Volume 15, Number 12—December 2009
Dispatch
Bartonella rochalimae in Raccoons, Coyotes, and Red Foxes
Cite This Article
Citation for Media
Abstract
To determine additional reservoirs for Bartonella rochalimae, we examined samples from several wildlife species. We isolated B. rochalimae from 1 red fox near Paris, France, and from 11 raccoons and 2 coyotes from California, USA. Co-infection with B. vinsonii subsp. berkhoffii was documented in 1 of the coyotes.
Twelve Bartonella species/subspecies have been recognized as zoonotic agents (1,2), including B. rochalimae isolated from a woman who had traveled from the United States to Peru before becoming ill with fever, splenomegaly, mild anemia, and rash (3). B. rochalimae, previously described as a B. clarridgeiae–like organism, has also been isolated from rural domestic dogs and gray foxes (Urocyon cinereoargenteus) from northern California (4,5). A case of fatal endocarditis in a domestic dog was associated with a B. clarridgeiae–like strain (6), later determined to be identical to B. rochalimae (5). Recently, B. rochalimae DNA was detected in a dog from Greece (7). High (43%) prevalence of bacteremia observed in gray foxes in California suggests that they might act as a wildlife reservoir for this newly identified species. Furthermore, several B. clarridgeiae–like and B. rochalimae genes have been detected in fleas collected from humans (8), rodents (9,10), red foxes (Vulpes vulpes) (11), and the environment in the Democratic Republic of Congo (12) during a plague outbreak. To determine whether other wildlife reservoirs exist, we tested samples from 3 additional wildlife species: coyotes, raccoons, and red foxes.
From 1996 through 1999 in central coastal California, 21 Canis latrans coyotes (3 juveniles [<1 year of age] and 18 adults) and 42 Procyon lotor raccoons (11 juveniles and 31 adults) were trapped. In 2002, a blood sample was collected from a road-killed red fox near Paris, France. All samples were collected in EDTA tubes and frozen at −70°C until plated on heart infusion agar containing 5% rabbit blood and incubated in 5% CO2 at 35°C for up to 4 weeks (13); subsequently, extracted DNA was tested for Bartonella spp. by PCR. In addition, from May 2003 through September 2004, blood was collected from 42 red foxes (23 females and 19 males; 2 kits [<1 year] and 40 adults) in Israel, and extracted DNA was tested for Bartonella spp. by PCR.
Bartonella isolates from 2 (9.5%) coyotes (coyote 004 [7-month-old male] and coyote 22 [adult female captured in central California], which yielded 2 different-size colonies: coyote 22/sub1, large size; coyote 22/sub2, small size), 11 (26%) of the raccoons (7 adult females and 4 adult males), the 1 (100%) red fox from France, and DNA from the blood of 2 (5%) foxes from Israel were compared with B. rochalimae strains isolated from a human, rural dogs, and gray foxes. Bartonella isolates were analyzed by PCR restriction fragment length polymorphism (RFLP) of the 16S–23S intergenic transcribed spacer (ITS) region (all strains) and the gltA, rpoB and ftsZ genes (raccoons, gray foxes, coyotes, and dogs), as previously described (5). For the isolate from the red fox from France, extracted DNA was also amplified for fragments of the groEL gene by using the primer sets HSPps1, HSPps2, and HSPsp4 (11,14). Sequencing was done in both directions by using a fluorescence-based automated sequencing system (Davis Sequencing, Davis, CA, USA). Sequences were imported into Vector NTI Suite 9.0 software (Invitrogen, Carlsbad, CA, USA) to obtain a consensus sequence. Align X in Vector NTI was used for aligning sequence variants with each other and other known Bartonella spp. for each of the 4 genes. A neighbor-joining tree was constructed in MEGA version 3.0 (www.megasoftware.net) by concatenating the 4 sequences. Bootstrap replicates were performed to estimate node reliability of the phylogenetic tree; values were obtained from 1,000 randomly selected samples of the aligned sequence data. Sequence data for the groEL gene of the isolate from the fox in France (GenBank accession no. FJ545656) was compared with sequences of DNA extracted from fleas collected on 4 foxes from Hungary (11) and deposited in GenBank under accession no. DQ522300.
