Volume 20, Number 4—April 2014
CME ACTIVITY - Research
Travel-associated Antimicrobial Drug–Resistant Nontyphoidal Salmonellae, 2004–2009
Cite This Article
Citation for Media
Introduction

Medscape, LLC is pleased to provide online continuing medical education (CME) for this journal article, allowing clinicians the opportunity to earn CME credit.
This activity has been planned and implemented in accordance with the Essential Areas and policies of the Accreditation Council for Continuing Medical Education through the joint sponsorship of Medscape, LLC and Emerging Infectious Diseases. Medscape, LLC is accredited by the ACCME to provide continuing medical education for physicians.
Medscape, LLC designates this Journal-based CME activity for a maximum of 1 AMA PRA Category 1 Credit(s)TM. Physicians should claim only the credit commensurate with the extent of their participation in the activity.
All other clinicians completing this activity will be issued a certificate of participation. To participate in this journal CME activity: (1) review the learning objectives and author disclosures; (2) study the education content; (3) take the post-test with a 70% minimum passing score and complete the evaluation at www.medscape.org/journal/eid; (4) view/print certificate.
Release date: March 18, 2014; Expiration date: March 18, 2015
Learning Objectives
Upon completion of this activity, participants will be able to:
• Describe the epidemiology and clinical characteristics of nontyphoidal Salmonella (NTS) infections
• Distinguish antibiotic resistance associated with NTS cases in Oregon between 2004 and 2009
• Identify factors positively and negatively associated with antibiotic resistance among NTS cases in Oregon between 2004 and 2009.
CME Editor
P. Lynne Stockton-Taylor, DVM, MA, Technical Writer/Editor, Emerging Infectious Diseases. Disclosure: Carol E. Snarey, MA, has disclosed no relevant financial relationships.
CME Author
Laurie Barclay, MD, freelance writer and reviewer, Medscape, LLC. Disclosure: Laurie Barclay, MD, has disclosed no relevant financial relationships.
Authors
Disclosures: Russell S. Barlow, MPH; Emilio E. DeBess, DVM, MPVM; Jodi A. Lapidus, PhD; Robert Vega, MS(SM,AAM); and Paul R. Cieslak, MD, have disclosed no relevant financial relationships. Kevin L. Winthrop, MD, MPH, has disclosed the following relevant financial relationships: served as an advisor or consultant for Pfizer, UCB, Genentech, AbbVie, Regeneron; received grants for clinical research from Pfizer.
Abstract
To evaluate trends in and risk factors for acquisition of antimicrobial-drug resistant nontyphoidal Salmonella infections, we searched Oregon surveillance data for 2004–2009 for all culture-confirmed cases of salmonellosis. We defined clinically important resistance (CIR) as decreased susceptibility to ampicillin, ceftriaxone, ciprofloxacin, gentamicin, or trimethoprim/sulfamethoxazole. Of 2,153 cases, 2,127 (99%) nontyphoidal Salmonella isolates were obtained from a specific source (e.g., feces, urine, blood, or other normally sterile tissue) and had been tested for drug susceptibility. Among these, 347 (16%) isolates had CIR. The odds of acquiring CIR infection significantly increased each year. Hospitalization was more likely for patients with than without CIR infections. Among patients with isolates that had been tested, we analyzed data from 1,813 (84%) who were interviewed. Travel to eastern or Southeast Asia was associated with increased CIR. Isolates associated with outbreaks were less likely to have CIR. Future surveillance activities should evaluate resistance with respect to international travel.
Each year, nontyphoidal salmonellae (NTS) are responsible for >1 million infections in the United States and an estimated 98 million cases globally (1–4). Each year in the United States, infections result in an estimated 168,000 physician visits, 19,000 hospitalizations, and 380 deaths at a cost of $US 2.3 billion (1–3,5). Data suggest that 85.6% of NTS infections are foodborne and that the remaining infections occur by the fecal–oral route in human-to-human transmission and zoonotic transmission (2). For healthy persons, infections commonly result in self-limiting acute gastroenteritis that resolves without antimicrobial drug therapy. However, antimicrobial drugs can be life saving for immunologically vulnerable populations, such as infants, elderly persons, immunocompromised persons, and persons with invasive infection (6–8). The drugs most commonly prescribed in developing countries are ampicillin and chloramphenicol; those most commonly prescribed in the United States are trimethoprim/sulfamethoxazole, fluoroquinolones, and cephalosporins (9).
