Volume 13, Number 3—March 2007
Dispatch
Clostridium difficile in Retail Ground Meat, Canada
Cite This Article
Citation for Media
Abstract
Clostridium difficile was isolated from 12 (20%) of 60 retail ground meat samples purchased over a 10-month period in 2005 in Canada. Eleven isolates were toxigenic, and 8 (67%) were classified as toxinotype III. The human health implications of this finding are unclear, but with the virulence of toxinotype III strains further studies are required.
Clostridium difficile is an important spore-forming human pathogen associated with serious enteric diseases worldwide (1–3). Recently, the epidemiology of C. difficile–associated diarrhea (CDAD), appears to have changed; increased illness and relapse rates have been reported (1,3). Much of this change has been attributed to the emergence of 1 toxigenic strain, classified according to PCR as ribotype 027/toxinotype III and pulsed-field gel electrophoresis (PFGE) as NAP1 (2).
Toxigenic strains of C. difficile typically produce 2 major toxins, A and B, although a small percentage produce only toxin B (3). Certain strains may also produce a binary toxin (known as CDT), whose clinical relevance is under investigation. PCR ribotype 027 strains produce all 3 toxins and have a mutated toxin regulatory gene, tcdC, which is thought to be associated with increased toxin production in vitro (2).
C. difficile is also associated with enteric diseases in animals, including horses, dogs, and pigs (4,5). Recent reports indicating that human and animal isolates are often indistinguishable (4,6) and that PCR ribotype 027 has been isolated from a dog (7) have created concerns regarding potential public health implications. C. difficile, including PCR ribotype 027 (4), has also been isolated from dairy calves, beef calves, veal calves, and adult cattle in Ontario (A. Rodriguez-Palacios et al., unpub. data).
The presence of C. difficile spores in bovine feces indicates the potential for contamination of retail meat products. Although contamination does not necessarily mean foodborne transmission, the possibility of C. difficile being a foodborne pathogen should be investigated. We therefore evaluated the prevalence of C. difficile contamination of retail ground meat samples and characterized the isolates.
A convenience sampling scheme was used whereby meat samples (beef, n = 53 and veal, n = 7) were purchased from 5 grocery retailers in Ontario (4 stores, 57 samples) and Quebec (1 store, 3 samples), Canada. The number of meat packages purchased per month was 12, 2, 4, 4, 2, 2, 1, 11, 21, and 1, from January to October 2005, respectively.
C. difficile were isolated by using C. difficile culture agar supplemented with C. difficile moxalactam norfloxacin (CDMN) and 5% horse blood (CM0601, SR0173E, and SR0048C, Oxoid, Basingstoke, United Kingdom) (8). C. difficile broth was prepared by mixing the ingredients of CM0601, except for the agar, with 0.1% sodium taurocholate (Sigma-Aldrich, Inc., St. Louis, MO, USA). Briefly, 4–5 g of each sample was added to 20 mL of prereduced CDMN broth and incubated anaerobically at 37°C for 10–15 days. Alcohol shock for spore selection was performed by mixing 2 mL homogenized culture broth and 96% ethanol (1:1 [v/v]) for 50 min. After centrifugation (3,800 × g for 10 min), the sediment was streaked onto C. difficile agar. Up to 2 suspected colonies (swarming, rough, nonhemolytic) were subcultured from each plate. C. difficile was presumptively identified on the basis of Gram stain and detection of L-proline aminopeptidase activity (Pro Disc, Remel, Lenexa, KS, USA) and confirmed by identification of the triose phosphate isomerase gene (9).
PCR ribotyping and gene identification for toxins A (tcdA) and B (tcdB), the binding component of CDT (cdtB), and the tcdC gene were performed as previously described (4,10). Toxinotyping of selected isolates was also performed (11). Antimicrobial drug susceptibility to metronidazole, clindamycin, levofloxacin, and vancomycin was determined for all isolates by using the E-test method (AB Biodisk, Solna, Sweden) on Mueller-Hinton agar (12).
C. difficile was isolated from 12 (20%) of 60 meat samples; 11 (20.8%) of 53 ground beef samples, and 1 (14.3%) of 7 ground veal samples (Table 1). Duplicate analysis was performed on 4 samples, and isolation of C. difficile was repeatable.
