Volume 19, Number 6—June 2013
Research
Active Surveillance for Influenza A Virus among Swine, Midwestern United States, 2009–2011
Cite This Article
Citation for Media
Abstract
Veterinary diagnostic laboratories identify and characterize influenza A viruses primarily through passive surveillance. However, additional surveillance programs are needed. To meet this need, an active surveillance program was conducted at pig farms throughout the midwestern United States. From June 2009 through December 2011, nasal swab samples were collected monthly from among 540 groups of growing pigs and tested for influenza A virus by real-time reverse transcription PCR. Of 16,170 samples, 746 were positive for influenza A virus; of these, 18.0% were subtype H1N1, 16.0% H1N2, 7.6% H3N2, and 14.5% (H1N1)pdm09. An influenza (H3N2) and (H1N1)pdm09 virus were identified simultaneously in 8 groups. This active influenza A virus surveillance program provided quality data and increased the understanding of the current situation of circulating viruses in the midwestern US pig population.
Influenza A virus has become a major pathogen, causing epidemics of respiratory disease in humans, which not only result in increased deaths but also raise public health organization alarms regarding the need for further understanding and control of this virus (1). Additionally, the ability of the virus to cross species barriers has raised more concern over the probability of reassortment and generation of highly transmissible viruses that might pose a threat to humans (2). Despite evidence of reassortment in other species, swine have been most often labeled as the “mixing vessel” because avian- and mammalian-type receptors for influenza A virus have been found in pig tracheas, making swine a potential source of new viruses through reassortment (3,4). Because these viruses can infect humans, influenza A virus in swine should be monitored for public health reasons (5).
In the United States, influenza A virus has been present in swine for almost a century (6). Within the US pig population, 3 major subtypes of influenza A virus (H1N1, H1N2, H3N2) circulate, causing widespread respiratory disease characterized by dry coughing, sneezing, fever, anorexia, rhinorrhea, and lethargy (7). Swine influenza viruses have been monitored through seroprevalence studies. Such studies from the 1970s through the 1990s revealed that influenza virus subtypes H1N1 and H3N2 circulated in the US pig population (8–11).
At the turn of the 21st century, 2 new viruses were detected in the swine population. These viruses were the result of either double or triple reassortment between human, avian, and swine viruses (11–13). Since 1998, circulating influenza viruses in pigs have been able to change because of mutations and the propensity for influenza A virus of swine with the triple reassortant genotype to frequently reassort and generate new genotypes, therefore increasing the diversity of influenza A virus in swine (14). Virologic and seroprevalence studies have provided valuable information about influenza A virus in swine, but the epidemiology of influenza A virus in swine is not fully understood.
Newly emerged pathogens can be detected through passive or active surveillance. Passive surveillance is driven by laboratory submission of samples after outbreaks of respiratory disease, whereas active surveillance is based on purposely collecting and screening field samples regardless of clinical status. In Asia, active surveillance for influenza conducted through collection of nasal swabs at slaughter plants has reportedly detected uncommon influenza viruses (i.e., subtypes H3N1, H7N2, H9N2) in the local pig population (15–17). In the United States, similar studies, following the same sample collection method, during the early and late 1990s have been reported (10,18). However, a surveillance program that will identify and report newly emerged viruses in a timely manner is still needed (14).
Overall, studies have elucidated epidemiologic features of the virus in swine, such as the constant circulation of influenza A virus in the swine population and sporadic infections with rare subtypes. However, absence of a proactive approach leaves a gap that needs to be filled (19). Therefore, the objectives of this study were to 1) conduct an active surveillance program to better characterize the presence of influenza viruses in the swine population and 2) make live viruses available for genetic characterization, potential vaccine, and diagnostic use. Procedures and protocols used in this study were approved by the University of Minnesota Institutional Animal Care and Use Committee
Farm Selection
Veterinarians who agreed to participate in the study were asked to enroll growing-pig farms (i.e., farms that house pigs 3–30 weeks of age) in the midwestern United States that were representative of modern swine production systems and that were owned by producers interested in participating in the study. Producers were allowed to withdraw from the study at any time.
Sample Collection
From June 2009 through December 2011, participating farms were visited every month for 12–24 consecutive months. At each visit, the investigator would meet with the farm manager/owner to decide which pigs were to be sampled. If pigs were all in 1 age group, samples were collected from that group; if pigs were in >1 age group, samples would be collected from the age group closest to 10 weeks, the group most likely to yield the most influenza A virus–positive pigs, per previous reports (20).