In the 1980s, studies demonstrated alarming increases in the prevalence of antimicrobial drug resistance among NTS infections (10,11). This increase was associated with indiscriminate use of antimicrobial drugs in animal husbandry and in humans (10–12). A retrospective study conducted during 1996–2001 associated antimicrobial drug resistance with increased disease severity, highlighting the risk to public health (13).
During the past decade, few population-level analyses have identified risk factors for acquiring a resistant NTS infection outside of outbreak clusters and retail meat supplies. A recently identified risk factor is international travel (14,15). Bacteriologic studies from Europe identified differences in resistance among Salmonella enterica serotypes Stanley, Concord, and Typhimurium isolated from patients with a history of international travel (16–20). However, these studies did not estimate the magnitude of or risk for antimicrobial drug–resistant NTS acquisition among international travelers. We hypothesized that international travel is a risk factor for acquisition of a resistant NTS infection. To test our hypothesis, we analyzed surveillance data from Oregon for 2004–2009 and quantified trends in antimicrobial drug resistance, investigated the relationship between resistance and case outcomes, and assessed whether international travel was associated with acquisition of NTS infections with clinically important resistance (CIR).
Reportable Infectious Disease Surveillance in Oregon
The Oregon Health Authority conducts active, laboratory-based surveillance for all cases of NTS infection. Physicians and laboratories are required by law to report laboratory-confirmed and clinically suspected cases of salmonellosis to the patient’s local health department; reports should contain the patient’s date of birth, sex, diagnosis, date of symptom onset, date of specimen collection, and laboratory test results. All Salmonella isolates are forwarded to the Oregon State Public Health Laboratory (OSPHL), where they are serotyped. Local health department officials interview patients about hospitalization, clinical outcomes, additional demographic information, and exposure history for the 7 days before illness onset. Risk-factor questions ask about specific travel, human, animal, and high-risk food exposures. International travel was considered a risk factor only if it had taken place in the 30 days before illness onset. Patients with recurrent infection or multiple Salmonella isolates (of same serotype within a plausible time frame for the original infection) are interviewed only once, at the time of initial illness onset.
During 2004–2009, the population of Oregon was 3.6–3.8 million persons, which is ≈1.2% of the US population (21,22). The surveillance system in Oregon is estimated to capture >99% of laboratory-confirmed cases of salmonellosis; however, for every 1 case confirmed, an estimated 25 additional cases are not detected (2).
Antimicrobial Drug Susceptibility Testing
For 2004 and 2005, all confirmed isolates were forwarded to the Oregon State University Veterinary Diagnostic Laboratory for susceptibility testing. From 2006 through 2009, susceptibility testing was performed by OSPHL. All isolates were tested by using broth microdilution to determine MICs for the following 10 antimicrobial agents: ampicillin, ceftriaxone, chloramphenicol, ciprofloxacin, gentamicin, nalidixic acid, nitrofurantoin, sulfamethoxazole, tetracycline, and trimethoprim/sulfamethoxazole. Susceptibilities were determined according to Clinical and Laboratory Standards Institute interpretative criteria (23). To ascertain cephalosporin resistance, OSPHL tested isolates for ceftriaxone susceptibility; and the Oregon State University Veterinary Diagnostic Laboratory tested for susceptibility to cefuroxime and cephalothin by using analogous broth microdilution methods. MIC results were dichotomized as susceptible or resistant.
Analyses
Isolates were included in analyses only if they were cultured from specific specimens, such as feces, urine, or blood or other normally sterile tissues (i.e., cerebrospinal fluid). CIR was defined as resistance to at least 1 of the following: ampicillin, ceftriaxone, ciprofloxacin, gentamicin, or trimethoprim/sulfamethoxazole (13). We used the Cochran-Armitage test for trend to analyze NTS case data for 2004–2009 to determine whether the proportion of Salmonella isolates with CIR increased significantly. Demographic and exposure risk factors, specifically international travel, were evaluated as risk factors for acquisition of a resistant isolate. The 9 most common Salmonella serotypes were fixed (included in all models regardless whether they met the p<0.05 level of significance) in all analyses, and remaining serotypes were grouped as “all other.” Serotype Enterditis is the most frequently isolated serotype in Oregon and was therefore used as the referent for all comparisons.