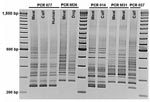
PCR ribotyping showed distinct patterns (Table 1, Figure). The most common ribotype, which accounted for 8 (67%) of 12 isolates, was different from any ribotype previously identified in our laboratory among ≈1,500 human and animal isolates. This ribotype, designated M31, had genes for toxins A, B and CDT; an 18-bp deletion in the tcdC gene and was toxinotype III. These are all molecular characteristics of PCR ribotype 027; however, ribotype pattern M31 was different from ribotype pattern 027 (Figure). PFGE with SmaI indicated that although this strain was distinguishable from prototypic strains NAP1, it had ≈80% similarity and was classified as NAP1 (B. Limbago, pers. comm.).
Two of the remaining 3 ribotypes had classic tcdC PCR fragments and did not have the cdtB gene. One group (n = 2), classified as PCR ribotype 077/toxinotype 0, had been isolated from calves, dogs, and humans (4). Another isolate from Quebec, classified as PCR ribotype 014/toxinotype 0, had also been isolated from calves and humans (3,4). The fourth isolate, nontoxigenic ribotype M26, had been isolated from dogs (6) but could not be toxinotyped because there was no detectable pathogenicity locus (M. Rupnik, pers. comm.). Overall, 3 (25%) of 12 meat C. difficile isolates were indistinguishable from Ontario human isolates.
All meat isolates were susceptible to metronidazole and vancomycin and resistant to levofloxacin and clindamycin (Table 2). These results are in agreement with previous findings for bovine-derived strains (4)
This is the first study to identify C. difficile spores in retail ground meat intended for human consumption. Previously, a study investigating the role of psychrotrophic clostridia on “blown pack” spoilage of commercial packages of chilled vacuum-packed meats and dog rolls reported 2 incidental isolates of C. difficile (13). More recently, a C. difficile isolate was identified in a commercial turkey-based raw diet intended for dogs (14).
The proportion of meat samples contaminated with C. difficile in our study (20%, 12/60) seems higher than those in the aforementioned reports. Possible reasons include the use of a more selective culture protocol in this study (8) and a potential temporary cluster of isolates with PCR ribotype M31 (Table 1). Those meat samples may have originated from the same larger contaminated batch or were subsequently contaminated at the store level during repackaging of retail products. PCR ribotype M31 was not identified in other samples or stores, which may suggest contamination at the retail level. Because PCR ribotype M31/toxinotype III had not been isolated in our laboratory, contamination during processing is unlikely.
The identification of PCR ribotypes 077 and 014, which are recognized human pathogens (3,15), is of concern, although the actual risk for disease is unclear. Of additional concern is isolation of toxinotype III strains that have many similarities with PCR ribotype 027, an important cause of CDAD in humans (2). This similarity was highlighted by classification of this strain by PFGE as NAP1.
The presence of meat-derived PCR ribotypes indistinguishable from human, bovine, and canine ribotypes further supports the potential risk for cross-transmission among species and suggests that ingestion of viable spores might occur. Although proper cooking of meat is emphasized for reducing the risk for foodborne disease, the fact that C. difficile is a spore former complicates this issue because spores can survive in ground beef at recommended cooking temperatures (71°C), even when that temperature is maintained for 120 min (A. Rodriguez-Palacios et al., unpub. data).
The clinical and epidemiologic relevance of these microbiologic findings remains unknown. The isolation of C. difficile from meat samples does not necessarily mean that CDAD is a foodborne disease. Additional studies are required to determine the prevalence of contamination and its clinical relevance.
Dr Rodriguez-Palacios recently completed a doctorate degree in veterinary science at the Ontario Veterinary College, University of Guelph. His research interests include infectious diseases of large animals and the study of foodborne pathogens.
Acknowledgment
We thank Marie-Pierre Schippers and Gerardo Quiroz-Rocha for their help collecting samples, Joyce Rousseau for laboratory assistance, Maja Rupnik for toxinotyping of selected strains, and Angela Thompson, Brandy Limbago, and L. Clifford McDonald for PFGE and scientific collaboration.
