Volume 11, Number 2—February 2005
Research
Carbapenemase-producing Enterobacteriaceae, U.S. Rivers
Cite This Article
Citation for Media
Abstract
Our study was initiated by previous isolation of 30 imipenem-resistant, gram-negative rods from 7 of 16 U.S. rivers sampled from 1999 to 2001. Imipenem hydrolysis was detected in 22 of those isolates identified as Enterobacter asburiae. Random amplified polymorphism DNA analysis showed that these E. asburiae isolates were genetically indistinguishable. An identical clavulanic acid–inhibited β-lactamase IMI-2 was identified from each isolate that shared 99% and 97% amino acid identity with the chromosome-encoded β-lactamases IMI-1 and NmcA, respectively, from E. cloacae clinical isolates. The blaIMI-2 gene was located on a self-transferable 66-kb plasmid. Sequence analysis of a cloned 5.5-kb DNA fragment obtained from 1 of the imipenem-resistant E. asburiae isolates identified an upstream LysR-type regulator gene that explained inducibility of IMI-2 expression. β-Lactamase IMI-2 is the first inducible and plasmid-encoded carbapenemase. Identification of clonally related E. asburiae isolates from distant rivers indicates an environmental and enterobacterial reservoir for carbapenemase genes.
Carbapenems, such as imipenem and meropenem, are the most potent β-lactam antimicrobial drugs for avoiding resistance in gram-negative rods. Resistance to carbapenems is rare in Enterobacteriaceae and may be mediated by 3 mechanisms: hyperproduction of an AmpC-type cephalosporinase combined with decreased drug permeability through the outer membrane, decreased affinity of penicillin-binding proteins that constitute target proteins for carbapenems, and carbapenem-hydrolyzing β-lactamases (1–3). These rare carbapenemases may be either plasmid-mediated metallo-β-lactamases (IMP- and VIM-type) or chromosomally encoded and clavulanate-inhibited enzymes (NmcA, IMI-1, Sme-1/Sme-2) (2,4–9). The latter group of enzymes shares consistent percentage of identity and belongs to the Ambler class A of β-lactamases (2,10). Very recently, plasmid-mediated and clavulanate-inhibited carbapenemases have been reported as a source of nosocomial infections in U.S. hospitals (11–15).
While the role of animals in the emergence of clinically important, antimicrobial-resistant strains has been extensively shown (e.g., in Salmonella spp.), the role of aquatic environment as a reservoir of antimicrobial-resistance genes is less established (16–21). A recent study described high levels of antimicrobial-resistant strains from U.S. rivers (22). We identified the imipenem-resistant, gram-negative strains recovered from that study and analyzed the molecular mechanism involved in carbapenem resistance of the imipenem-resistant enterobacterial strains. Clonally related Enterobacter asburiae strains were identified in midwestern U.S. rivers. E. asburiae naturally produces a cephalosporinase but no carbapenemase and may be responsible for nosocomial infections (23). Here, the strains expressed a novel plasmid-encoded and clavulanate-inhibited carbapenemase.
Bacterial Isolates
A previous study identified 30 imipenem-resistant, gram-negative strains out of 1,861 ampicillin-resistant, gram-negative isolates from 7 out of 16 U.S. rivers that were sampled from 1999 to 2001 (22). Identification of these imipenem-resistant isolates was performed by conventional biochemical techniques (API-20E and API-NE systems [bioMérieux, Marcy-l’Etoile, France]), and confirmed by 16S rDNA sequencing (24).
E. asburiae CIP 103358 and E. asburiae CIP 105006 were used as reference strains (Institut Pasteur strain collection, Paris, France). E. cloacae NOR-1 and E. cloacae 1413B were used as strains that produce the chromosome-encoded, clavulanate-inhibited carbapenemases NmcA and IMI-1, respectively (5,8). One of the E. asburiae isolates recovered from a river (strain MS7) was used for cloning experiments. Streptomycin-resistant Escherichia coli DH10B strain was used in cloning and conjugation experiments (Life Technologies, Eragny, France).