We sought to evaluate whether resistance was associated with increased disease severity, including hospitalization and invasive infection. Invasive infection was defined as isolation of Salmonella from a normally sterile body site or tissue, such as blood (13). Multiple logistic regression models were constructed with variables that were significant at the p<0.25 level in unadjusted analyses. Salmonella serotype and patient race, age, and year of illness onset were fixed in all models. Other variables were given further consideration according to disease severity or relevance for external validity. Predictor variables significant at p<0.05 were retained in the final model, and adjusted log odds ratios (aORs) were calculated. Model fit was assessed by using the Hosmer–Lemeshow goodness-of-fit test.
All analyses were performed by using SAS version 9.2 (SAS Institute Inc., Cary, NC, USA). Because this study involved more extensive analysis only of data collected routinely as part of public health surveillance, it was not considered human subjects research.
Descriptive Epidemiology and Resistance Trends
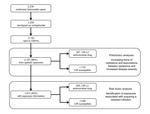
From 2004 through 2009, a total of 2,255 laboratory-confirmed cases of nontyphoidal salmonellosis were reported in Oregon. In accordance with Oregon law, 2,153 isolates were forwarded to OSPHL, and 2,127 (98.8% of all NTS isolates) were cultured from a specific source and had antimicrobial drug susceptibility testing information (Figure). Of these isolates, 26 (1.2%) were obtained through a nonspecific source, such as lesions or sputum, and were excluded from analysis.
The most common Salmonella serotypes detected were Enteritidis (18.4%), Typhimurium (14.3%), Heidelberg (8.2%), Typhimurium var. Copenhagen (5.1%), and Newport (4.5%). Cases that were part of identified outbreaks represented 24.1% of the cohort; the remaining 75.9% were considered sporadic cases. The median age of patients was 29 years (interquartile range 9–51 years), and 53.1% of patients were female. Information about race was not available for 9%; of patients for whom race was known, 91.8% were white and 9.2% were not white. Similarly, information about ethnicity was not available for 10.5% of patients. Among patients for whom ethnicity was known, 87.4% were not Hispanic and 12.6% were Hispanic. Isolates from 1,213 (57%) patients were susceptible to all antimicrobial drugs screened (pansusceptible), and isolates from 347 (16.3%) had CIR (Table 1). Of the 2,127 patients, 412 (19.4%) were hospitalized and 110 (5.2%) had invasive disease.
The proportion of isolates that were pansusceptible significantly decreased from 69.5% in 2004 to 53.6% in 2009 (p<0.01). CIR did not significantly increase during this study period (p = 0.27). Stratification by serotype revealed that CIR increased among the 3 most common serotypes: Enteritidis (3% to 8%, p = 0.02), Typhimurium (19% to 34%, p = 0.03), and Heidelberg (6% to 30%, p<0.01). Significant increases were identified for resistance to ciprofloxacin (p<0.05), nalidixic acid (p<0.01), sulfamethoxazole (p<0.01), tetracycline (p<0.01), and trimethoprim/sulfamethoxazole (p<0.01). Cephalosporin resistance increased, although not significantly (p = 0.06).
We suspected that these findings were confounded by serotype and therefore used logistic regression to model the odds of acquiring a resistant infection for each of the clinically important antimicrobial drugs (ampicillin, ceftriaxone, ciprofloxacin, gentamicin, or trimethoprim/sulfamethoxazole) as well as CIR. Serotype-adjusted log odds ratios were generated with year of infection entered as a discrete continuous variable. After adjusting for serotype, we found that with each subsequent year, patients were 30% more likely to acquire an infection that was resistant to quinolones (nalidixic acid or ciprofloxacin) and trimethoprim/sulfamethoxazole (Table 2). Resistance to ampicillin and cephalosporin also increased, although not significantly. We also found that with each year, odds of acquiring an infection with CIR increased by 13%.