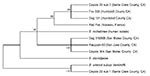
Amplified PCR products were obtained from the ITS region and the gltA, rpoB, and ftsZ genes of all isolates. Isolates from coyote 004 and coyote 22/sub2, the red fox from France, and the 11 raccoons had identical RFLP profiles, also identical to those observed in the rural dogs and gray foxes from California (5). Isolate sub1 from coyote 22 had banding patterns identical to those of Bartonella vinsonii subsp. berkhoffii (American Type Culture Collection 51672), indicating co-infection with B. vinsonii subsp. berkhoffii and B. rochalimae. Consensus sequences were obtained for isolates from all coyotes and the red fox from France. Because the RFLP profiles for the 11 raccoon isolates were identical, only 2 isolates (from raccoons 60 and 75, adult females from central California) were selected for sequencing and were identical for all genes. Partial sequences from the 4 genes were identical for the isolates from coyote 004 and the 2 raccoons. These isolates were 100% similar to a strain recovered from the dog with endocarditis (GenBank accession nos. DQ676488–DQ676491) (5,6). The B. rochalimae isolate sub2 from coyote 22 was identical to isolates from rural dogs and gray foxes from northern California (accession nos. DQ676484–DQ676487). Similarity of isolates from these regions ranged from 99.5% to 100% (Tables 1, 2). A tree constructed from the merged set of concatenated sequences (Figure 1) demonstrates that isolate sub2 from coyote 22 clustered with isolates from the dog and gray fox from northern California; those from coyote 004 and raccoon 60 grouped with those from the dog with endocarditis.
According to PCR results and comparison of a 571-bp sequence amplified from the ITS region, the sequences from 2 foxes (1 male, 1 female) from 2 villages in northern Israel were identical to each other and to that from the fox from France (Figure 2). Differences of 2–5 bp were observed among ITS region sequences when comparing those from the foxes from Israel and France with those from B. rochalimae from gray foxes and raccoons from California. When the groEL partial sequence FJ545656 from the red fox from France was compared with sequence DQ522300 from a Pulex irritans flea collected from foxes from Hungary, the 156-bp fragment (based on the consensus sequence from both directions) from the red fox from France was 100% identical to that of the flea.
We report the isolation or detection of B. rochalimae from red foxes, raccoons, and coyotes from North America, Europe, and the Middle East. Sequence analysis of 4 genes identified small variations in the B. rochalimae isolates from these different geographic regions. A relatively high percentage (26%) of raccoons had B. rochalimae bacteremia compared with only 9.5% (2/21) coyotes. A previous study found that of 109 coyotes, none were infected with B. rochalimae, but 31 (28%) harbored B. vinsonii subsp. berkhoffii (13). In raccoons, bacteremia was found in adults only, which is surprising because for all animals in general, Bartonella spp. bacteremia is detected more frequently in younger animals (1). Gray foxes from northern California had B. rochalimae bacteremia prevalence of 42% (4), suggesting that gray foxes and raccoons could be natural reservoirs of B. rochalimae in California and that infection of coyotes and domestic dogs could result from occasional spillover. Co-infection of a coyote also illustrates that wild canids can simultaneously harbor >1 species of Bartonella. Co-infection of humans with B. henselae and B. vinsonii subsp. berkhoffii has also been reported (15). Additionally, co-infection with 2 zoonotic Bartonella species in this coyote raises the possibility that humans and domestic dogs could also be co-infected with these species, making appropriate diagnosis more difficult. Pulex fleas collected on red foxes from Hungary (11) were indeed infected with a strain of Bartonella that was identical, at least for the groEL partial sequence, with that of the isolate from the red fox from France, suggesting that red foxes from central Europe may also be infected with B. rochalimae. Future studies with larger sample sizes will be needed to better define the role of these wild carnivores—red foxes, raccoons, and coyotes—in maintaining B. rochalimae in the environment.
Dr Henn is an epidemiologist with Napa County Health and Human Services and conducted this study for her PhD thesis. She has a strong interest in the epidemiology of emerging zoonoses.
Acknowledgments
We thank Darren Simpson, Laurie Frazer, Alain Henault, and Amit Dolev for their help with sample collection.
This work was supported by a grant from the Center for Companion Animal Health at the University of California, Davis, and by the American Kennel Club Canine Health Foundation.
