Volume 18, Number 10—October 2012
CME ACTIVITY - Research
Methicillin-Resistant Staphylococcus aureus Sequence Type 239-III, Ohio, USA, 2007–20091
Figure 3
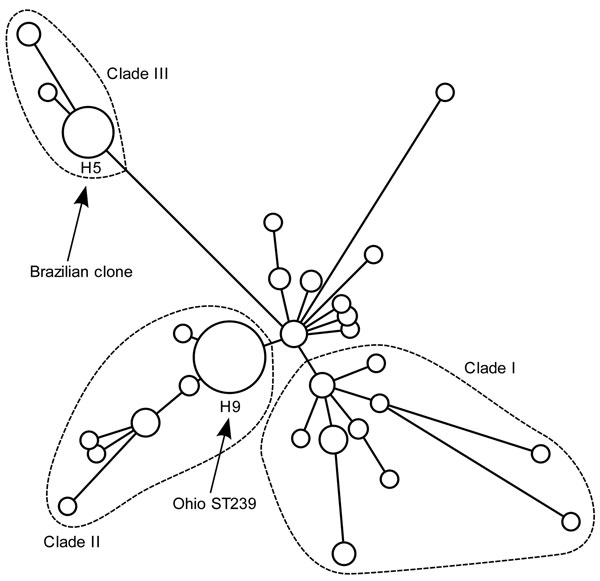
Figure 3. . . Single-nucleotide polymorphism (SNP) haplotype map showing position of methicillin-resistant Staphylococcus aureus sequence type 239-III (MRSA ST239-III) isolates, Ohio, USA, 2007–2009, within the global population structure of the MRSA ST239-III clonal group. Circles indicate distinct haplotypes, as defined by using a panel of 43 SNPs (9). Sizes of circles indicate relative frequency of different haplotypes. Arrows indicate haplotype 5 (H5), which includes the Brazilian clone, and haplotype 9 (H9), which includes the 22 MRSA ST239-III isolates from Ohio. Relationships between haplotypes were determined by using maximum-parsimony analysis (9).
References
- Cosgrove SE, Sakoulas G, Perencevich EN, Schwaber MJ, Karchmer AW, Carmeli Y. Comparison of mortality associated with methicillin-resistant and methicillin-susceptible Staphylococcus aureus bacteremia: a meta-analysis. Clin Infect Dis. 2003;36:53–9. DOIPubMedGoogle Scholar
- Melzer M, Eykyn SJ, Gransden WR, Chinn S. Is methicillin-resistant Staphylococcus aureus more virulent than methicillin-susceptible S. aureus? A comparative cohort study of British patients with nosocomial infection and bacteremia. Clin Infect Dis. 2003;37:1453–60. DOIPubMedGoogle Scholar
- McDougal LK, Steward CD, Killgore GE, Chaitram JM, McAllister SK, Tenover FC. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: establishing a national database. J Clin Microbiol. 2003;41:5113–20. DOIPubMedGoogle Scholar
- Feil EJ, Nickerson EK, Chantratita N, Wuthiekanum V, Srisomang P, Cousind R, Rapid detection of the pandemic methicillin-resistant Staphylococcus aureus clone ST 239, a dominant strain in Asian hospitals. J Clin Microbiol. 2008;46:1520–2. DOIPubMedGoogle Scholar
- Holden MT, Lindsay JA, Corton C, Quail MA, Cockfield JD, Pathak S, Genome sequence of a recently emerged, highly transmissible, multi-antibiotic- and antiseptic-resistant variant of methicillin-resistant Staphylococcus aureus, sequence type 239 (TW). J Bacteriol. 2010;192:888–92. DOIPubMedGoogle Scholar
- Aires de Sousa M, Conceição T, Simas C, de Lencastre H. Comparison of genetic backgrounds of methicillin-resistant and -susceptible Staphylococcus aureus isolates from Portuguese hospitals and the community. J Clin Microbiol. 2005;43:5150–7. DOIPubMedGoogle Scholar
- Monecke S, Coombs G, Shore AC, Coleman DC, Akpaka P, Borg M, A field guide to pandemic, epidemic and sporadic clones of methicillin-resistant Staphylococcus aureus. PLoS ONE. 2011;6:e17936. DOIPubMedGoogle Scholar
- Smyth DS, McDougal LK, Gran FW, Manoharan A, Enright MC, Song JH, Population structure of a hybrid clonal group of methicillin-resistant Staphylococcus aureus, ST239-MRSA-III. PLoS ONE. 2010;5:e8582. DOIPubMedGoogle Scholar