CIR was associated with hospitalization (odds ratio [OR] 1.5, 95% CI 1.1–2.0). This association was preserved after adjustment for serotype, patient age, patient race, and year (aOR 1.7, 95% CI 1.2–2.1). For patients with CIR infections, odds of invasive infection were increased, although not significantly, according to unadjusted or adjusted analyses (OR 1.4, 95% CI 0.9–2.2 and aOR 1.5, 95% CI 0.9–2.5, respectively).
Risk Factors
Of the 2,127 patients included in the previous analyses, 1,813 (84.2% of all Oregon patients with NTS) were interviewed. For 305 (16.8%) of these patients, isolates had CIR, and for the remaining 1,508 (83.2%), isolates were susceptible to all clinically important antimicrobial drugs (Figure). Of the 1,508 isolates susceptible to clinically important drugs, 1,002 (55.3%) were susceptible to all drugs screened and 506 (27.9%) were resistant to at least 1 non-CIR drug.
According to the unadjusted analysis, several serotypes were more likely than the referent serotype, Enteritidis, to be resistant to >1 clinically important antimicrobial drug (Table 3). Patient sex, race, age, and ethnicity were not significantly associated with resistance.
CIR was not associated with any other demographic risk factors or high-risk food or animal exposures. However when international travel was examined by individual countries or applicable United Nations region, CIR was significantly associated with travel to Southeast Asia (24). Associations of resistance with travel to Mexico and eastern Asia also approached significance (Table 4). The most common travel destinations in Asia where resistant infections were acquired were Thailand, China, and Malaysia/Indonesia. On the basis of these findings, we analyzed international travel by 3 destinations: Central America, including Mexico (135 patients), eastern and Southeast Asia (46 patients), and all other international travel destinations (77 patients).
Patients who were part of identified outbreak clusters were significantly less likely than patients with sporadic infections to have a resistant infection (OR 0.5, 95% CI 0.4–0.8). During our study period, 131 outbreaks (406 cases) occurred, and for 25 of these outbreaks a causative vehicle was successfully identified. To assess whether oversampling of cases from outbreak clusters could explain this association, we first restricted cases to 1 isolate per outbreak where a causative vehicle was implicated while retaining all cases from outbreaks for which a vehicle was not implicated (302 cases). Second, we further restricted cases to 1 isolate per outbreak, regardless whether a vehicle was identified (131 cases). In each of these analyses the magnitude, direction, and significance of the association was preserved (OR 0.6, 95% CI 0.4–0.8 and OR 0.5, 95% CI 0.3–0.9, respectively), suggesting that oversampling could not have explained this association. Furthermore, 53.6% of outbreaks had intra-outbreak cases for which the antimicrobial drug susceptibility profiles of the isolates differed.
The resultant main-effects model included the fixed variables of serotype, patient age, year of onset, and patient race, along with travel to eastern or Southeast Asia, and outbreak association (Table 3). We hypothesized that effect modification occurred between the variables of outbreak cases and travel to eastern or Southeast Asia, as well as between travel to eastern or Southeast Asia, serotypes, and outbreaks. However, no significant interactions at the p<0.25 level were identified. No other variables or exposures were significantly associated with CIR. This model had excellent goodness-of-fit (p = 0.87). The association between resistance and travel to eastern or Southeast Asia was preserved after exclusion of all outbreak-associated cases (aOR 4.6, 95% CI 2.3–9.4; Table 5). Similarly, the association between resistance and outbreak-associated cases was preserved after exclusion of patients with a history of international travel (aOR 0.5, 95% CI 0.4–0.8; Table 6). Therefore, these results suggest that inclusion of patients with a history of travel to Asia, as well as patients with outbreak-associated infections for the main analysis, was appropriate.
Patients with a history of recent travel to eastern or Southeast Asia were >5 times more likely to acquire a CIR infection than were patients with no history of recent international travel. The most common serotypes acquired among persons with a history of travel to Asia were Enteriditis (n = 13, 54% CIR), Typhimurium (n = 5, 60% CIR), Newport (n = 4, 25% CIR), I 4, 5, 12:i:- (n = 4, 50% CIR), Stanley (n = 3, 33% CIR), and Typhimurium var. Copenhagen (n = 2, 100% CIR). Patients with outbreak-associated infections were half as likely as those with sporadic infections to have CIR (Table 3).
