Volume 22, Number 4—April 2016
Dispatch
Neisseria meningitidis Serogroup X in Sub-Saharan Africa
Figure 1
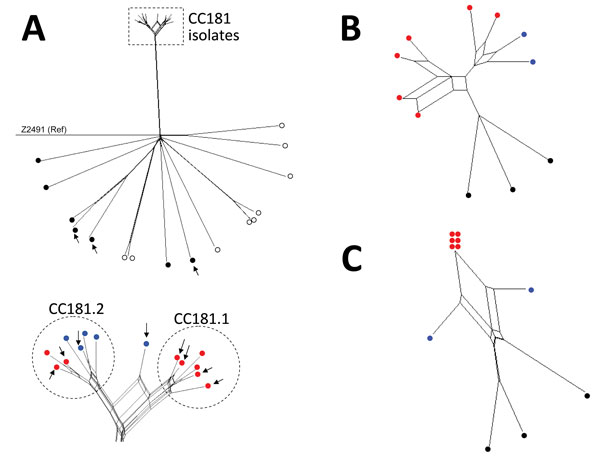
Figure 1. Neighbor-Net SplitsTree graphs generated using SplitsTree4 version 4.13.1 (http://www.splitstree.org) to visualize trees of Neisseria meningitidis serogroup X isolates. A) All 30 Neisseria meningitidis serogroup X isolates available on BIGSdb (11) were analyzed, including the 11 isolates from this study (8 from sub-Saharan Africa and 3 from France), 9 carriage isolates, 3 invasive isolates from Europe, 1 isolate from the United States, and 6 isolates from sub-Saharan African countries. Open (white) circles indicate carriage isolates; black circles indicate invasive isolates obtained outside of Africa; red circles indicate isolates obtained in sub-Saharan Africa since 2006; blue circles indicate isolates obtained from sub-Saharan Africa during the 1990s. Arrows indicate the 11 isolates obtained during this study. All isolates were compared with reference (Ref) meningococcal strain Z2491. The dashed rectangle indicates the cluster of all the clonal complex (CC) 181 isolates from sub-Saharan Africa (enlarged view at bottom of panel). B) The 11 isolates obtained during this study were compared for iron-acquisition genes (tpbA, tbpB, hpuA, hpuB, lbpA, lbpB, and hmbR). C) Genes of the 11 isolates obtained during this study compared with the 41 genes that differed between all the isolates obtained in Africa since 2006 and the isolate LNP13407 or the isolate LNP14354 (obtained during the 1990s).
References
- World Health Organization. Meningococcal disease control in countries of the African meningitis belt, 2013. Wkly Epidemiol Rec. 2014;89:206–14.PubMedGoogle Scholar
- Maiden MC, Bygraves JA, Feil E, Morelli G, Russell JE, Urwin R, Multilocus sequence typing: a portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci U S A. 1998;95:3140–5. DOIPubMedGoogle Scholar
- Etienne J, Sperber G, Adamou A, Picq JJ. Epidemiological notes: meningococcal meningitis of serogroup X in Niamey (Niger) [in French]. Med Trop (Mars). 1990;50:227–9.PubMedGoogle Scholar
- Boisier P, Nicolas P, Djibo S, Taha MK, Jeanne I, Mainassara HB, Meningococcal meningitis: unprecedented incidence of serogroup X–related cases in 2006 in Niger. Clin Infect Dis. 2007;44:657–63. DOIPubMedGoogle Scholar
- Hong E, Giuliani MM, Deghmane AE, Comanducci M, Brunelli B, Dull P, Could the multicomponent meningococcal serogroup B vaccine (4CMenB) control Neisseria meningitidis capsular group X outbreaks in Africa? Vaccine. 2013;31:1113–6. DOIPubMedGoogle Scholar
- European Centre for Disease Prevention and Control. Annual epidemiological report 2012. Reporting on 2010 surveillance data and 2011 epidemic intelligence data. Stockholm: The Centre; 2013. p. 168–72.
- Maiden MC, van Rensburg MJ, Bray JE, Earle SG, Ford SA, Jolley KA, MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol. 2013;11:728–36. DOIPubMedGoogle Scholar
- Jolley KA, Hill DM, Bratcher HB, Harrison OB, Feavers IM, Parkhill J, Resolution of a meningococcal disease outbreak from whole-genome sequence data with rapid Web-based analysis methods. J Clin Microbiol. 2012;50:3046–53. DOIPubMedGoogle Scholar
- Kellogg DS Jr, Peacock WL Jr, Deacon WE, Brown L, Pirkle DI. Neisseria gonorrhoeae. I. Virulence genetically linked to clonal variation. J Bacteriol. 1963;85:1274–9.PubMedGoogle Scholar
- Veyrier FJ, Hong E, Deghmane AE, Taha MK. Draft genome sequence of a Neisseria meningitidis serogroup C isolate of sequence type 11 linked to an outbreak among men who have sex with men. Genome Announc. 2013;1:pii:e00795-13.
- Jolley KA, Maiden MC. BIGSdb: scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics. 2010;11:595. DOIPubMedGoogle Scholar
- Harrison OB, Bennett JS, Derrick JP, Maiden MC, Bayliss CD. Distribution and diversity of the haemoglobin–haptoglobin iron-acquisition systems in pathogenic and non-pathogenic Neisseria. Microbiology. 2013;159:1920–30. DOIPubMedGoogle Scholar
- Szatanik M, Hong E, Ruckly C, Ledroit M, Giorgini D, Jopek K, Experimental meningococcal sepsis in congenic transgenic mice expressing human transferrin. PLoS ONE. 2011;6:e22210. DOIPubMedGoogle Scholar
- Guiddir T, Deghmane AE, Giorgini D, Taha MK. Lipocalin 2 in cerebrospinal fluid as a marker of acute bacterial meningitis. BMC Infect Dis. 2014;14:276. DOIPubMedGoogle Scholar
- Rune Andersen S, Kolberg J, Hoiby EA, Namork E, Caugant DA, Oddvar Froholm L, Lipopolysaccharide heterogeneity and escape mechanisms of Neisseria meningitidis: possible consequences for vaccine development. Microb Pathog. 1997;23:139–55. DOIPubMedGoogle Scholar
1Current affiliation: Ministry of Health Panama and Instituto de Ciencias Médicas, Las Tablas, Panama.