Antimicrobial Agents and Resistance Study
The antimicrobial agents and their sources were as follows: amoxicillin, ceftazidime, clavulanic acid, and ticarcillin (GlaxoSmithKline, Nanterre, France); aztreonam (Bristol-Myers Squibb, Paris La Defense, France); cephalothin (Eli Lilly, Saint-Cloud, France); piperacillin and tazobactam (Lederle, Les Oullins, France); cefotaxime (Aventis, Romainville, France); imipenem (without cilastatin) (Merck Sharp and Dohme, Paris, France); meropenem (AstraZeneca, Paris, France); ampicillin and streptomycin (Sigma, Paris, France).
MICs were determined by an agar dilution technique on Mueller-Hinton (MH) agar (Sanofi Diagnostics Pasteur, Marnes-La-Coquette, France) with an inoculum of 104 CFU per spot (25). Carbapenemase activity was determined by UV spectrophotometry with culture extracts of each of the imipenem-resistant, gram-negative rods and imipenem (100 μmol) as substrate, as reported previously (26). One unit of enzyme activity corresponded to the hydrolysis of 1 μmol of substrate per min. Inducibility of the β-lactamase expression was determined with imipenem and cefoxitin as β-lactamase inducers, as described (27). Briefly overnight culture of each imipenem-resistant Enterobacter asburiae isolate was diluted (1:10) in a prewarmed trypticase soy broth, allowed to culture in an antimicrobial-free medium for 2 h, and further cultured for 6 h with cefoxitin (2–50 mg/L) or imipenem (10–50 mg/L). β-Lactamase culture extracts were obtained after centrifugation and sonication, as detailed (26).
Nucleic Acid Techniques and Conjugation
Genotype comparison of the imipenem-resistant E. asburiae strains was performed by using the random amplified polymorphism detection (RAPD) technique as described with primer 6MW (CCGACTCGAGNNNNNNATGTGG) and primers UBC 245 and UBC 282 (26,28,29). Transfer of the imipenem resistance marker from each imipenem-resistant E. asburiae isolate to Escherichia coli DH10B was attempted by using the immobilization filter mating out technique, as described (26). Briefly, equal volume (0.1 mL) of overnight cultures of each Enterobacter asburiae isolate and Escherichia coli DH10B were put onto a paper filter that was placed on an MH agar plate. Twenty-four hours later, the filter was removed, washed with water (0.2 mL), and the bacterial suspension was spread onto MH agar plates containing ampicillin (100 mg/L) and streptomycin (50 mg/L) for selecting transconjugants after 24 h (26).
Plasmid extraction was performed for each Enterobacter asburiae strain and their transconjugants and compared to reference plasmid sizes of Escherichia coli NCTC 50192 by using the Kieser technique designed to extract large size plasmids (30,31). Whole-cell DNA of Enterobacter spp. reference strains and of an Enterobacter asburiae strain MS7 was extracted as described (26).
Southern hybridization of plasmid DNA (26) of the transconjugants was performed as described by the manufacturer with the ECL nonradioactive kit (Amersham, Les Ulis, France). An 818–bp internal probe for blaIMI-1 was obtained by using primers IMI-A (5′-ATAGCCATCCTTGTTTAGCTC-3′) and IMI-B (5′-TCTGCGATTACTTTATCCTC-3′) and standard polymerase chain reaction (PCR) amplification procedures (5,26).
Primers designed to hybridize to the ends of the blaNmcA, blaIMI-1, and blaSme-1/Sme-2 genes were used for standard PCR amplification experiments (5,7,8) with plasmid DNA of each imipenem-resistant E. asburiae isolate and of their transconjugants as templates. Cloning experiments were then performed with BamHI restricted whole-cell DNA of E. asburiae MS7 followed by ligation of DNA fragments into the BamHI-site of cloning vector pGB2 (32). Recombinant plasmids were transformed by electroporation into Escherichia coli DH10B electrocompetent cells (26). E. coli DH10B harboring recombinant plasmids was selected on MH agar plates containing ampicillin (100 mg/L) and streptomycin (100 mg/L).
DNA sequencing of both strands of PCR fragments amplified with the primers for blaIMI-1 and plasmid DNA of Enterobacter asburiae isolates as templates and of the cloned fragment of a recombinant plasmid was determined with an Applied Biosystems sequencer (ABI377). The nucleotide sequences and the deduced protein sequences were analyzed with software available on the Internet from the National Center for Biotechnology Information Web site (http://www.ncbi.nlm.nih.gov/BLAST).