To identify risk factors for resistance to individual antimicrobial drugs, we constructed models with each of the clinically important antimicrobial drugs. Travel to eastern or Southeast Asia was significantly associated with resistance to ampicillin, quinolones (nalidixic acid or ciprofloxacin), and trimethoprim/sulfamethoxazole (Table 7). Only individual serotypes were associated with resistance to cephalosporins or gentamicin, and no other risk factors were significantly associated with resistance to ampicillin, quinolones, or trimethoprim/sulfamethoxazole.
We found that NTS infections were more likely to have CIR with each subsequent year of our study. In Oregon during 2004–2009, the proportion of isolates susceptible to all antimicrobial drugs significantly decreased. Travel to eastern and Southeast Asia was associated with acquisition of Salmonella with CIR. Such travel was specifically associated with resistance to ampicillin, quinolones, and trimethoprim/sulfamethoxazole. Isolates from patients who were part of identified outbreak clusters were significantly less likely to be resistant, suggesting that resistance estimates based on outbreak cases alone may underestimate the true level of resistance. We also report that resistance is associated with increased hospitalization (13,25).
Our analysis was performed by using Salmonella susceptibility data from a surveillance system that captures ≈100% of confirmed infections, has antimicrobial drug susceptibility information for >95% of confirmed cases, and includes exposure histories for >84% of patients. This study is strengthened by having collected data on several known and potential confounders before the drug-susceptibility profiles were known. Our study design complements a previous National Antimicrobial Resistance Monitoring System/FoodNet study that analyzed antimicrobial drug resistance and increased disease severity (13). However, we determined where resistant infections were acquired (exposures) and patient outcomes associated with resistant infections by using an entire population. The ability to integrate resistance and serotype data with case-specific demographic and risk-factor data improves the generalizability and plausibility of our study and provides population-level risk estimates (14–20).
Widespread quinolone resistance in Southeast Asia has been reported (26); a better understanding of global use of antimicrobial drugs might suggest where resistant salmonellae are prevalent. Examination of serotype profiles among patients who had traveled to eastern or Southeast Asia and multivariate analyses adjusted for serotype provided strong evidence that the increased resistance in this region is widespread and not specifically attributable to a single serotype or regional serotype differences.
Increasing antimicrobial drug resistance has widespread implications for human health. We confirm the results of Varma et al. and Lee et al., who found antimicrobial drug resistance to be associated with increased likelihood of hospitalization (13,25). More severe infections can lead to treatment failure, sepsis, meningitis, and even death. If resistance to clinically important antimicrobial drugs continues to increase by 13% per year, as our data suggest, we can expect more severe illnesses, hospitalizations, and deaths, along with the accompanying higher economic costs.
The association between resistance and outbreak cases persisted after restricting the data in unadjusted and adjusted analyses. The lack of effect modification between outbreak cases and a history of travel to Asia in the multiple logistic regression modeling suggests that this finding is independent of travel. Resistant isolates might be less infectious and therefore less likely to cause recognizable outbreaks. Alternatively, common sources of resistant isolates might be less likely to cause widespread contamination.
Our study had limitations. We did not have information about previous antimicrobial drug use (11,27,28). However, this exposure would be expected to confound the observed associations nondifferentially, thereby resulting in lower point estimates. Reporting lags could have delayed risk-factor interviews, resulting in nondifferential recall bias. This bias would not be expected to explain the association between resistance and international travel and would ultimately lead to underestimation of the true effect size. Case ascertainment among persons with a history of travel to eastern or Southeast Asia could have been biased. This bias could have affected our analyses if more severe illness developed in travelers with resistant infections, who were more likely to seek health care or be reported than were travelers without resistant infections. However, according to a subanalysis, not presented here, we found that patients who traveled to eastern or Southeast Asia were less likely to be hospitalized than were those who had not recently traveled internationally (OR 0.4, 95% CI 0.2–1.2). This finding might be explained by a healthy traveler effect; thus, the association between resistance and travel to eastern and Southeast Asia cannot be explained by biased case ascertainment (29). The use of 2 laboratories for susceptibility testing could have resulted in systematic bias. Both laboratories were licensed and were using Clinical and Laboratory Standards Institute standardized methods, suggesting that this bias, if present, would be minimal and could not explain the observed associations.
