Volume 16, Number 6—June 2010
Dispatch
Novel Norovirus in Dogs with Diarrhea
Cite This Article
Citation for Media
Abstract
To identify the prevalence and genetic variability of noroviruses in dogs, we tested fecal samples by using reverse transcription–PCR. We found canine norovirus in 40% and 9% of dogs with and without diarrhea, respectively. The virus was genetically unrelated to other noroviruses and constitutes a tentative new genogroup.
Human noroviruses (NoVs) are the most frequent cause of epidemic and sporadic acute gastroenteritis worldwide among humans of all ages (1,2). The virus is transmitted through ingestion of contaminated food or water or from person to person through the fecal–oral route. The close genetic relatedness of swine NoV with human NoVs of genogroup (G) II suggests the potential for transfer from animals to humans (3–5). In addition, recent findings of viruses genetically related to human NoVs, as well as to animal NoV sequences in pigs and calves, have raised concerns about the possible emergence of recombinant viruses (4).
NoVs are genetically heterogeneous viruses that belong to the family Caliciviridae. The viral capsid encloses a single-stranded, positive-sense RNA genome of 7.3–7.7 kb that is organized in 3 open reading frames (ORFs), of which ORF1 encodes a polyprotein that is proteolytically cleaved into 6 nonstructural proteins, including RNA-dependent RNA polymerase (RdRp), helicase, and protease (1). ORF2 and ORF3 encode major (viral protein [VP] 1) and minor (VP2) capsid proteins. The GLPSG and YGDD motifs of the RdRp protein are conserved among all members of the family Caliciviridae (6). The genus Norovirus currently comprises 5 genogroups, designated GI–GV, which can be grouped into at least 32 genetic clusters (2,7). Only viruses from GI, GII, and GIV have been associated with human disease. Recently, the finding of a novel GIV norovirus in a young dog (5) indicated that pets can be infected with NoVs. To identify the prevalence and genetic variability of NoVs in dogs, we tested fecal specimens from dogs with and without diarrhea.
A total of 105 fecal samples from dogs in municipal dog shelters, veterinary clinics, and pet shops from 3 districts (Porto, Viseu, and Guarda) in Portugal were collected during December 2007–November 2008. Veterinarians evaluated the dogs for diarrhea at the time of the visit. The fecal panel consisted of 63 samples from dogs with diarrhea and 42 samples from dogs with formed, normal brown feces (i.e., controls). All samples were kept at –20°C until processed. Fecal suspensions (10%) were made in phosphate-buffered saline pH 7.2, and solids were removed by centrifugation at 8,000 × g for 5 min. Nucleic acid was extracted by using the QIAamp viral RNA mini kit (QIAGEN, Hilden, Germany) according to the manufacturer's instructions and tested for the presence of NoV RNA by 2 broadly reactive NoV conventional reverse transcription–PCR (RT-PCR) assays (8,9), selective for a partial RdRp region of the genome, by using One-Step RT-PCR kit (QIAGEN) and 37°C as annealing temperature.
Two (2%) of the 105 fecal samples tested positive for NoV with primer pair JV12y/13i, whereas no RT-PCR products were found using the p289-p290 primer pair. The 2 JV12y/13i sequences were identical and contained the GLPSG amino acid motif characteristic of viral RNA polymerases.
We designed specific canine norovirus oligonucleotide primers JV102 (5′-TGG GAT TCA ACA CAG CAG AG-3′) and JV103 (5′-TGC GCA ATA GAG TTG ACC TG-3′) and retested all 105 samples. Fecal samples from 25 (40%) of the 63 dogs with diarrhea and 4 (9%) of the 42 controls tested positive for a new canine NoV (Viseu strain) (Table). An ≈3.3-kb fragment from the RdRp region in ORF1 to the poly-A tail, including the complete ORF2 and ORF3 genes, was generated by long-template RT-PCR by using previously described methods (10). Gel-purified PCR products were cloned and sequenced by primer walking, and a 3,357-nt sequence including partial ORF1, full-length ORF2 and ORF3, and the 3′ end noncoding region from the Viseu strain was obtained. The consensus sequence from 5 different clones was submitted to GenBank and assigned accession no. GQ443611. All 105 nucleic acid extracts also were tested by RT-PCR for the recently reported canine NoV strain (5), canine coronavirus (CCV) (11), and by PCR for canine parvovirus (CPV-2) combining oligonucleotide primers for the detection of types 2a and 2b (12). None of the samples tested positive for the reported GIV.2 canine NoV strain (GIV.2/170/07/ITA). A total of 32 (51%) samples from dogs with diarrhea tested positive for CCV, whereas 2 (5%) of the control samples were positive. In addition, 58 (55%) samples tested positive for CPV-2, of which 18 also were positive for the novel canine NoV (Table). Univariate analysis (SPSS version 17.0, SPSS Inc., Chicago, IL, USA) demonstrated that the novel canine NoV and CCV were significantly associated with diarrhea (Table).
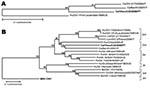
We detected novel norovirus sequences in 40% of samples from dogs with diarrhea and 9% of specimens from dogs without diarrhea. All canine norovirus RdRp sequences had a high nucleotide sequence identity (range 98%–100%) and differed 19%–22% from recently reported canine norovirus strains (5,13) (Figure, panel A).
In the complete VP1, the Viseu strain was most closely related to canine NoV strain Bari/91/07/IT (63.2% amino acid identity) and human strain Chiba/040502/2004/JP (55.1% identity) (Figure, panel B). Therefore, we tentatively classifed the Viseu strain, together with the Bari and Chiba strains, as a novel genogroup (GVI). To elucidate the potential pathogenic enteric role of the new canine NoV, we also tested all fecal specimens for CPV-2 and CCV, which are well-established enteric pathogens of dogs. Our results show that the novel canine NoV and CCV, but not CPV-2, were significantly associated with diarrhea.