Bacterial Identification
Twenty-nine of the 30 imipenem-resistant isolates substantially hydrolyzed imipenem, i.e., 10.5 ± 1.6 U/mg of protein of culture extracts. These isolates were a single Aeromonas hydrophila isolate, 6 Stenotrophomonas maltophilia isolates known to naturally produce carbapenemases, and 22 Enterobacter spp. isolates identified as E. asburiae that were further analyzed.
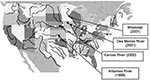
As reported in Table 1, E. asburiae strains were isolated at different times from several rivers in the midwest. Other tested rivers had ampicillin-resistant isolates that were not imipenem-resistant (Figure). These rivers were Arkansas (Little Rock), Canadian (Oklahoma City), Hudson (New York), Chicago (Chicago), Colorado (Glenwood Springs), Missouri (Parkville), Cuyahoga (Cleveland), Mississippi (New Orleans, St. Louis), Ohio (Cincinnati, Louisville, Pittsburgh, Wheeling), Platte (Grand Island), Scioto (Columbus), Wabash (Terre Haute), and White (Indianapolis). RAPD analysis was then performed to compare all imipenem-resistant E. asburiae isolates. Using a series of different primers, this genotyping technique identified clonally indistinguishable E. asburiae isolates, although they were from various geographic origins (data not shown).
β-Lactam Resistance Marker
The imipenem resistance marker was transferred from each imipenem-resistant E. asburiae isolate to Escherichia coli DH10B by conjugation. Plasmid analysis identified a 66-kb plasmid (pNat) from cultures of each imipenem-resistant Enterobacter asburiae isolate, whereas this plasmid was not isolated from E. cloacae and E. asburiae reference strains (data not shown). PCR experiments with primers for the blaIMI-1 gene were positive with plasmid DNA of each E. asburiae isolate and transconjugants as templates, whereas primers designed to amplify blaNmcA and blaSme-1/Sme-2 failed to give PCR product. The Southern blot analysis confirmed that the blaIMI-like gene was located on the natural plasmid pNat (data not shown).
Sequencing PCR products with primers hybridizing at the ends of the blaIMI-1 gene and plasmid DNA of each imipenem-resistant E. asburiae isolate identified the same β-lactamase IMI-2 in all cases. This novel enzyme had 2 amino acid substitutions (tyrosine to histidine at position Ambler 105 and asparagine to aspartic acid at position Ambler 35) compared to the chromosomally encoded carbapenemase IMI-1 (5). β-Lactamase IMI-1 had been isolated from an E. cloacae isolate from Minnesota close to locations where IMI-2–producing isolates have been found (5). However, the blaIMI-2 gene was not just a point-mutant derivative of the blaIMI-1 gene, since these genes differ by 11 nucleotide substitutions. β-Lactamase IMI-2 was also related to NmcA (97% amino acid identity) (8).
Cloning BamHI-restricted DNA of whole-cell DNA of E. asburiae MS7 gave recombinant plasmid pIMI-2 that had a 5.5-kb insert that allowed identification of the surrounding sequence of the blaIMI-2 gene. A gene encoding a LysR-type regulator named IMIR-2 was found just upstream of blaIMI-2. It shared 95% amino acid identity with IMIR-1, which is located upstream of the blaIMI-1 gene (5). The surrounding sequences of blaIMI-2 shared significant nucleotide identity with transposable elements. Part of an open reading frame that shared 97% nucleotide identity with that of the transposase gene tnpA of the transposon Tn2501 (Tn3 family) was identified downstream of blaIMI-2 (33). Upstream of imiR-2, a 142-bp sequence shared 76% nucleotide identity with part of the insertion sequence IS2.
Susceptibility Testing and Expression of Resistance
MICs of several β-lactams, including carbapenems for the IMI-2–positive E. asburiae MS7 and for Escherichia coli DH10B expressing the blaIMI-2 gene were high (Table 2). The MICs of β-lactams for all imipenem-resistant clinical isolates were identical (data not shown). Much higher level of resistance to aztreonam than to expanded-spectrum cephalosporins was found for the IMI-2–positive strains, as reported for the other producers of class A carbapenemases (2). The activity of β-lactamase IMI-2 was partially inhibited by clavulanate and tazobactam. Induction studies showed increase of β-lactamase expression from 17- to 30-fold (170 to 300 U/mg of protein) (for each Enterobacter asburiae isolate when imipenem (50 mg/L) and cefoxitin (50 mg/L) were used as inducers, respectively. These induction results were consistent with location and functionality of a LysR-type regulator gene upstream of the blaIMI-2 gene in the imipenem-resistant E. asburiae isolates. No other antimicrobial resistance marker was carried by natural plasmid pNat.