This study demonstrates that antimicrobial drug resistance among NTS is increasing and has clinical and public health implications. Our analyses elucidated that travel to Asia is strongly associated with antimicrobial drug resistance. When considering antimicrobial drug therapy, providers should evaluate patient travel history and Salmonella serotype. Our results highlight the need for enhanced domestic surveillance for antimicrobial drug resistance and suggest a need for increased prudence regarding the use of antimicrobial drugs.
Mr Barlow is a graduate student in the Departments of Immunology and Global Health, University of Washington, Seattle, Washington, USA. His research interests include host–pathogen interactions and disease surveillance relating to emerging infectious diseases.
Acknowledgments
We thank Julie Hatch for helping with serotype classifications and legacy data abstraction, Steven Fiala for providing statistical advice, and Hillary Booth for helping with retrieval of outbreak case information.
This research was supported in part by the Pacific Northwest Regional Center of Excellence for Biodefense and Emerging Infectious Diseases Research and by Cooperative Agreement no. 3U01CI000306 from the Centers for Disease Control and Prevention.
References
- Mead PS, Slutsker L, Dietz V, McCaig LF, Bresee JS, Shapiro C, Food-related illness and death in the United States. Emerg Infect Dis. 1999;5:607–25. DOIPubMedGoogle Scholar
- Scallan E, Hoekstra RM, Angulo FJ, Tauxe RV, Widdowson MA, Roy SL, Foodborne illness acquired in the United States—major pathogens. Emerg Infect Dis. 2011;17:7–15. DOIPubMedGoogle Scholar
- Voetsch AC, Van Gilder TJ, Angulo FJ, Farley MM, Shallow S, Marcus R, FoodNet estimate of the burden of illness caused by non-typhoidal Salmonella infections in the United States. Clin Infect Dis. 2004;38(suppl 3):S127–34. DOIPubMedGoogle Scholar
- Majowicz SE, Musto J, Scallan EJ, Angulo FJ, Kirk M, O'Brien SJ, The global burden of non-typhoidal Salmonella gastroenteritis. Clin Infect Dis. 2010;50:882–9. DOIPubMedGoogle Scholar
- Frenzen PD, Riggs TL, Buzby JC, Breuer T, Roberts T, Voetsch D, Salmonella cost estimate updated using FoodNet data. Food Review. 1999;22:10–5.
- Gordon MA, Banda HT, Gondwe M, Gordon SB, Boeree MJ, Walsh AL, Non-typhoidal Salmonella bacteraemia among HIV-infected Malawian adults: high mortality and frequent recrudescence. AIDS. 2002;16:1633–41. DOIPubMedGoogle Scholar
- Gordon MA, Graham SM, Walsh AL, Wilson L, Phiri A, Molyneux E, Epidemics of invasive Salmonella enterica serovar Enteritidis and S. enterica serovar Typhimurium infection associated with multidrug resistance among adults and children in Malawi. Clin Infect Dis. 2008;46:963–9. DOIPubMedGoogle Scholar
- Cheesbrough JS, Taxman BC, Green SD, Mewa F, Numbi I. A clinical definition for invasive Salmonella infection in African children. Pediatr Infect Dis J. 1997;16:277–83. DOIPubMedGoogle Scholar
- Guerrant RL, Van Gilder T, Seiner TS, Thielman NM. Slutsker L, Tauxe RV, et al. Practice guidelines for the management of infectious diarrhea. Clin Infect Dis. 2001;32:331–51.
