Volume 20, Number 6—June 2014
Dispatch
Dengue Virus Type 3, South Pacific Islands, 2013
Cite This Article
Citation for Media
Abstract
After an 18-year absence, dengue virus serotype 3 reemerged in the South Pacific Islands in 2013. Outbreaks in western (Solomon Islands) and eastern (French Polynesia) regions were caused by different genotypes. This finding suggested that immunity against dengue virus serotype, rather than virus genotype, was the principal determinant of reemergence.
In contrast to circulation in countries in Southeast Asia, where dengue is hyperendemic and ≤4 dengue virus (DENV) serotypes might co-circulate, it is rare for >1 DENV serotype to sustainably circulate in any South Pacific island country or territory. The pattern of single-serotype predominance has been historically observed in the entire South Pacific region (Figure 1); 1 serotype circulates for 4–5 years before being displaced by another serotype, i.e., DENV-3 (1989–1996), DENV-2 (1996–2000), DENV-1 (2001–2009), and DENV-4 (2008–2009) (1–5). Because DENV-3 had not circulated in the South Pacific Islands since 1996, a large nonimmune human population susceptible to infection with this serotype was present, and it had been suggested that this serotype would reemerge in ≈2012 (4).
In January 2013, an outbreak of DENV-3 infections was reported in the Solomon Islands (6). Two months later, DENV-1 and DENV-3 infections were identified in patients in French Polynesia. DENV-3 isolated in the Solomon Islands belonged to genotype I, and DENV-3 isolated in French Polynesia belonged to genotype III. This finding is an example of dengue outbreaks in the South Pacific Islands caused by introductions of multiple genotypes of the same DENV serotype into susceptible populations in the South Pacific Islands (2), in this instance, genotype I from Southeast Asia and genotype III from South America.
A dengue outbreak was reported by clinicians at the National Referral Hospital in Honiara, Solomon Islands, in January 2013. By July, >6,000 cases of suspected dengue had been reported, and 7 deaths were attributed to this outbreak. Serum samples from 3,141 patients were tested for DENV nonstructural protein 1 (NS1) and IgM against DENV (Dengue Duo, Standard Diagnostics Inc., Suwon, South Korea), and 1,220 (39%) samples were positive. DENV-3 was isolated by cell culture from 4 NS1-positive and IgM-negative samples by the World Health Organization Collaborating Centre for Arbovirus Reference and Research (Brisbane, Queensland, Australia). Ten additional NS1-positive samples were positive by reverse transcription PCR (RT-PCR) for DENV-3 when tested at the Institut Louis Malardé (Papeete, Tahiti, French Polynesia) (6).
In the first week of March 2013, DENV-3 was detected in French Polynesia in 2 patients in the same family cluster, 1 of whom had returned with a fever from Cayenne, French Guiana, 2 weeks earlier. Patients infected with DENV-1 were also detected in French Polynesia in February 2013. By July 2013, a total of 1,326 suspected dengue cases had been reported, of which 258 were laboratory confirmed by NS1 ELISA (Bio-Rad, Hercules, CA, USA), IgM ELISA (Bio-Rad), or RT-PCR. Serotype identification confirmed 170 DENV-1 infections, 73 DENV-3 infections, and 1 co-infection with both virus serotypes.
Envelope genes from DENV-3 isolated in the Solomon Islands and French Polynesia were amplified by using RT-PCR and sequenced as described (7–10). The 2 sequenced DENV-3 isolates from the Solomon Islands were obtained in February 2013 from patients who lived in the capital (Honiara), where the outbreak occurred. The 5 sequenced DENV-3 isolates from French Polynesia were obtained at different times and distances from the initially detected case. One virus (PF13/120213) was obtained in February from the initial family cluster in Mahina, Tahiti; 2 viruses (PF13/280213 and PF13/260313) were obtained in February and March from patients who lived in Mahina; 1 virus (PF13/040313) was obtained in March from a patient who lived in Papeete, Tahiti; and 1 virus (PF13/230413) was obtained in April from a patient who lived on Moorea Island.