- de Lencastre H, Severina EP, Roberts RB, Kreiswirth BN, Tomasz A. Testing the efficacy of a molecular surveillance network: methicillin-resistant Staphylococcus aureus (MRSA) and vancomycin-resistant Enterococcus faecium (VREF) genotypes in six hospitals in the metropolitan New York City area. The BARG Initiative Pilot Study Group. Bacterial Antibiotic Resistance Group. Microb Drug Resist. 1996;2:343–51. DOIPubMedGoogle Scholar
- Klevens RM, Morrison MA, Nadle J, Petit S, Gershman K, Ray S, Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA. 2007;298:1763–71. DOIPubMedGoogle Scholar
- Chung M, Dickinson G, de Lencastre H, Tomasz A. International clones of methicillin-resistant Staphylococcus aureus in two hospitals in Miami, Florida. J Clin Microbiol. 2004;42:542–7. DOIPubMedGoogle Scholar
- Tenover FC, Tickler IA, Goering RV, Kreiswirth BN, Mediavilla JR, Persing DH. Characterization of nasal and blood culture isolates of methicillin-resistant Staphylococcus aureus from patients in United States hospitals. Antimicrob Agents Chemother. 2012;56:1324–30. DOIPubMedGoogle Scholar
- Mermel LA, Eells SJ, Acharya MK, Cartony JM, Dacus D, Fadem S, Quantitative analysis and molecular fingerprinting of methicillin-resistant Staphylococcus aureus nasal colonization in different patient populations: a prospective, multicenter study. Infect Control Hosp Epidemiol. 2010;31:592–7. DOIPubMedGoogle Scholar
- Clincal and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. 19th informational supplement, vol 29, no. 3. M100–S19. Wayne (PA): The Institute; 2009.
- Healy M, Huong J, Bittner T, Lising M, Frye S, Raza S, Microbial DNA typing by automated repetitive-sequence-based PCR. J Clin Microbiol. 2005;43:199–207. DOIPubMedGoogle Scholar
- Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.PubMedGoogle Scholar
- Shopsin B, Gomez M, Montgomery SO, Smith DH, Waddington M, Dodge DE, Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J Clin Microbiol. 1999;37:3556–63.PubMedGoogle Scholar
- Mathema B, Mediavilla J, Kreiswirth BN. Sequence analysis of the variable number tandem repeat in Staphylococcus aureus protein A gene: spa typing. Methods Mol Biol. 2008;431:285–305.PubMedGoogle Scholar
- Chen L, Mediavilla JR, Oliveira DC, Willey BM, de Lencastre H, Kreiswirth BN. Multiplex real-time PCR for rapid staphylococcal cassette chromosome mec typing. J Clin Microbiol. 2009;47:3692–706. DOIPubMedGoogle Scholar
- Goering RV, Morrison D, Al-Doori Z, Edwards GF, Gemmell CG. Usefulness of mec-associated direct repeat unit (dru) typing in the epidemiological analysis of highly clonal methicillin-resistant Staphylococcus aureus in Scotland. Clin Microbiol Infect. 2008;14:964–9. DOIPubMedGoogle Scholar
- Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol. 2000;38:1008–15.PubMedGoogle Scholar
- Saïd-Salim B, Mathema B, Braughton K, Davis S, Sinsimer D, Eisner W, Differential distribution and expression of Panton-Valentine leucocidin among community-acquired methicillin-resistant Staphylococcus aureus strains. J Clin Microbiol. 2005;43:3373–9. DOIPubMedGoogle Scholar