A total of 30 nasal swab samples were collected at each visit, enabling us to be 95% confident of detecting at least 1 positive sample when influenza prevalence was at least 10%. Clinically healthy pigs were restrained by a snare, and a nasal swab (Starswab II, Starplex Scientific Inc., Etobicoke, Ontario, Canada) was inserted 2–3 inches into the back of each nostril while being rotated. Nasal swabs were labeled with a specific code containing the farm identification number, month, 2-letter state abbreviation, and nasal swab sample number.
During the visit, the age of the pigs and respiratory clinical signs (absence or presence of sneezing, coughing, and nasal secretion) among the group members were recorded. Nasal swabs and submission sheets were placed into a Styrofoam container with ice packs and shipped overnight to the laboratory for testing.
Sample Testing
All nasal swab samples were tested at the virology department laboratory of St. Jude Children’s Research Hospital (Memphis, TN, USA). Nasal swab samples were initially screened for influenza A virus by real-time reverse transcription (RRT-PCR) selective for the matrix gene. Samples that were positive by RRT-PCR underwent further diagnostics for determination of subtype, including the influenza A(H1N1)pdm09 virus and swine H1 and H3 viruses (21–23).
Statistical Analyses
A farm was considered positive for a given month if any of the 30 individually collected swab samples tested positive. Farm-level data, such as farm size, were analyzed by repeated measures logistic regression, and differences between farms were accounted for by including farm as a random effect and allowing for an autoregressive effect by month within the farm. Model building was performed by first screening independent variables through univariate analysis. Variables with a p value <0.25 were retained for the multivariable model. All selected variables were forced in the model including 2-way interactions and were sequentially removed if p value was >0.05.
We built 2 models. The first model assessed the relationships between farm status (positive vs. negative) for influenza virus and age, year, clinical signs, and season. The second model assessed the relationship between respiratory clinical signs (presence vs. absence) and subtype and season. Season was included in the models by categorizing the 4 seasons as follows: winter (January–March), spring (April–June), summer (July–September), and fall (October–December). Statistical procedures were performed in SAS 9.2 (SAS Institute Inc., Cary, NC, USA).
Farms were enrolled in the program as agreement to participate (by veterinarians and producers) was obtained; thus, the program started in June 2009 and ended in December 2011. The 33 producers who agreed to participate were located throughout the midwestern United States: 17 farms in Iowa, 4 in Illinois, 8 in Indiana, and 4 in Minnesota. Each of the 33 farms housed 1,000–13,000 pigs. One group of 7 farms in Iowa withdrew from the program in August 2009 after the influenza A(H1N1)pdm09 virus–associated crisis in the swine industry.
Sample Test Results
A total of 16,170 nasal swab samples from 540 groups of growing pigs from the remaining 32 farms were collected. From the total number of samples collected, 746 (4.6%) were positive for influenza A virus by RRT-PCR, and 178 viruses were isolated from these samples. At least 1 positive sample was detected in 117 (21.7%) groups of pigs; thus, these groups were classified as positive. Of the 32 farms, 29 (90.6%) had at least 1 positive group throughout the study. Of the 117 groups with RRT-PCR–positive results for influenza A virus, complete or partial subtype details were obtained for 99 (84.6%) pig groups (Figure 1). Influenza A virus infection with just 1 subtype (H1N1, H1N2, H3N2, or H1N1pdm09) was detected in 21, 19, 9, and 17 groups, respectively. Dual infections were detected in 10 groups, of which 8 concurrently harbored influenza virus subtypes H1N2 and H1N1pdm09, 1 harbored subtype H1N1 and an H3N-untypeable virus, and the remaining group harbored subtype H1N1 and an H1N-untypeable virus. Partial subtyping information was obtained for 16 pig groups, from which 11, 4 and 1 were infected with an H1N-untypeable, H3N-untypeable, and an H1N-untypeable with pandemic matrix gene virus subtype, respectively. In 18 groups, a subtype could not be defined through either RRT-PCR or sequencing. At most farms in our study, groups of pigs were identified as having multiple and different influenza A viruses detected throughout the surveillance period (Figure 1). Viruses were isolated from 178 swab samples that originated from 62 pig groups.