References
- Chomel BB, Boulouis HJ, Maruyama S, Breitschwerdt EB. Bartonella spp. in pets and effect on human health. Emerg Infect Dis. 2006;12:389–94.PubMedGoogle Scholar
- Raoult D, Roblot F, Rolain JM, Besnier JM, Loulergue J, Bastides F, First isolation of Bartonella alsatica from a valve of a patient with endocarditis. J Clin Microbiol. 2006;44:278–9. DOIPubMedGoogle Scholar
- Eremeeva ME, Gerns HL, Lydy SL, Goo JS, Ryan ET, Mathew SS, Bacteremia, fever, and splenomegaly caused by a newly recognized Bartonella species. N Engl J Med. 2007;356:2381–7. DOIPubMedGoogle Scholar
- Henn JB, Gabriel MW, Kasten RW, Brown RN, Theis JH, Foley JE, Gray foxes (Urocyon cinereoargenteus) as a potential reservoir of a Bartonella clarridgeiae–like bacterium and domestic dogs as sentinels for zoonotic arthropod-borne pathogens in northern California. J Clin Microbiol. 2007;45:2411–8. DOIPubMedGoogle Scholar
- Henn JB, Gabriel MW, Kasten RW, Brown RN, Koehler JE, McDonald KA, Infective endocarditis in a dog and the phylogenetic relationship of the associated “Bartonella rochalimae” strain with isolates from dogs, gray foxes, and a human. J Clin Microbiol. 2009;47:787–90. DOIPubMedGoogle Scholar
- MacDonald KA, Chomel BB, Kittleson MD, Kasten RW, Thomas WP, Pesavento P. A prospective study of canine infective endocarditis in northern California (1999–2001): emergence of Bartonella as a prevalent etiologic agent. J Vet Intern Med. 2004;18:56–64.PubMedGoogle Scholar
- Diniz PP, Billeter SA, Otranto D, De Caprariis D, Petanides T, Mylonakis ME, Molecular documentation of Bartonella infection in dogs in Greece and Italy. J Clin Microbiol. 2009;47:1565–7. DOIPubMedGoogle Scholar
- Parola P, Shpynov S, Montoya M, Lopez M, Houpikian P, Zeaiter Z, First molecular evidence of new Bartonella spp. in fleas and a tick from Peru. Am J Trop Med Hyg. 2002;67:135–6.PubMedGoogle Scholar
- Loftis AD, Reeves WK, Szumlas DE, Abbassy MM, Helmy IM, Moriarity JR, Surveillance of Egyptian fleas for agents of public health significance: Anaplasma, Bartonella, Coxiella, Ehrlichia, Rickettsia, and Yersinia pestis. Am J Trop Med Hyg. 2006;75:41–8.PubMedGoogle Scholar
- Inoue K, Maruyama S, Kabeya H, Hagiya K, Izumi Y, Une Y, Exotic small mammals as potential reservoirs of zoonotic Bartonella spp. Emerg Infect Dis. 2009;15:526–32. DOIPubMedGoogle Scholar
- Sréter-Lancz Z, Tornyai K, Széll Z, Sréter T, Márialigeti K. Bartonella infections in fleas (Siphonaptera: Pulicidae) and lack of bartonellae in ticks (Acari: Ixodidae) from Hungary. Folia Parasitol (Praha). 2006;53:313–6.PubMedGoogle Scholar
- Sackal C, Laudisoit A, Kosoy M, Massung R, Eremeeva ME, Karpathy SE, Bartonella spp. and Rickettsia felis in fleas, Democratic Republic of Congo. Emerg Infect Dis. 2008;14:1972–4. DOIPubMedGoogle Scholar
- Chang CC, Kasten RW, Chomel BB, Simpson DC, Hew CM, Kordick DL, Coyotes (Canis latrans) as the reservoir for a human pathogenic Bartonella sp.: molecular epidemiology of Bartonella vinsonii subsp. berkhoffii infection in coyotes from central coastal California. J Clin Microbiol. 2000;38:4193–200.PubMedGoogle Scholar
- Zeaiter Z, Fournier PE, Raoult D. Genomic variation of Bartonella henselae strains detected in lymph nodes of patients with cat scratch disease. J Clin Microbiol. 2002;40:1023–30. DOIPubMedGoogle Scholar
- Breitschwerdt EB, Maggi RG, Duncan AW, Nicholson WL, Hegarty BC, Woods CW. Bartonella species in blood of immunocompetent persons with animal and arthropod contact. Emerg Infect Dis. 2007;13:938–41.PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 15, Number 12—December 2009
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Bruno B. Chomel, Department of Population Health and Reproduction, 1114 Tupper Hall, School of Veterinary Medicine, University of California, Davis, CA 95616, USA
Top