Volume 14, Number 12—December 2008
Letter
Parachlamydia acanthamoebae Infection and Abortion in Small Ruminants
Cite This Article
Citation for Media
To the Editor: Abortion in ruminants is of worldwide economic importance. Moreover, several abortigenic agents have a zoonotic potential, i.e., Brucella abortus, Coxiella burnetii, and Chlamydophila abortus. C. abortus, which causes ovine enzootic abortion, may also infect pregnant women who have had contact with C. abortus–infected sheep and goats, and such infection can lead to miscarriage (1).
Parachlamydia acanthamoebae (2) is a Chlamydia-related organism considered as an emerging agent of pneumonia in humans. Recently, we reported its role in the setting of bovine abortion (3). Here, we investigated the prevalence of C. abortus and P. acanthamoebae infections in abortions in small ruminants.
Formalin-fixed placenta, fetal lung and liver, or both, were available from abortion products from 144 goats and 86 sheep (n = 211). These specimens had previously been investigated for several abortigenic agents (4). Placentas and fetal organs were analyzed by histopathologic examination and by specific real-time PCR and immunohistochemical protocols that detect members of the Chlamydiaceae family and P. acanthamoebae.
DNA from paraffin blocks was extracted as described (5) by using the DNeasy Tissue kit (QIAGEN, Hilden, Germany). The real-time PCR for Chlamydiaceae was conducted on an ABI 7500 (Applied Biosystems, Foster City, CA, USA) by using a modified version of Everett’s PCR (6). Primers Ch23S-F (5′-CTGAAACCAGTAGCTTATAAGCGGT-3′), Ch23S-R (5′-ACCTCGCCGTTTAACTTAACTCC-3′), and probe Ch23S-p (5′-FAM-CTCATCA TGCAAAAGGCACGCCG-TAMRA-3′) were used to amplify and detect a 111-bp product specific for members of the family Chlamydiaceae. Chlamydial species identification of real-time PCR positive cases was performed with the ArrayTube Microarray (Clondiag, Jena, Germany) as described (7).
The Parachlamydia-specific real-time PCR was performed with the ABI Prism 7000 sequence detection system (Applied Biosystems), as reported (8). This PCR is genus-specific, as demonstrated by the absence of PCR positivity with DNA extracted from other Parachlamydiaceae (Protochlamydia spp./Neochlamydia hartmannellae). To confirm positive results, another specific PCR, which targeted the tlc gene, was performed (9).
Paraffin sections from specimens positive in real-time PCR were further examined by immunohistochemical tests. A Chlamydiaceae-specific mouse monoclonal antibody directed against the chlamydial lipopolysaccharide (Progen, Heidelberg, Germany) and a specific mouse polyclonal antibody against Parachlamydia spp. was used as described (3,5,10). These antibodies were applied at dilutions of 1:200 and 1:1,000, respectively. Detection was performed with a detection kit (ChemMate; Dako, Glostrup, Denmark). Antigen retrieval was performed by enzyme digestion for 10 minutes (Pronase; Dako) for the Chlamydiaceae antibody and repeated microwave treatment in citrate buffer (ChemMate; Dako) for the Parachlamydia antibody, respectively. Double immunohistochemical labeling was performed on the sheep abortion specimen identified as simultaneously infected with Chlamydiaceae and Parachlamydia spp. Immunohistochemical analysis for both pathogens was performed subsequently by using diaminobenzidine as substrate for the Chlamydiaceae antibody (brown labeling) and by using 3-amino-9-ethylcarbazole as substrate for the Parachlamydia antibody (red labeling). Specificity of PCR and immunohistochemical tests for Chlamydiaceae and Parachlamydia spp., respectively, was assessed by using negative control placentas taken from 2 healthy ruminants (both specimens were negative in all tests).
Results of real-time PCR showed that 55 (26.1%) of 211 specimens were positive for Chlamydiaceae. All 55 cases could be identified as C. abortus by ArrayTube Microarray (Clondiag). Of these, 42 (76.4%) could be confirmed by immunohistochemical analysis with the anti-Chlamydiaceae antibody.
Of the 211 specimens, only 2 (0.9%) were positive for Parachlamydia spp. by real-time PCR, and both cases could be confirmed by immunohistochemical testing with the parachlamydial antibody. These 2 specimens were negative for other common abortigenic agents such as Toxoplasma gondii, C. burnetii, and border disease virus (data not shown). One case was recorded among the 144 goat samples investigated. This placenta displayed necrotizing placentitis and was positive for Parachlamydia spp. by 16S rRNA-specific real-time PCR (cycle threshold [Ct] 40.5) and immunohistochemical testing, but negative for Chlamydiaceae. Results of this PCR was confirmed by another PCR, targeting the tlc gene (Ct 36.7), which excluded false-positive results because of amplicon contamination.