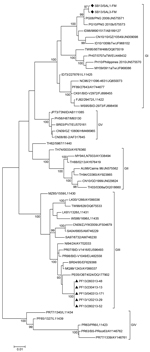
All sequenced DENV-3 strains isolated in the Solomon Islands belonged to genotype I, and all sequenced DENV-3 strains isolated in French Polynesia belonged to genotype III (Figure 2, Appendix). Phylogenetic analyses suggested that the recent DENV-3 outbreak in the Solomon Islands resulted from introduction of a strain from Indonesia that might have been circulating unrecognized in Papua New Guinea for 4–5 years. Previous reemergence of DENV-4 into the Pacific Islands might have resulted from introduction of a strain from Indonesia into the Solomon Islands (4,5,11).
In contrast, epidemiologic and phylogenetic evidence suggested that the outbreak of DENV-3 infection in French Polynesia resulted from introduction of virus from South America (French Guiana). Before 1990, most dengue outbreaks in the South Pacific region were caused by DENV that originated in Latin America. However, since that time most outbreaks have been caused by DENV from Southeast Asia (1–4,12,13). Recent transmission of DENV-3 from South America into French Polynesia is a timely reminder of the ongoing risk for DENV infection from a region in which DENV transmission is becoming hyperendemic. It remains to be seen whether, in the South Pacific, genotypes I and III of DENV-3 will maintain independent areas of transmission, they will co-circulate in the same localities, one will become dominant, or populations of the South Pacific Island nations are large enough for these sorts of dynamics to occur.
As anticipated by Li et al. (4), reemergence of DENV-3 in the South Pacific Islands in 2013 supports the contention that birth and immigration rates (1.2% and 1.1%, respectively, in French Polynesia in 2010) in these island nations provide sufficient susceptible hosts for DENV to cause an epidemic every 4–5 years and for a given DENV serotype to reappear every 12–15 years. Moreover, the finding that recent DENV-3 outbreaks were caused by multiple genotypes supports observations made during outbreaks of DENV-1 infection in 2001–2004 (2). These observations also suggest that the principal determinant of DENV reemergence in the South Pacific Islands is herd immunity, rather than the genotype of the DENV being introduced.
Genotype III of DENV-3 is believed to have originated in Sri Lanka, spread to eastern Africa in the mid-1980s, and then to South America in the mid-1990s (14). The discovery that DENV-3 genotype III has recently been introduced into the South Pacific Islands from South America indicates that this lineage might be the first DENV lineage to have circumnavigated the globe. Although the reemergence of DENV-3 in the South Pacific Islands in 2013 was anticipated, the reemergence of DENV-1 in New Caledonia in 2012 (15) and in French Polynesia in 2013 was not anticipated. Reports of co-circulation of multiple DENV serotypes are becoming more frequent in the South Pacific region and might reflect consequences of incomplete vector control (i.e., sufficient to prevent a major dengue outbreak but also preventing sufficient DENV transmission for development of effective herd immunity). Whether DENV-3 genotype I, genotype III, or both genotypes, will displace DENV-1 is not known.
Dr Cao-Lormeau is a research scientist at the Institut Louis Malardé, Papeete, Tahiti, French Polynesia. Her research focuses on the genetic evolution of dengue viruses in Pacific Island countries, in particular, dengue virus intrahost genetic diversity.
Acknowledgment
We thank the staff of the Ministry of Health, Honiara, Solomon Islands, for assisting in the provision of data; and the Medical Laboratory at the National Referral Hospital, Honiara, Solomon Islands, for assisting in provision of diagnostic material and data; and the laboratory staff at the Institut Louis Malardé, French Polynesia, for providing technical support.
References
- Chungue E, Deparis X, Murgue B. Dengue in French Polynesia: major features, surveillance, molecular epidemiology and current situation. Pac Health Dialog. 1998;5:154–62.