References
- Pepin J, Alary ME, Valiquette L, Raiche E, Ruel J, Fulop K, Increasing risk of relapse after treatment of Clostridium difficile colitis in Quebec, Canada.Clin Infect Dis. 2005;40:1591–7. DOIPubMedGoogle Scholar
- Warny M, Pepin J, Fang A, Killgore G, Thompson A, Brazier J, Toxin production by an emerging strain of Clostridium difficile associated with outbreaks of severe disease in North America and Europe.Lancet. 2005;366:1079–84. DOIPubMedGoogle Scholar
- Pituch H, Brazier JS, Obuch-Woszczatynski P, Wultanska D, Meisel-Mikolajczyk F, Luczak M. Prevalence and association of PCR ribotypes of Clostridium difficile isolated from symptomatic patients from Warsaw with macrolide-lincosamide-streptogramin B (MLSB) type resistance.J Med Microbiol. 2006;55:207–13. DOIPubMedGoogle Scholar
- Rodriguez-Palacios A, Stämpfli H, Duffield T, Peregrine AS, Trotz-Williams LA, Arroyo LG, Clostridium difficile PCR ribotypes in calves, Canada.Emerg Infect Dis. 2006;12:1730–6.PubMedGoogle Scholar
- Baverud V. Clostridium difficile infections in animals with special reference to the horse. A review.Vet Q. 2002;24:203–19.PubMedGoogle Scholar
- Arroyo LG, Kruth SA, Willey BM, Staempfli HR, Low DE, Weese JS. PCR ribotyping of Clostridium difficile isolates originating from human and animal sources.J Med Microbiol. 2005;54:163–6. DOIPubMedGoogle Scholar
- Lefebvre SL, Arroyo LG, Weese JS. Epidemic Clostridium difficile strain in hospital visitation dog.Emerg Infect Dis. 2006;12:1036–7.PubMedGoogle Scholar
- Aspinall ST, Hutchinson DN. New selective medium for isolating Clostridium difficile from faeces.J Clin Pathol. 1992;45:812–4. DOIPubMedGoogle Scholar
- Lemee L, Dhalluin A, Testelin S, Mattrat MA, Maillard K, Lemeland JF, Multiplex PCR targeting tpi (triose phosphate isomerase), tcdA (toxin A), and tcdB (toxin B) genes for toxigenic culture of Clostridium difficile.J Clin Microbiol. 2004;42:5710–4. DOIPubMedGoogle Scholar
- Bidet P, Barbut F, Lalande V, Burghoffer B, Petit JC. Development of a new PCR-ribotyping method for Clostridium difficile based on ribosomal RNA gene sequencing.FEMS Microbiol Lett. 1999;175:261–6. DOIPubMedGoogle Scholar
- Rupnik M. Clostridium difficile toxinotypes. Maribor, Slovenia: University of Maribor. 2006 [cited 2006 May 9]. Available from http://www.mf.uni-mb.si/Mikro/tox
- Bishara J, Bloch Y, Garty M, Behor J, Samra Z. Antimicrobial resistance of Clostridium difficile isolates in a tertiary medical center, Israel.Diagn Microbiol Infect Dis. 2006;54:141–4. DOIPubMedGoogle Scholar
- Broda DM, DeLacy KM, Bell RG, Braggins TJ, Cook RL. Psychrotrophic Clostridium spp. associated with ‘blown pack’ spoilage of chilled vacuum-packed red meats and dog rolls in gas-impermeable plastic casings.Int J Food Microbiol. 1996;29:335–52. DOIPubMedGoogle Scholar
- Weese JS, Rousseau J, Arroyo L. Bacteriological evaluation of commercial canine and feline raw diets.Can Vet J. 2005;46:513–6.PubMedGoogle Scholar
- Terhes G, Brazier JS, Urban E, Soki J, Nagy E. Distribution of Clostridium difficile PCR ribotypes in regions of Hungary.J Med Microbiol. 2006;55:279–82. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 13, Number 3—March 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Alexander Rodriguez-Palacios, Department of Clinical Studies, University of Guelph, Guelph, Ontario N1G 2W1, Canada;
Top