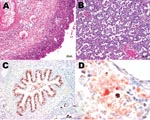
The second case was identified among the 86 sheep investigated. Placenta and fetal lung and liver exhibited necrotizing placentitis and vasculitis (Figure, panel A), interstitial pneumonia (Figure, panel B), and mixed cellular periportal hepatitis. Fetal liver was negative by parachlamydial 16S rRNA real-time PCR and immunohistochemical analysis, but the fetal lung was positive by parachlamydial 16S rRNA real-time PCR (Ct 40.7) and immunohistochemical tests (Figure, panel C), but negative with the tlc PCR. Fetal lung and liver were positive by real-time PCR for Chlamydiaceae (mean Ct for both organs 36.7), but negative by immunohistochemical tests. The placenta was positive for Chlamydiaceae by immunohistochemical tests and real-time PCR (mean Ct 23.3), and C. abortus was identified by ArrayTube Microarray. Brown (Chlamydiaceae) and red (Parachlamydia spp.) granular reaction was demonstrated within the necrotic lesions of the placenta by double immunohistochemical labeling (Figure, panel D).
We report Parachlamydia infection in small ruminant abortion. C. abortus and Parachlamydia spp. were simultaneously present in an aborted sheep placenta. Parachlamydia spp. could be further detected in the lung of the aborted sheep fetus by real-time PCR and immunohistochemistry. Parachlamydia was also detected in a goat placenta. Thus, Parachlamydia spp. should be considered as a new abortigenic agent in sheep and goats. Persons in contact with small ruminants should be informed about the zoonotic potential of these abortigenic agents.
Acknowledgments
We are grateful to the laboratory staff (in particular, Carmen Kaiser) of the Institute of Veterinary Pathology, Vetsuisse Faculty, University of Zurich.
This study was supported by the State Secretary for Education and Research, Bern, Switzerland (project number C05.0141), as part of the COST Action 855 (Animal Chlamydiosis and Zoonotic Implications).
G.G. is supported by the Leenards Foundation through a career award entitled “Bourse Leenards pour la relève académique en médecine clinique à Lausanne.”
References
- Longbottom D, Coulter LJ. Animal chlamydioses and zoonotic implications. J Comp Pathol. 2003;128:217–44. DOIPubMedGoogle Scholar
- Greub G, Raoult D. Parachlamydiaceae: potential emerging pathogens. Emerg Infect Dis. 2002;8:625–30.PubMedGoogle Scholar
- Borel N, Ruhl S, Casson N, Kaiser C, Pospischil A, Greub G. Parachlamydia spp. and related Chlamydia-like organisms and bovine abortion. Emerg Infect Dis. 2007;13:1904–7.PubMedGoogle Scholar
- Chanton-Greutmann H, Thoma R, Corboz L, Borel N, Pospischil A. Abortion in small ruminants in Switzerland: investigations during two lambing seasons (1996–1998) with special regard to chlamydial abortions [in German]. Schweiz Arch Tierheilkd. 2002;144:483–92. DOIPubMedGoogle Scholar
- Borel N, Thoma R, Spaeni P, Weilenmann R, Teankum K, Brugnera E, Chlamydia-related abortions in cattle from Graubunden, Switzerland. Vet Pathol. 2006;43:702–8. DOIPubMedGoogle Scholar
- Everett KD, Bush RM, Andersen AA. Emended description of the order Chlamydiales, proposal of Parachlamydiaceae fam. nov. and Simkaniaceae fam. nov., each containing one monotypic genus, revised taxonomy of the family Chlamydiaceae, including a new genus and five new species, and standards for the identification of organisms. Int J Syst Bacteriol. 1999;49:415–40.PubMedGoogle Scholar
- Borel N, Kempf E, Hotzel H, Schubert E, Torgerson P, Slickers P, Direct identification of chlamydiae from clinical samples using a DNA microarray–a validation study. Mol Cell Probes. 2008;22:55–64. DOIPubMedGoogle Scholar
- Casson N, Posfay-Barbe KM, Gervaix A, Greub G. New diagnostic real-time PCR for specific detection of Parachlamydia acanthamoebae DNA in clinical samples. J Clin Microbiol. 2008;46:1491–3. DOIPubMedGoogle Scholar
- Greub G, Berger P, Papazian L, Raoult D. Parachlamydiaceae as rare agents of pneumonia. Emerg Infect Dis. 2003;9:755–6.PubMedGoogle Scholar
- Casson N, Entenza JM, Greub G. Serological cross-reactivity between different Chlamydia-like organisms. J Clin Microbiol. 2007;45:234–6. DOIPubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 14, Number 12—December 2008
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Nicole Borel, Institute of Veterinary Pathology, Vetsuisse Faculty, University of Zurich, 8057 Zurich, Switzerland
Top