This report indicates that several U.S. rivers may be a reservoir for broad-spectrum carbapenemases. Here, we report a novel clavulanic-acid inhibited Ambler class A β-lactamase IMI-2 that has an usual spectrum of hydrolysis for this type of β-lactamase, including penicillins, carbapenems, and aztreonam (2). β-Lactamase IMI-2 is closely related to several Ambler class A carbapenemases whose genes are chromosomally located, including blaIMI-1 and blaNmcA, and found in several clinical isolates (5,8). While this work was in progress, a clinical case of an NmcA-producing E. cloacae isolate was reported from Seattle (34). An extended epidemiologic survey identified Sme-1 type–producing Serratia marcescens isolates from the West Coast to the East Coast, which indicates that these isolates may also represent a reservoir for carbapenemases in Enterobacteriaceae (9). Thus, identification of carbapenemase genes in enterobacterial strains from rivers may have clinical importance.
In the present study, the β-lactamase gene was plasmid-encoded and was adjacent to mobile sequences that may play an additional role in gene transfer. The E. asburiae isolates were clonally related and may correspond to a single clone, although they were obtained from distantly related midwestern rivers. The reason for the presence of these antimicrobial-resistant strains in this region is unknown. Taking into account the small number of specimens withdrawn from the rivers and the selection technique for imipenem-resistant isolates (ampicillin- and not imipenem-containing plates), the prevalence of carbapenemase-producing enterobacterial strains may be high in the environment, at least in the Midwest.
Cloning experiments led to identification of a regulatory gene from an E. asburiae strain (found in the other E. asburiae strains as well, [data not shown]) that explained inducibility of carbapenemase expression. Whatever the level of imipenem resistance is, failure of an imipenem-containing regimen may occur when treating infections caused by similar carbapenemase-producing strains, as deduced from results obtained with an animal model of pneumonia (35). Finally, this study raises the question of the importance of this reservoir in Enterobacteriaceae as well as the origin of this plasmid-located carbapenemase gene that may be transferred among other enterobacterial pathogens.
Dr. Aubron is studying antimicrobial resistance mechanisms at the Hospital Bicêtre, South-Paris Medical School, University Paris XI, France. She is a resident specializing in infectious diseases.
Acknowledgments
We thank K. Bush for providing the E. cloacae NOR-1 and E. cloacae 1413B that produced the chromosome-encoded, clavulanate-inhibited carbapenemases NmcA and IMI-1.
This work was funded by a grant from the Ministère de l’Education Nationale et de la Recherche, (UPRES EA 3539) Université Paris XI, Paris, France, and by the European Community (6th PCRD, LSHM-CT-2003-503-335).
References
- Lee EH, Nicolas MH, Kitzis MD, Pialoux G, Collatz E, Gutmann L. Association of two resistance mechanisms in a clinical isolate of Enterobacter cloacae with high level resistance to imipenem. Antimicrob Agents Chemother. 1991;35:1093–8.PubMedGoogle Scholar
- Nordmann P, Poirel L. Emerging carbapenemases in gram-negative aerobes. Clin Microbiol Infect. 2002;8:321–31. DOIPubMedGoogle Scholar
- De Champs C, Henquell C, Guelon D, Sirot D, Gazuy N, Sirot J. Clinical and bacteriological study of nosocomial infections due to Enterobacter aerogenes resistant to imipenem. J Clin Microbiol. 1993;31:123–7.PubMedGoogle Scholar