- Holmberg SD, Wells JG, Cohen ML. Animal-to-man transmission of antimicrobial-resistant Salmonella: investigations of U.S. outbreaks, 1971–1983. Science. 1984;225:833–5. DOIPubMedGoogle Scholar
- MacDonald KL, Cohen ML, Hargrett-Bean NT, Wells JG, Puhr ND, Collin SF, Changes in antimicrobial resistance of Salmonella isolated from humans in the United States. JAMA. 1987;258:1496–9. DOIPubMedGoogle Scholar
- Tauxe RV. Antimicrobial resistance in human salmonellosis in the United States. J Anim Sci. 1986;62(Suppl 3):65–73.
- Varma JK, Molbak K, Barrett TJ, Beebe JL, Jones TF, Rabatsky-Ehr T, Antimicrobial-resistant non-typhoidal Salmonella is associated with excess bloodstream infections and hospitalizations. J Infect Dis. 2005;191:554–61. DOIPubMedGoogle Scholar
- Hendriksen RS, Mikoleit M, Kornschober C, Rickert RL, Duyne SV, Kjelsø C, Emergence of multidrug-resistant Salmonella Concord infections in Europe and the United States in children adopted from Ethiopia, 2003–2007. Pediatr Infect Dis J. 2009;28:814–8. DOIPubMedGoogle Scholar
- Hendriksen RS, Le Hello S, Bortolaia V, Pulsrikarn C, Nielsen EM, Pornruangmong S, Characterization of isolates of Salmonella enterica serovar Stanley, a serovar endemic to Asia and associated with travel. J Clin Microbiol. 2012;50:709–20. DOIPubMedGoogle Scholar
- Threlfall EJ, Ward LR, Skinner JA, Graham A. Antimicrobial drug resistance in non-typhoidal Salmonella from humans in England and Wales in 1999: decrease in multiple resistance in non-typhoidal Salmonella enterica serotypes Typhimurium, Virchow and Hadar. Microb Drug Resist. 2000;6:319–25. DOIPubMedGoogle Scholar
- Kariuki S, Revathi G, Kariuki N, Kiiru J, Mwituria J, Muyodi J, Invasive multidrug resistant non-typhoidal Salmonella infections in Africa: zoonotic or anthroponotic transmission? J Med Microbiol. 2006;55:585–91. DOIPubMedGoogle Scholar
- Whichard JM, Gay K, Stevenson JE, Joyce KJ, Cooper KL, Omondi M, Human Salmonella and concurrent decreased susceptibility to quinolones and extended-spectrum cephalosporins. Emerg Infect Dis. 2007;13:1681–8. DOIPubMedGoogle Scholar
- Hendriksen RS, Bangtrakulnonth A, Pulsrikarn C, Pornreongwong S, Hasman H, Song SW, Antimicrobial resistance and molecular epidemiology of Salmonella Rissen from animals, food products, and patients in Thailand and Denmark. Foodborne Pathog Dis. 2008;5:605–19. DOIPubMedGoogle Scholar
- Torpdahl M, Lauderdale TL, Liang SY, Li I, Wei SH, Chiou CS. Human isolates of Salmonella enterica serovar Typhimurium from Taiwan displayed significantly higher levels of antimicrobial resistance than those from Denmark. Int J Food Microbiol. 2013;161:69–75 and. DOIPubMedGoogle Scholar
- Portland State University. Population estimates [cited 2011 Oct 9]. http://pdx.edu/prc/population-estimates-0
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. Sixteenth informational supplement (M100-s16). Wayne (PA): The Institute:2006.
- United Nations. Composition of macro geographical (continental) regions, geographical sub-regions, and selected economic and other groupings [cited 2013 Oct 11]. http://millenniumindicators.un.org/unsd/methods/m49/m49regin.htm#americas
- Lee LA, Puhr N, Maloney E, Bean N, Tauxe R. Increase in antimicrobial-resistant Salmonella infections in the United States, 1989–1990. J Infect Dis. 1994;170:128–34. DOIPubMedGoogle Scholar
- Lee HY, Su LH, Tsai MH, Kim SW, Chang HH. Jung SI, et al. High rate of reduced susceptibility to ciprofloxacin and ceftriaxone among nontyphoid Salmonella clinical isolates in Asia. Antimicrob Agents Chemother. 2009;53:2696–9.
