Volume 19, Number 12—December 2013
Dispatch
Novel Variants of Clade 2.3.4 Highly Pathogenic Avian Influenza A(H5N1) Viruses, China
Cite This Article
Citation for Media
Abstract
We characterized 7 highly pathogenic avian influenza A(H5N1) viruses isolated from poultry in China during 2009–2012 and found that they belong to clade 2.3.4 but do not fit within the 3 defined subclades. Antigenic drift in subtype H5N1 variants may reduce the efficacy of vaccines designed to control these viruses in poultry.
Infection with the Asian lineage of highly pathogenic avian influenza (HPAI) A(H5N1) virus (prototype strain A/goose/Guangdong/1/96 [Gs/GD]) has resulted in substantial losses in the poultry industry and poses a threat to public health worldwide. According to the World Health Organization (WHO)/World Organisation for Animal Health (OIE)/Food and Agriculture Organization of the United Nations (FAO) H5N1 Evolution Working Group, 10 distinct clades of these viruses (0–9) were initially designated in 2008 to characterize the phylodynamics of the hemagglutinin (HA) gene of the Gs/GD-like viruses circulating during 1996–2007 (1). On the basis of these nomenclature criteria, new second-, third-, and fourth-order clades have been identified within the previously defined clades in the phylogenetic analyses that were updated in 2009 and 2011 (2,3). Therefore, as HPAI A(H5N1) virus continues to undergo substantial evolution, extensive genetic divergence is expected to periodically accumulate to form novel monophyletic groups. To identify continued divergence of clade 2.3.4 other than the recognized subclades 2.3.4.1, 2.3.4.2, and 2.3.4.3, we characterized 7 HPAI A(H5N1) viruses isolated from poultry during 2009–2012 in China (2).
As part of continuous national avian influenza virus surveillance, we performed a monthly collection of cloacal swabs from various poultry species (chicken, duck, goose, quail, and pigeon) at a wholesale live-bird market (LBM) in Yangzhou, Jiangsu Province, in eastern China. Birds offered for retail sale in the LBM were mainly from local farms in Jiangsu and the neighboring provinces in eastern China; some were transported from regions in southern or northern China. Virus isolation and identification were conducted as described (4). During December 2009–September 2012, avian influenza virus isolates belonging to 8 HA subtypes (H1, H3–H6, H9–H11) were identified; 7 of the isolates were subtype H5N1 (Table 1).
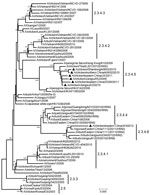
To characterize these 7 isolates, we sequenced the HA genes to determine clade distribution. In the reconstructed phylogenetic tree (Figure) using reference sequences retrieved from the GenBank database and partially recommended by WHO/OIE/FAO (2), the 7 isolates belonged to clade 2.3.4 (the “Fujian-like” sublineage), which has been prevalent in China since 2005 (5). However, none of the isolates could be further classified into previously identified subclades 2.3.4.1, 2.3.4.2, or 2.3.4.3. Six of the viruses closely resembled A/peregrine falcon/Hong Kong/810/2009, and the remaining virus was highly homologous with recent H5 viruses bearing various neuraminidase (NA) subtypes (N1, N2, N5, and N8).
According to WHO/OIE/FAO guidelines (1–3), new clades (including subclades) were specified not only with a bootstrap value of >60 at the clade-defining node in which sequences monophyletically arose from a common ancestor but also with average between-clade and within-clade nucleotide divergences of >1.5% and <1.5%, respectively. Apart from subclades 2.3.4.1, 2.3.4.2, and 2.3.4.3, we found 3 additional monophyletic categories—the A/peregrine falcon/Hong Kong/810/2009-like viruses, the HPAI subtype H5N5–like reassortants, and the HPAI subtype H5N2/H5N8–like reassortants—that grouped clearly within the tree (Figure). The bootstrap values and average within-clade and between-clade distances for these 3 groups were 81, 1.0%, 4.2%; 100, 1.0%, 5.2%; and 100, 1.3%, 5.3%, respectively.