- Nordmann P, Mariotte S, Naas T, Labia R, Nicolas MH. Biochemical properties of a carbapenem-hydrolyzing β-lactamase from Enterobacter cloacae and cloning of the gene into Escherichia coli. Antimicrob Agents Chemother. 1993;37:939–46.PubMedGoogle Scholar
- Rasmussen BA, Bush K, Keeney D, Yang Y, Hare R, O’Gara C, Characterization of IMI-1 β-lactamase, a class A carbapenem-hydrolyzing enzyme from Enterobacter cloacae. Antimicrob Agents Chemother. 1996;40:2080–6.PubMedGoogle Scholar
- Naas T, Livermore DM, Nordmann P. Characterization of an LysR family protein, SmeR from Serratia marcescens S6, its effect on expression of the carbapenem-hydrolyzing β-lactamase Sme-1, and comparison of this regulator with other β-lactamase regulators. Antimicrob Agents Chemother. 1995;39:629–37.PubMedGoogle Scholar
- Naas T, Vandel L, Sougakoff W, Livermore DM, Nordmann P. Cloning and sequence analysis of the gene for a carbapenem-hydrolyzing class A β-lactamase, Sme-1, from Serratia marcescens S6. Antimicrob Agents Chemother. 1994;38:1262–70.PubMedGoogle Scholar
- Naas T, Nordmann P. Analysis of a carbapenem-hydrolyzing class A β-lactamase from Enterobacter cloacae and of its LysR-type regulatory protein. Proc Natl Acad Sci U S A. 1994;91:7693–7. DOIPubMedGoogle Scholar
- Queenan AM, Torres-Vierra C, Gold HS, Carmeli Y, Eliopoulos GM, Moellering RC Jr, SME-type carbapenem-hydrolyzing class A β-lactamases from geographically diverse Serratia marcescens strains. Antimicrob Agents Chemother. 2000;44:3035–9. DOIPubMedGoogle Scholar
- Ambler RP, Coulson AF, Frère JM, Ghuyssen JM, Joris B, Forsman M, A standard numbering scheme for the class A beta-lactamase. Biochem J. 1991;276:269–72.PubMedGoogle Scholar
- Miriagou V, Tzouvelekis LS, Rossiter S, Tzelepi E, Angulo FJ, Whichard JM. Imipenem resistance in Salmonella clinical strain due to plasmid-mediated class A carbapenemase KPC-2. Antimicrob Agents Chemother. 2003;47:1297–300. DOIPubMedGoogle Scholar
- Yigit H, Queenan AM, Anderson GJ, Domenech-Sanchez A, Biddle JW, Steward CD, Novel carbapenem-hydrolyzing β-lactamase, KPC-1, from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob Agents Chemother. 2001;45:1151–61. DOIPubMedGoogle Scholar
- Smith-Moland E, Black JA, Ourada J, Reisbig MD, Hanson ND, Thomson KS. Occurrence of newer β-lactamases in Klebsiella pneumoniae isolates from 24 U.S. hospitals. Antimicrob Agents Chemother. 2002;46:3837–42. DOIPubMedGoogle Scholar
- Smith-Moland E, Hanson ND, Herrera VL, Black AJ, Lockhart T, Hossain A, Plasmid-mediated, carbapenem-hydrolysing β-lactamase, KPC-2, in Klebsiella pneumoniae isolates. J Antimicrob Chemother. 2003;51:711–4. DOIPubMedGoogle Scholar
- Poirel L, Weldhagen GF, Naas T, De Champs C, Dove MG, Nordmann P. GES-2, class A β-lactamase from Pseudomonas aeruginosa with increased hydrolysis of imipenem. Antimicrob Agents Chemother. 2001;45:2598–603. DOIPubMedGoogle Scholar
- Sherwood L. Antimicrobial use in animal feed—time to stop. N Engl J Med. 2001;345:1202–3. DOIPubMedGoogle Scholar
- White DG, Shaohua Z, Sudler R, Sherry A, Friedman S, Chen S, The isolation of antibiotic-resistant salmonella from retail ground meats. N Engl J Med. 2001;345:1147–54. DOIPubMedGoogle Scholar
- Mac Arthur JV, Tuckfield RC. Spatial patterns in antibiotic resistance among stream bacteria; effect of industrial pollution. Appl Environ Microbiol. 2000;66:3722–6. DOIPubMedGoogle Scholar
- Munesia M, Garcia A, Miro E, Prats G, Jofre J, Navarro F. Bacteriophages and diffusion of β-lactamase genes. Emerg Infect Dis. 2004;10:1134–7.PubMedGoogle Scholar
- Goni-Urriza M, Capdepuy M, Arpin C, Raymond N, Caumette P, Quentin C. Impact of an urban effluent on antibiotic resistance of riverine Enterobacteriaceae and Aeromonas spp. Appl Environ Microbiol. 2000;66:125–32. DOIPubMedGoogle Scholar