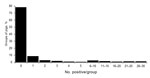
Of the positive groups, the mean and median numbers of samples positive for influenza A virus by RRT-PCR were 6.4 and 2, respectively. The numbers of positive samples ranged from 1 to 30; most groups (n = 48) had 1 positive sample (Figure 2). There were 15, 10, 4, and 2 groups that had 2, 3, 4, and 5 positive samples, respectively. A total of 13 groups had >20 positive samples, 2 had 29 positive samples, and 1 had 30 positive samples. Although most samples collected from these 13 groups were positive for influenza A virus by RRT-PCR, pigs in only 7 of these groups exhibited clinical signs of influenza-like illness.
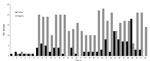
The number of positive groups per farm ranged from 1 to 18. On average, 31% of the groups tested by farm throughout the program were classified as positive (Figure 3). Farms with no influenza A virus–positive results were monitored for ≈1 year but lacked consistency in the testing frequency and time intervals between tests.
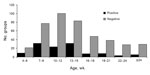
The average age of the pigs in the 540 groups was 13.7 ± 5.7 weeks. Influenza virus was detected in pigs as young as 4 weeks and as old as 30–32 weeks of age. Mean age in positive groups was 12.4 ± 5.2 weeks and in negative groups was 13.9 ± 5.8 SD (Figure 4). However, age was not a statistically significant predictor of influenza A virus test status.
Clinical Signs
Respiratory clinical signs were observed in pigs in 187 (34.6%) of 540 groups. From these 187 groups that reportedly exhibited clinical signs, 43 (22.9%) were positive for influenza A virus. From the 353 groups that exhibited no clinical signs, 74 (20.9%) were positive for influenza virus (Table 1). Even when within-group prevalence of influenza A virus was high, such as in the 13 groups in which >20 samples were positive for influenza A virus, clinical signs of respiratory disease were low. Indeed, clinical signs of influenza-like illness were noted in only 7 of those 13 groups.
Season
Throughout the study, the numbers (proportions) of groups sampled in each season were similar. In winter, 124 (23%) were sampled; in spring, 137 (25%); in summer, 150 (28%); and in fall, 129 (24%) (Table 2).
Logistic Regression
In the first model, univariate analysis of all variables (age, season, and year) except clinical signs yielded p<0.25. The multivariable model retained 2 variables: season and age. Odds of positive results for influenza A virus were 2 (95% CI 1.1–3.8) times and 1.9 (95% CI 1.0–3.5) times higher for groups of pigs tested during the spring and summer, respectively, than for groups tested during the fall. There was no association between the winter season and influenza virus detection. Age was not significantly (p = 0.09) associated with influenza A virus group status; however, the variable was left in the model because of confounding (Table 3).
In the second model, complete subtype information was available for 81 monthly test results. Because neither subtype nor season was significantly associated with presence of clinical signs at the univariate level (Table 4), no attempts were made to build a multivariate model.
Active surveillance at 32 farms demonstrated that influenza A virus is commonly present in the nasal secretions of pigs; 29 (90.6%) farms had at least 1 positive group throughout the study. Despite this high group or population prevalence, detection of swine influenza A virus in individual samples was low and thereby compatible with previously published findings (10,15–18) in which swine influenza A virus detection rates through either virus isolation or RRT-PCR on individual nasal swab samples was ≤5%, presenting a challenge for surveillance programs. However, new sample collection techniques in swine, such as the collection of pen-based oral fluids (e.g., saliva) for antibody and antigen detection are becoming more commonly used because of their practicality and lower testing costs (24); other studies have shown an increased probability of detection, making oral fluid sampling a suitable and essential tool for population surveillance on pig farms (25,26). Asymptomatic carriers of influenza A virus can be detected more efficiently through oral fluid sampling of clinically healthy populations than through nasal swab sampling.
Influenza A virus persistence in pig populations is not fully understood. In our study, 41% of the positive groups had only 1 positive nasal swab sample. One possible explanation could be that these groups of pigs were sampled either at the beginning or end of an infection. Another possible explanation could be that underlying passive or active immunity enabled transmission to occur at a rather low rate and that transmission remained continuous in and thus perpetuated the infection within the population. However, our study was not designed to measure transmission. In addition, our study did not obtain information regarding the entrance into or exit from the farm by groups of pigs. Therefore, it is possible that the same group of pigs was sampled on consecutive months.