Because of the compulsory vaccination practice against HPAI in China (6), we examined serologic cross-reactivity between the 7 subtype H5N1 isolates and the diagnostic antigen of the widely used inactivated reassortant H5N1/PR8 vaccine Re-5 (Table 1). Although Re-5 derived its HA and NA genes from a clade 2.3.4 representative virus A/duck/Anhui/1/2006, the hemagglutination inhibition (HI) titers of Re-5 antiserum against the 7 subtype H5N1 viruses were as much as 6–7 log2 lower than that against the homologous antigen. In contrast, the antiserum of A/chicken/northern China/k0602/2010 (k0602) showed limited reaction to Re-5 and A/chicken/eastern China/AH/2012. Moreover, antigenic variation also existed among the 6 A/peregrine falcon/Hong Kong/810/2009-like viruses, as highlighted by the HI assay using k0602 antiserum (Table 1).
To explore whether these antigenic variations can be translated into protection efficacy difference in vivo, we selected A/chicken/northern China/k0602/2010 (k0602) and A/chicken/Shandong/k0603/2010 (k0603) viruses to evaluate the bivalent inactivated reassortant H5N1/PR8 vaccine Re-4/Re-5 (the HA and NA genes of Re-4 are from a clade 7 virus A/chicken/Shanxi/2/2006). This vaccine has been extensively used to control the prevalence of clade 2.3.4 and clade 7 viruses in China since 2008 (6). In addition, a reassortant rk0602 virus, which carries the HA and NA genes of k0602 virus and the internal genes of PR8, was recovered by using reverse genetics and the inactivated rk0602 vaccine was applied to evaluate the homologous protection. Four-week-old specific-pathogen free chickens were vaccinated with Re-4/Re-5 or the rk0602 vaccine and readily developed specific antibodies against the component viruses by day 28 after vaccination (Table 2). The birds were then intranasally challenged with 106.0 50% egg infectious dose of k0602 or k0603 virus. During the 10-day observation period, the Re-4/Re-5 vaccinated birds displayed clinical signs including severe depression, ruffled feathers, huddling, decreased feed and water consumption, and diarrhea; moreover, only 14.3% (1/7 birds in the k0602 group) and 10% (1/10 birds in the k0603 group) of the challenged chickens survived, reflecting poor protection by the Re-4/Re-5 vaccine. In addition, shed virus was detected in tracheal and cloacal swabs from most of the tested chickens on 3 and 5 days postchallenge. By contrast, the rk0602-vaccinated chickens all survived the challenge, and no virus was recovered from tracheal and cloacal samples (Table 2).
The location of the 7 HPAI A(H5N1) virus variants in the HA gene tree (Figure) suggests that novel monophyletic subclades other than the previously identified 2.3.4.1, 2.3.4.2, and 2.3.4.3 subclades continue to emerge within clade 2.3.4. As a result of our findings, we suggest that these groups should be assigned new fourth-order clades of 2.3.4.4, 2.3.4.5, and 2.3.4.6 to reflect the wide divergence of clade 2.3.4 viruses.
In China, 1 of the 6 countries to which subtype H5N1 virus is endemic (7), multiple distinct clades (2.2, 2.5, 2.3.1, 2.3.2, 2.3.3, 2.3.4, 7, 8, and 9) were identified by surveillance during 2004–2009 (5). In particular, clades 2.3.2, 2.3.4, and 7 viruses have gained ecologic niches and have continued circulating by further evolving into new subclades (2). In addition, various NA subtypes of H5 viruses (H5N5, H5N8, and H5N2) bearing the genetic backbone of clade 2.3.4 A(H5N1) viruses have been detected in ducks, geese, quail, and chickens (8–12). These findings highlight the importance of periodic updates of the WHO/OIE/FAO classification of Asian A(H5N1) viruses by continuous surveillance to better understand the dynamic nature of the viral evolution.
Our findings have implications for the effectiveness of vaccination of chickens against HPAI A(H5N1) viruses. The results of cross-HI assays (Table 1) and vaccine efficacy experiments (Table 2) indicate antigenic drift in subtype H5N1 variants, as compared with the vaccine strain specifically designed to control the prevalent clade 2.3.4 virus infection in poultry. Although previous studies by Tian et al. (13) and Kumar et al. (14) proposed that vaccinated chickens with HI antibody titers of >4 log2 could be protected from virus challenge, our data demonstrate that vaccine efficacy is substantially influenced by antigenic matching between the vaccine strain and circulating viruses in preventing the replication and transmission of influenza virus, especially when the induced antibodies are of moderate titers.