- Baya AM, Brayton PR, Brown VL, Grimes DJ, Russek-Cohen E, Colwell RR. Coincident plasmids and antimicrobial resistance in marine bacteria isolated from polluted and unpolluted Atlantic Ocean samples. Appl Environ Microbiol. 1986;51:1285–92.PubMedGoogle Scholar
- Ash RJ, Mauck B, Morgan M. Antibiotic resistance of gram-negative bacteria in rivers, United States. Emerg Infect Dis. 2002;8:713–6.PubMedGoogle Scholar
- Brenner DJ, McWorther AC, Kai A, Steigerwalt AG, Farmer JJ. Enterobacter asburiae sp. nov. a new species found in clinical specimens and reassignement of Erwinia dissolvens and Erwinia nimipressuralis to the genus Enterobacter as Enterobacter dissolvens comb. nov. and Enterobacter nimipressuralis comb. nov. J Clin Microbiol. 1986;23:1114–20.PubMedGoogle Scholar
- Avidor B, Kletter Y, Abulafia S, Golan Y, Ephros M, Giladi M. Molecular diagnosis of cat scratch disease: a two-step approach. J Clin Microbiol. 1997;35:1924–30.PubMedGoogle Scholar
- National Committee for Clinical Laboratory Standards. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically. 6th ed. Approved standard M7-A5. Wayne (PA): The Committee; 2003.
- Poirel L, Guibert M, Bellais S, Naas T, Nordmann P. Integron- and carbenicillinase-mediated reduced susceptibility to amoxicillin-clavulanic acid in isolates of multidrug-resistant Salmonella enterica serotype Typhimurium DT104 from French patients. Antimicrob Agents Chemother. 1999;43:1098–104.PubMedGoogle Scholar
- Poirel L, Naas T, Guibert M, Chaibi EB, Labia R, Nordmann P. Molecular and biochemical characterization of VEB-1, a novel class A extended-spectrum β-lactamase encoded by an Escherichia coli integron gene. Antimicrob Agents Chemother. 1999;43:573–81.PubMedGoogle Scholar
- Nazarowec-White M, Farber JM. Phenotypic and genotypic typing food and clinical isolates of Enterobacter sakazakii. J Med Microbiol. 1999;48:559–67. DOIPubMedGoogle Scholar
- Telenius H, Carter NP, Bebb CE, Nordenskjold M, Ponder BA, Tunnacliffe A. Degenerate oligonucleotide-primer PCR: general amplification of target DNA by single degenerate primer. Genomics. 1992;13:718–25. DOIPubMedGoogle Scholar
- Kieser T. Factors affecting the isolation of CCC DNA from Streptomyces lividans and Escherichia coli. Plasmid. 1984;12:19–36. DOIPubMedGoogle Scholar
- Danel F, Hall LM, Gur D, Livermore DM. OXA-14, another extended-spectrum variant of OXA-10 (PSE-2) β-lactamase from Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1995;39:1881–4.PubMedGoogle Scholar
- Churchward G, Belin D, Nagamine Y. A pSC101-derived plasmid which shows no sequence homology to other commonly used cloning vectors. Gene. 1984;31:165–71. DOIPubMedGoogle Scholar
- Michiels T, Cornelis G, Ellis K, Grinsted J. Tn2501, a component of the lactose transposon Tn951, is an example of a new category of class II transposable elements. J Bacteriol. 1987;169:624–31.PubMedGoogle Scholar
- Pottumarthy S, Smith-Moland E, Juretschko S, Swanzy SR, Thomson KS, Fritsche TR. NmcA carbapenem-hydrolyzing enzyme in Enterobacter cloacae in North America. Emerg Infect Dis. 2003;9:999–1002.PubMedGoogle Scholar
- Mimoz O, Léotard S, Jacolot A, Padoin C, Louchahi K, Petitjean O, Efficacies of imipenem, meropenem, cefepime, and ceftazidime in rats with experimental pneumonia due to a carbapenem-hydrolyzing β-lactamase-producing strain of Enterobacter cloacae. Antimicrob Agents Chemother. 2000;44:885–90. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 11, Number 2—February 2005
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Nordmann P. Service de Bactériologie-Virologie, Hôpital de Bicêtre, 78, rue du Général Leclerc, 94275 Le Kremlin-Bicêtre, France; fax: 33-1-45-21-63-40
Top