For several reasons, our findings should be interpreted with caution. Viral shedding from dogs without diarrhea could represent asymptomatic infection, a resolution stage of the disease, or detection of CPV-2 vaccine virus. In addition, other canine enteric pathogens for which we did not test could bias the results. Further research that includes electron microscopy or infectivity studies is needed to confirm the existence of this new canine NoV.
The new virus sequences were detected in dogs from several dog owners throughout central Portugal and from a dog shelter in Viseu. Because Portuguese law requires official shelters to provide sanctuary for unwanted animals and keep them as adoptable pets for a certain period, humans potentially could be exposed to canine NoV that could increase the possibility for the emergence of canine/human NoV recombinants and increase the likelihood of zoonotic transmission. Additional serologic studies are needed to determine the level of exposure to canine NoVs in dogs and humans similar to previous studies on bovine NoV (14). Zoonotic transmission between dogs and humans is not new, and the close and often intimate interactions between these 2 species have been suggested as a major disease risk for humans (15). Only a few studies on NoV infections in dogs have been reported (5,13). Experimental infections in gnotobiotic dogs could ultimately provide essential data for describing the prevalence of canine NoVs in dogs and their causal relationship with diarrhea.
Dr Mesquita is a veterinary assistant at the Agrarian Superior School of Viseu and a PhD fellow at the Faculty of Pharmacy, University of Porto. His research interests include the epidemiology of enteric viruses in animals and humans.
Acknowledgments
We thank Jennifer Cannon and Nicole Gregoricus for technical support and Veronica Costantini for statistical analysis.
The study was partly supported by grant SFRH/BD/45407/2008 to J.R.M. from Fundação para a Ciência e a Tecnologia.
References
- Glass RI, Parashar UD, Estes MK. Norovirus gastroenteritis. N Engl J Med. 2009;361:1776–85. DOIPubMedGoogle Scholar
- Patel MM, Hall AJ, Vinjé J, Parashar UD. Noroviruses: a comprehensive review. J Clin Virol. 2009;44:1–8. DOIPubMedGoogle Scholar
- van der Poel WH, Vinjé J, van der Heide R, Herrera MI, Vivo A, Koopmans MP. Norwalk-like calicivirus genes in farm animals. Emerg Infect Dis. 2000;6:36–41.PubMedGoogle Scholar
- Mattison K, Shukla A, Cook A, Pollari F, Friendship R, Kelton D, Human noroviruses in swine and cattle. Emerg Infect Dis. 2007;13:1184–8.PubMedGoogle Scholar
- Martella V, Lorusso E, Decaro N, Elia G, Radogna A, D’Abramo M, Detection and molecular characterization of a canine norovirus. Emerg Infect Dis. 2008;14:1306–8. DOIPubMedGoogle Scholar
- Farkas T, Sestak K, Wei C, Jiang X. Characterization of a rhesus monkey calicivirus representing a new genus of Caliciviridae. J Virol. 2008;82:5408–16. DOIPubMedGoogle Scholar
- Martella V, Campolo M, Lorusso E, Cavicchio P, Camero M, Bellacicco A, Norovirus in captive lion cub (Panthera leo). Emerg Infect Dis. 2007;13:1071–3.PubMedGoogle Scholar
- Jiang X, Huang PW, Zhong WM, Farkas T, Cubitt DW, Matson DO. Design and evaluation of a primer pair that detects both Norwalk- and Sapporo-like caliciviruses by RT-PCR. J Virol Methods. 1999;83:145–54. DOIPubMedGoogle Scholar
- Vennema H, Bruin E, Koopmans M. Rational optimization of generic primers used for Norwalk-like virus detection by reverse transcriptase polymerase chain reaction. J Clin Virol. 2002;25:233–5. DOIPubMedGoogle Scholar
- Vinjé J, Koopmans MPG. Simultaneous detection and genotyping of “Norwalk-like viruses” by oligonucleotide array in a reverse line blot hybridization format. J Clin Microbiol. 2000;38:2595–601.PubMedGoogle Scholar
- Pratelli A, Tempesta M, Greco G, Martella V, Buonavoglia C. Development of a nested PCR assay for the detection of canine coronavirus. J Virol Methods. 1999;80:11–5. DOIPubMedGoogle Scholar
- Pereira CA, Monezi TA, Mehnert DU, D’Angelo M, Durigon EL. Molecular characterization of canine parvovirus in Brazil by polymerase chain reaction assay. Vet Microbiol. 2000;75:127–33. DOIPubMedGoogle Scholar
- Martella V, Decaro N, Lorusso E, Radogna A, Moschidou P, Amorisco F. Genetic heterogeneity and recombination in canine noroviruses. J Virol. 2009;83:11391–6. DOIPubMedGoogle Scholar
- Widdowson MA, Rockx B, Schepp R, van der Poel WH, Vinje J, van Duynhoven YT, Detection of serum antibodies to bovine norovirus in veterinarians and the general population in the Netherlands. J Med Virol. 2005;76:119–28. DOIPubMedGoogle Scholar
- Grøndalen J, Sævik B, Sørum H. Companion animals as reservoir for zoonotic diseases. European Journal of Companion Animal Practice. 2008;18:213–22.
Figure
Table
Cite This ArticleTable of Contents – Volume 16, Number 6—June 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jan Vinjé, Centers for Disease Control and Prevention, 1600 Clifton Rd NE, Mailstop G04, Atlanta, GA 30333, USA
Top