In our study, the absence of respiratory clinical signs in groups of pigs harboring influenza A virus is notable. This lack of signs could be the result of events such as the presence of antibodies conferred by colostrum ingestion (e.g., maternal immunity), vaccination, or previous exposure; other studies have suggested that low levels of exposure to virus might preclude clinical signs in pigs (27–29). Subclinical infections in pigs have public health implications because humans can become infected after coming in contact with apparently healthy pigs that are shedding enough infectious viral particles (5,30–32). Subclinical infections are perhaps one of the most common routes for influenza A virus entry into a pig farm because replacement breeding stock or recently weaned animals are constantly moved within and between states and countries. In fact, there is evidence that movement of pigs might have been the cause of dissemination of certain virus lineages within the United States, and subclinical infections might have played a role (33).
Pathogenicity of swine influenza A virus can vary among strains within the same subtype (34). From a clinical signs standpoint, experimental infection with influenza A virus has resulted in great variability (29). When swine influenza A virus is part of a co-infection (i.e., with other viruses and/or bacteria), clinical signs are evident (35,36); such co-infections might reflect the health status situation of the groups of pigs used in this study because the likelihood of co-infections in the field was high. However, in our study, the lack of association between virus subtype and clinical signs is difficult to explain. More studies are needed to better understand the role that virus subtypes play on the presence of respiratory signs.
Our study demonstrated that influenza A virus is present in growing pigs throughout the year and that groups of pigs are more likely to have positive test results during the spring and summer than in the fall. This finding is contrary to what has been suggested (7). The previously suggested seasonal trend could have been based on presence of clinical signs that appear during a time of the year when other factors are present (e.g., cold weather, bad air quality inside barns, co-infections) (7,37), which led veterinarians to submit samples to diagnostic laboratories for the detection of influenza A virus. However, as mentioned earlier, subclinical infections might have occurred during warm months, thereby leading to misinterpretation of the information available at that time. Another possible explanation for finding more influenza A virus–positive groups in the spring and summer is the increase of pigs born to primiparous females. This increase in potentially more susceptible growing pigs is a result of increased breeding of gilts (female pigs that have not had their first litter) during late summer, when swine producers often increase the number of gilts bred. These primiparous females often have a lower level of antibody protection to offer to their first litter of piglets. These relatively immunologically naive piglets would often be the group studied in the spring and summer seasons (38), providing a source of susceptible individuals in which viruses circulate during these seasons.
A limitation of our study might be the locations of the farms, which were all in the midwestern United States. Weather conditions and seasons in the midwestern United States might not accurately reflect conditions in other swine-producing areas of the United States, namely, the southeastern and south-central regions. For logistical reasons, neither of these regions was included in this study.
Surveillance will continue to be a useful tool for infectious disease epidemiology because the data it provides will aid in the understanding of the determinants of infection, enabling scientists and practitioners to work toward generation of disease prevention and control strategies (39). Surveillance studies contribute to science by generating data about zoonotic and emerging pathogens (40). Such contributions are true for influenza A virus in swine because emerging influenza viruses have been identified through surveillance programs (15,16). In summary, our study has led to the following 3 conclusions: 1) different influenza viruses circulate simultaneously within pig populations; 2) influenza is present in pigs of different ages at a rather low prevalence throughout the year; and 3) subclinical infections are frequent among groups of pigs. More studies are needed to add to understanding of influenza A virus.
Dr Corzo was a PhD student at the University of Minnesota College of Veterinary Medicine, Saint Paul, Minnesota, during the study period . His research interests include the epidemiology of swine influenza viruses.
Acknowledgments
We thank the participating swine producers and Aaron Roakey, Keith Kinsley, Max Rodibaugh, Jeff Harker, Cameron Schmitt, Todd Distad, Joe Rudolphi and Tom Gillespie for their willingness to participate in this project. We also thank Michael Kuhn, Nicole Grate-Hurt, Jess Waddell, Chase Stahl, Micah Taylor, Britni Miller, Samantha Shields, Dayle Dillon, Lauren Hughes, Dave Davies, Jamie Coss, and Lance Barton for their help with logistics and sample collection. We acknowledge Jeri Carol Crumpton, Daniel Darnell, Adam Rubrum, Jerry Parker, Rich Elia, Pamela McKenzie, Scott Krauss for their excellent technical assistance and their help with data management.
This study was supported by the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services (contract no. HHSN266200700005C) and by the American Lebanese Syrian Associated Charities.