Dr Gu is a faculty member at the College of Veterinary Medicine, Yangzhou University. Her primary research interest is the epidemiology and evolutionary mechanism of avian influenza viruses.
Acknowledgment
This work was supported by the Major State Basic Research Development Program of China (973 Program; (grant no. 2011CB505003), the Jiangsu Provincial Natural Science Foundation of China (grant no. BK20130442), the earmarked fund for Modern Agro-industry Technology Research System (nycytx-41-G07), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
References
- WHO/OIE/FAO H5N1 Evolution Working Group. Toward a unified nomenclature system for highly pathogenic avian influenza virus (H5N1). Emerg Infect Dis. 2008;14:e1. DOIGoogle Scholar
- WHO/OIE/FAO H5N1 Evolution Working Group. Continued evolution of highly pathogenic avian influenza A (H5N1): updated nomenclature. Influenza Other Respi Viruses. 2012;6:1–5. DOIGoogle Scholar
- WHO/OIE/FAO H5N1 Evolution Working Group. Continuing progress towards a unified nomenclature for the highly pathogenic H5N1 avian influenza viruses: divergence of clade 2.2 viruses. Influenza Other Respi Viruses. 2009;3:59–62 . DOIGoogle Scholar
- Zhao G, Pan J, Gu X, Zhao K, Chen H, Chen S, Prevalence of low pathogenic avian influenza virus in one live bird market in Eastern China from 2002–2011. Avian Dis. 2013;57:155–8. DOIPubMedGoogle Scholar
- Li Y, Shi J, Zhong G, Deng G, Tian G, Ge J, Continued evolution of H5N1 influenza viruses in wild birds, domestic poultry, and humans in China from 2004 to 2009. J Virol. 2010;84:8389–97. DOIPubMedGoogle Scholar
- Chen H. Avian influenza vaccination: the experience in China. Rev Sci Tech. 2009;28:267–74 .PubMedGoogle Scholar
- Food and Agriculture Organization of the United Nations. Stamping out H5N1 avian influenza could take decades. 2011 [cited 2013 Oct 24]. http://www.fao.org/news/story/en/item/66118
- Zhao K, Gu M, Zhong L, Duan Z, Zhang Y, Zhu Y, Characterization of three H5N5 and one H5N8 highly pathogenic avian influenza viruses in China. Vet Microbiol. 2013;163:351–7. DOIPubMedGoogle Scholar
- Gu M, Huang J, Chen Y, Chen J, Wang X, Liu X. Genome sequence of a natural reassortant H5N2 avian influenza virus from domestic mallard ducks in eastern China. J Virol. 2012;86:12463–4. DOIPubMedGoogle Scholar
- Gu M, Liu W, Cao Y, Peng D, Wang X, Wan H, Novel reassortant highly pathogenic avian influenza (H5N5) viruses in domestic ducks, China. Emerg Infect Dis. 2011;17:1060–3 .PubMedGoogle Scholar
- Zhao G, Gu X, Lu X, Pan J, Duan Z, Zhao K, Novel reassortant highly pathogenic H5N2 avian influenza viruses in poultry in China. PLoS ONE. 2012;7:e46183. DOIPubMedGoogle Scholar
- Zou W, Guo X, Li S, Yang Y, Jin M. Complete genome sequence of a novel natural recombinant H5N5 influenza virus from ducks in central China. J Virol. 2012;86:13878. DOIPubMedGoogle Scholar
- Tian G, Zhang S, Li Y, Bu Z, Liu P, Zhou J, Protective efficacy in chickens, geese and ducks of an H5N1-inactivated vaccine developed by reverse genetics. Virology. 2005;341:153–62. DOIPubMedGoogle Scholar
- Kumar M, Chu HJ, Rodenberg J, Krauss S, Webster RG. Association of serologic and protective responses of avian influenza vaccines in chickens. Avian Dis. 2007;51(Suppl):481–3. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 19, Number 12—December 2013
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Xiufan Liu, College of Veterinary Medicine, Yangzhou University, 48 East Wenhui Road, Yangzhou, Jiangsu, 225009, ChinaXiufan Liu, College of Veterinary Medicine, Yangzhou University, 48 East Wenhui Road, Yangzhou, Jiangsu, 225009, ChinaXiufan Liu, College of Veterinary Medicine, Yangzhou University, 48 East Wenhui Road, Yangzhou, Jiangsu, 225009, China
Top