- A-Nuegoonpipat A. Berlioz-Arthaud A, Chow V, Endy T, Lowry K, Mai L, et al. Sustained transmission of dengue virus type 1 in the Pacific due to repeated introductions of different Asian strains. Virology. 2004;329:505–12.
- Descloux E, Cao-Lormeau VM, Roche C, de Lamballerie X. Dengue 1 diversity and microevolution, French Polynesia 2001–2006: connection with epidemiology and clinics. PLoS Negl Trop Dis. 2009;3:e493 . DOIPubMedGoogle Scholar
- Li DS, Liu W, Guigon A, Mostyn C, Grant R, Aaskov J. Rapid displacement of dengue virus type 1 by type 4, pacific region, 2007–2009. Emerg Infect Dis. 2010;16:123–5 . DOIPubMedGoogle Scholar
- Cao-Lormeau VM, Roche C, Aubry M, Teissier A, Lastere S, Daudens E, Recent emergence of dengue virus serotype 4 in French Polynesia results from multiple introductions from other South Pacific Islands. PLoS ONE. 2011;6:e29555. DOIPubMedGoogle Scholar
- Nogareda F, Joshua C, Sio A, Shortus M, Dalipanda T, Durski K, On-going outbreak of dengue serotype-3 in Solomon Islands: January–May 2013. Western Pac Surveill Response J. 2013;4:28–33.
- Aubry M, Roche C, Dupont-Rouzeyrol M, Aaskov J, Viallon J, Marfel M, Use of serum and blood samples on filter paper to improve the surveillance of Dengue in Pacific Island Countries. J Clin Virol. 2012;55:23–9 . DOIPubMedGoogle Scholar
- Liang H, Luo L, Yang Z, Di B, Bai Z, He P, Re-emergence of dengue virus type 3 in Canton, China, 2009–2010, associated with multiple introductions through different geographical routes. PLoS ONE. 2013;8:e55353 . DOIPubMedGoogle Scholar
- Delenda C, Staropoli I, Frenkiel MP, Cabanié L, Deubel V. Analysis of C-terminally truncated dengue 2 and dengue 3 virus envelope glycoproteins: processing in insect cells and immunogenic properties in mice. J Gen Virol. 1994;75:1569–78. DOIPubMedGoogle Scholar
- Lee E, Nestorowicz A, Marshall ID, Weir RC, Dalgarno L. Direct sequence analysis of amplified dengue virus genomic RNA from cultured cells, mosquitoes and mouse brain. J Virol Methods. 1992;37:275–88. DOIPubMedGoogle Scholar
- Shu PY, Su CL, Liao TL, Yang CF, Chang SF, Lin CC, Molecular characterization of dengue viruses imported into Taiwan during 2003–2007: geographic distribution and genotype shift. Am J Trop Med Hyg. 2009;80:1039–46 .PubMedGoogle Scholar
- Steel A, Gubler DJ, Bennett SN. Natural attenuation of dengue virus type-2 after a series of island outbreaks: a retrospective phylogenetic study of events in the South Pacific three decades ago. Virology. 2010;405:505–12. DOIPubMedGoogle Scholar
- Chungue E. Molecular epidemiology of dengue viruses. In: Saluzzo JF, Dbet B, editors. Factors in the emergence of arbovirus diseases. Amsterdam: Elsevier; 1997. p. 93–101.
- Messer WB, Gubler DJ, Harris E, Sivananthan K, de Silva AM. Emergence and global spread of a dengue serotype 3, subtype III virus. Emerg Infect Dis. 2003;9:800–9. DOIPubMedGoogle Scholar
- Western Pacific Regional Office of the World Health Organization. Dengue situation updates [cited 2013 Aug 21]. http://www.wpro.who.int/emerging_diseases/DengueSituationUpdates/en/index.html
Figures
Cite This ArticleTable of Contents – Volume 20, Number 6—June 2014
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Van-Mai Cao-Lormeau, Institut Louis Malardé, PO Box 30, 98713 Papeete, Tahiti, French Polynesia
Top