- Winokur PL, Canton R, Casellas JM, Legakis N. Variations in the prevalence of strains expressing an extended-spectrum beta-lactamase phenotype and characterization of isolates from Europe, the Americas, and the Western Pacific region. Clin Infect Dis. 2001;32(Suppl 2):S94–103 . DOIPubMedGoogle Scholar
- Brent AJ, Oundo JO, Mwangi I, Ochola L, Lowe B, Berkley JA. Salmonella bacteremia in Kenyan children. Pediatr Infect Dis J. 2006;25:230–6. DOIPubMedGoogle Scholar
- Koch K, Kristensen B, Holt HM, Ethelberg S, Mølbak K, Schønheyder HC. International travel and the risk of hospitalization with non-typhoidal Salmonella bacteremia. A Danish population-based cohort study, 1999–2008. BMC Infect Dis. 2011;11:277. DOIPubMedGoogle Scholar
Figure
Tables
Follow Up
Earning CME Credit
To obtain credit, you should first read the journal article. After reading the article, you should be able to answer the following, related, multiple-choice questions. To complete the questions (with a minimum 70% passing score) and earn continuing medical education (CME) credit, please go to www.medscape.org/journal/eid. Credit cannot be obtained for tests completed on paper, although you may use the worksheet below to keep a record of your answers. You must be a registered user on Medscape.org. If you are not registered on Medscape.org, please click on the New Users: Free Registration link on the left hand side of the website to register. Only one answer is correct for each question. Once you successfully answer all post-test questions you will be able to view and/or print your certificate. For questions regarding the content of this activity, contact the accredited provider, CME@medscape.net. For technical assistance, contact CME@webmd.net. American Medical Association’s Physician’s Recognition Award (AMA PRA) credits are accepted in the US as evidence of participation in CME activities. For further information on this award, please refer to http://www.ama-assn.org/ama/pub/category/2922.html. The AMA has determined that physicians not licensed in the US who participate in this CME activity are eligible for AMA PRA Category 1 Credits™. Through agreements that the AMA has made with agencies in some countries, AMA PRA credit may be acceptable as evidence of participation in CME activities. If you are not licensed in the US, please complete the questions online, print the certificate and present it to your national medical association for review.
Article Title: Travel-associated Antimicrobial Drug–Resistant Nontyphoidal Salmonellae, 2004–2009
CME Questions
1. Your patient is a 30-year-old, otherwise healthy white woman thought to have nontyphoidal Salmonella (NTS). According to the study by Dr. Barlow and colleagues, which of the following statements about the epidemiology and clinical characteristics of NTS infections is correct?
A. Annual incidence of NTS exceeds 1 million in the United States and is estimated to be 98 million worldwide
B. Most NTS infections are transmitted by droplet spray
C. Most NTS infections require antibiotic treatment
D. Ampicillin and chloramphenicol are commonly prescribed in the United States for patients with NTS infection
2. According to the study by Dr. Barlow and colleagues, which of the following statements about antibiotic resistance associated with NTS cases in Oregon between 2004 and 2009 is correct?
A. Clinically important resistance (CIR) occurred in more than one quarter of isolates
B. The odds of acquiring CIR did not change from year to year
C. CIR was defined as decreased susceptibility to ampicillin, ceftriaxone, ciprofloxacin, gentamicin, or trimethoprim-sulfamethoxazole
D. Among cases with travel to East or Southeast Asia, increased resistance was attributable to a single serotype
3. According to the study by Dr. Barlow and colleagues, which of the following variables would most likely be positively associated with antibiotic resistance among NTS cases in Oregon between 2004 and 2009?
A. Outbreaks
B. Travel to Eastern Europe
C. Outpatient treatment
D. Travel to Southeast Asia
Activity Evaluation
|
1. The activity supported the learning objectives. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
2. The material was organized clearly for learning to occur. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
3. The content learned from this activity will impact my practice. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
|
4. The activity was presented objectively and free of commercial bias. |
||||
|
Strongly Disagree |
|
|
|
Strongly Agree |
|
1 |
2 |
3 |
4 |
5 |
Related Links
Table of Contents – Volume 20, Number 4—April 2014
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Emilio E. DeBess, Acute and Communicable Disease Prevention, Oregon Health Authority, 800 NE Oregon St., Portland, OR 97232, USA
Top