- Johnson WM, Tyler SD, Ewan EP, Ashton FE, Pollard DR, Rozee KR. Detection of genes for enterotoxins, exfoliative toxins, and toxic shock syndrome toxin 1 in Staphylococcus aureus by the polymerase chain reaction. J Clin Microbiol. 1991;29:426–30.PubMedGoogle Scholar
- Diep BA, Stone GG, Basuino L, Graber CJ, Miller A, des Etages SA, The arginine catabolic mobile element and staphylococcal chromosomal cassette mec linkage: convergence of virulence and resistance in the USA300 clone of methicillin-resistant Staphylococcus aureus. J Infect Dis. 2008;197:1523–30. DOIPubMedGoogle Scholar
- Anthony RM, Connor AM, Power EG, French GL. Use of the polymerase chain reaction for rapid detection of high-level mupirocin resistance in staphylococci. Eur J Clin Microbiol Infect Dis. 1999;18:30–4. DOIPubMedGoogle Scholar
- Harris SR, Feil EJ, Holden MT, Quail MA, Nickerson EK, Chantratita N, Evolution of MRSA during hospital transmission and intercontinental spread. Science. 2010;327:469–74. DOIPubMedGoogle Scholar
- Amaral MM, Coelho LR, Flores RP, Souza RR, Silva-Carvalho MC, Teixeira LA, The predominant variant of the Brazilian epidemic clonal complex of methicillin-resistant Staphylococcus aureus has an enhanced ability to produce biofilm and to adhere to and invade airway epithelial cells. J Infect Dis. 2005;192:801–10. DOIPubMedGoogle Scholar
- Edgeworth JD, Yadegarfar G, Pathak S, Batra R, Cockfield JD, Wyncoll D, An outbreak in an intensive care unit of a strain of methicillin-resistant Staphylococcus aureus sequence type 239 associated with an increased rate of vascular access device-related bacteremia. Clin Infect Dis. 2007;44:493–501. DOIPubMedGoogle Scholar
- Cho DT, Cha HY, Chang HH, Kim SW, Chung JM, Kim J, Risk factors for specific methicillin-resistant Staphylococcus aureus clones in a Korean hospital. J Antimicrob Chemother. 2006;57:1122–7. DOIPubMedGoogle Scholar
- Li DZ, Chen YS, Yang JP, Zhang W, Hu CP, Li JS, Preliminary molecular epidemiology of the Staphylococcus aureus in lower respiratory tract infections: a multicenter study in China. Chin Med J (Engl). 2011;124:687–92.PubMedGoogle Scholar
- Smyth DS, Wong A, Robinson DA. Cross-species spread of SCCmec IV subtypes in staphylococci. Infect Genet Evol. 2011;11:446–53. DOIPubMedGoogle Scholar
- Van Bellum A, Struelens M, de Visser A, Verbrugh H, Tibayrenc M. Role of genomic typing in taxonomy, evolutionary genetics, and microbial epidemiology. Clin Microbiol Rev. 2001;14:547–60. DOIPubMedGoogle Scholar
- Olive DM, Bean P. Principles and applications of methods for DNA-based typing of microbial organisms. J Clin Microbiol. 1999;37:1661–9.PubMedGoogle Scholar
- Tumin R, Wang SH, Pancholi P, Khan Y, Hines L, Stevenson K. Relationship of social and medical factors to other antimicrobial resistance among methicillin-resistant Staphylococcus aureus strains. In: Abstracts of the 139th American Public Health Association Annual Meeting; Washington, DC, Oct 29–Nov 2, 2011. Washington (DC): American Public Health Association; 2011. Abstract 246417.
1Presented in part at the 48th Annual Meeting of the Infectious Diseases Society of America, Vancouver, British Columbia, Canada, October 21–24, 2010.