References
- Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y. Evolution and ecology of influenza A viruses. Microbiol Rev. 1992;56:152–79 .PubMedGoogle Scholar
- Ito T, Couceiro JN, Kelm S, Baum LG, Krauss S, Castrucci MR, Molecular basis for the generation in pigs of influenza A viruses with pandemic potential. J Virol. 1998;72:7367–73 .PubMedGoogle Scholar
- Ma W, Lager KM, Vincent AL, Janke BH, Gramer MR, Richt JA. The role of swine in the generation of novel influenza viruses. Zoonoses Public Health. 2009;56:326–37 .PubMedGoogle Scholar
- Myers KP, Olsen CW, Gray GC. Cases of swine influenza in humans: a review of the literature. Clin Infect Dis. 2007;44:1084–8 .PubMedGoogle Scholar
- Koen JS. A practical method for the field diagnosis of swine diseases. Am J Vet Med. 1919;14:468–70.
- Brown IH. The epidemiology and evolution of influenza viruses in pigs. Vet Microbiol. 2000;74:29–46 .PubMedGoogle Scholar
- Hinshaw VS, Bean WJ Jr, Webster RG, Easterday BC. The prevalence of influenza viruses in swine and the antigenic and genetic relatedness of influenza viruses from man and swine. Virology. 1978;84:51–62 .PubMedGoogle Scholar
- Chambers TM, Hinshaw VS, Kawaoka Y, Easterday BC, Webster RG. Influenza viral infection of swine in the United States 1988–1989. Arch Virol. 1991;116:261–5 .PubMedGoogle Scholar
- Olsen CW, Carey S, Hinshaw L, Karasin AI. Virologic and serologic surveillance for human, swine and avian influenza virus infections among pigs in the north-central United States. Arch Virol. 2000;145:1399–419 .PubMedGoogle Scholar
- Zhou NN, Senne DA, Landgraf JS, Swenson SL, Erickson G, Rossow K, Genetic reassortment of avian, swine, and human influenza A viruses in American pigs. J Virol. 1999;73:8851–6 .PubMedGoogle Scholar
- Webby RJ, Swenson SL, Krauss SL, Gerrish PJ, Goyal SM, Webster RG. Evolution of swine H3N2 influenza viruses in the United States. J Virol. 2000;74:8243–51 .PubMedGoogle Scholar
- Karasin AI, Olsen CW, Anderson GA. Genetic characterization of an H1N2 influenza virus isolated from a pig in Indiana. J Clin Microbiol. 2000;38:2453–6 .PubMedGoogle Scholar
- Webby RJ, Rossow K, Erickson G, Sims Y, Webster R. Multiple lineages of antigenically and genetically diverse influenza A virus co-circulate in the United States swine population. Virus Res. 2004;103:67–73 .PubMedGoogle Scholar
- Peiris JS, Guan Y, Markwell D, Ghose P, Webster RG, Shortridge KF. Cocirculation of avian H9N2 and contemporary “human” H3N2 influenza A viruses in pigs in southeastern China: potential for genetic reassortment? J Virol. 2001;75:9679–86 .PubMedGoogle Scholar
- Shieh HK, Chang PC, Chen TH, Li KP, Chan CH. Surveillance of avian and swine influenza in the swine population in Taiwan, 2004. J Microbiol Immunol Infect. 2008;41:231–42 .PubMedGoogle Scholar
- Kwon TY, Lee SS, Kim CY, Shin JY, Sunwoo SY, Lyoo YS. Genetic characterization of H7N2 influenza virus isolated from pigs. Vet Microbiol. 2011;153:393–7 .PubMedGoogle Scholar
- Wright SM, Kawaoka Y, Sharp GB, Senne DA, Webster RG. Interspecies transmission and reassortment of influenza A viruses in pigs and turkeys in the United States. Am J Epidemiol. 1992;136:488–97 .PubMedGoogle Scholar
- Beaudoin A, Johnson S, Davies P, Bender J, Gramer M. Characterization of influenza A outbreaks in Minnesota swine herds and measures taken to reduce the risk of zoonotic transmission. Zoonoses Public Health. 2012;59:96–106 .PubMedGoogle Scholar
- Richt JA, Lager KM, Clouser DF, Spackman E, Suarez DL, Yoon KJ. Real-time reverse transcription-polymerase chain reaction assays for the detection and differentiation of North American swine influenza viruses. J Vet Diagn Invest. 2004;16:367–73 .PubMedGoogle Scholar
