H, Allewelt M. Role of newer fluoroquinolones in lower respiratory tract infections. J Antimicrob Chemother. 2002;49:709–12. DOIPubMedGoogle Scholar
- Fenoll A, Jado I, Vicioso D, Pérez A, Casal J. Evolution of Streptococcus pneumoniae serotypes and antibiotic resistance in Spain (1990 to 1996). J Clin Microbiol. 1998;36:3447–54.PubMedGoogle Scholar
- Campos J, Chanyangam M, deGroot R, Smith AL, Tenover FC, Reig R. Genetic relatedness of antibiotic resistance determinants in multiply resistant Haemophilus influenzae. J Infect Dis. 1989;160:810–7. DOIPubMedGoogle Scholar
- Davidson R, Cavalcanti R, Brunton JL, Bast DJ, Azavedo JCS, Kibsey P, Resistance to levofloxacin and failure of treatment of pneumococcal pneumonia. N Engl J Med. 2002;346:747–50. DOIPubMedGoogle Scholar
- Campos J, Sáez-Nieto JA. Gram negative infections: Haemophilus and other clinically relevant Gram negative coccobacilli. In: Cimolai N, editor. Laboratory diagnosis of bacterial infections. New York: Marcel Dekker, Inc; 2001. p. 557–80.
- Performance standards for antimicrobial susceptibility testing: eleventh informational supplement. Wayne (PA): National Committee for Clinical Laboratory Standards (NCCLS document no. M100–S11); 2001.
- Sánchez L, Pan W, Viñas M, Nikaido H. The acrAB homolog of H. influenzae codes for a functional multidrug efflux pump. J Bacteriol. 1997;179:6855–7.PubMedGoogle Scholar
- Campos J, Román F, Georgiou M, García C, Gómez-Lus R, Cantón R, Long-term persistence of ciprofloxacin-resistant Haemophilus influenzae in patients with cystic fibrosis. J Infect Dis. 1996;174:1345–7. DOIPubMedGoogle Scholar
- Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269:496–512. DOIPubMedGoogle Scholar
- Georgiou M, Muñoz R, Román F, Cantón R, Gómez-Lus R, Campos J, Ciprofloxacin-resistant Haemophilus influenzae strains possess mutations in analogous positions of GyrA and ParC. Antimicrob Agents Chemother. 1996;40:1741–4.PubMedGoogle Scholar
- Pérez-Vázquez M, Camacho T, Román F, Campos J, Cantón R. Mutations in quinolone-resistant determining regions (QRDR) of gyrA, gyrB, parC and parE genes in Haemophilus influenzae clinical isolates. In: Abstracts of the 41st Interscience Conference on Antimicrobial Agents and Chemotherapy, Chicago, Illinois; December 16–19, 2001; Abstract 145. Washington: American Society for Microbiology; 2001.
- Zhao X, Drlica K. Restricting the selection of antibiotic-resistant mutants: a general strategy derived from fluoroquinolone studies. Clin Infect Dis. 2001;33(Suppl 3):S147–56. DOIPubMedGoogle Scholar
- Chien SC, Rogge MC, Gisclon LG, Curtin C, Wong F, Natarajan J, Pharmacokinetic profile of levofloxacin following once-daily 500-milligram oral or intravenous doses. Antimicrob Agents Chemother. 1997;41:2256–60.PubMedGoogle Scholar
Figure
Table
Cite This ArticleTable of Contents – Volume 9, Number 11—November 2003
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Carles Alonso-Tarrés, Servei d’Anàlisis Clíniques-Microbiologia, Hospital de l’Hospitalet, Consorci Sanitari Integral, Avda Josep Molins 29-41, L’Hospitalet de Llobregat, 08906 Barcelona, Spain; fax: +34.93.333.45.43
Top