- World Health Organization. CDC protocol of realtime RTPCR for influenza A(H1N1) [cited 2013 April 10]. http://www.who.int/csr/resources/publications/swineflu/realtimeptpcr/en/
- Ducatez MF, Hause B, Stigger-Rosser E, Darnell D, Corzo C, Juleen K, Multiple reassortment between pandemic (H1N1) 2009 and endemic influenza viruses in pigs, United States. Emerg Infect Dis. 2011;17:1624–9 .PubMedGoogle Scholar
- Prickett JR, Zimmerman JJ. The development of oral fluid–based diagnostics and applications in veterinary medicine. Anim Health Res Rev. 2010;11:207–16 .PubMedGoogle Scholar
- Romagosa A, Gramer M, Joo HS, Torremorell M. Sensitivity of oral fluids for detecting influenza A virus in populations of vaccinated and non-vaccinated pigs. Influenza Other Respi Viruses. 2012;6:110–8 .PubMedGoogle Scholar
- Ramirez A, Wang C, Prickett JR, Pogranichniy R, Yoon KJ, Main R, Efficient surveillance of pig populations using oral fluids. Prev Vet Med. 2012;104:292–300 .PubMedGoogle Scholar
- Van Reeth K, Labarque G, Nauwynck H, Pensaert M. Differential production of proinflammatory cytokines in the pig lung during different respiratory virus infections: correlations with pathogenicity. Res Vet Sci. 1999;67:47–52 .PubMedGoogle Scholar
- Van Reeth K. Cytokines in the pathogenesis of influenza. Vet Microbiol. 2000;74:109–16 .PubMedGoogle Scholar
- Van Reeth K. Avian and swine influenza viruses: our current understanding of the zoonotic risk. Vet Res. 2007;38:243–60 .PubMedGoogle Scholar
- Gray GC, Trampel DW, Roth JA. Pandemic influenza planning: shouldn't swine and poultry workers be included? Vaccine. 2007;25:4376–81 .PubMedGoogle Scholar
- Killian ML, Swenson SL, Vincent AL, Landgraf JG, Shu B, Lindstrom S, Simultaneous infection of pigs and people with triple-reassortant swine influenza virus H1N1 at a U.S. county fair. Zoonoses Public Health. 2012 Jul 9. [epub ahead of print].
- Newman AP, Reisdorf E, Beinemann J, Uyeki TM, Balish A, Shu B, Human case of swine influenza A (H1N1) triple reassortant virus infection, Wisconsin. Emerg Infect Dis. 2008;14:1470–2 .PubMedGoogle Scholar
- Nelson MI, Lemey P, Tan Y, Vincent A, Lam TT, Detmer S, Spatial dynamics of human-origin H1 influenza A virus in North American swine. PLoS Pathog. 2011;7:e1002077 .PubMedGoogle Scholar
- Vincent AL, Ma W, Lager KM, Janke BH, Richt JA. Swine influenza viruses: a North American perspective. Adv Virus Res. 2008;72:127–54 .PubMedGoogle Scholar
- Van Reeth K, Nauwynck H, Pensaert M. Dual infections of feeder pigs with porcine reproductive and respiratory syndrome virus followed by porcine respiratory coronavirus or swine influenza virus: a clinical and virological study. Vet Microbiol. 1996;48:325–35 .PubMedGoogle Scholar
- Thacker EL, Thacker BJ, Janke BH. Interaction between Mycoplasma hyopneumoniae and swine influenza virus. J Clin Microbiol. 2001;39:2525–30 .PubMedGoogle Scholar
- Straub OC. The important viral infections of pigs. Swine Health and Production. 1994;2:15–8.
- Leman AD. Optimizing farrowing rate and litter size and minimizing nonproductive sow days. Vet Clin North Am Food Anim Pract. 1992;8:609–21 .PubMedGoogle Scholar
- Salman MD, editor. Animal disease surveillance and survey systems. Methods and applications. 1st ed. Ames (IA): Iowa State Press; 2003.
- Kuiken T, Leighton FA, Fouchier RA, LeDuc JW, Peiris JS, Schudel A, Public health. Pathogen surveillance in animals. Science. 2005;309:1680–1 .PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 19, Number 6—June 2013
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Marie Culhane, University of Minnesota, Veterinary Diagnostic Laboratory, 1333 Gortner Ave, St. Paul, MN, USA 55108
Top