Volume 23, Number 12—December 2017
Research Letter
High Abundance and Genetic Variability of Atypical Porcine Pestivirus in Pigs from Europe and Asia
Cite This Article
Citation for Media
Abstract
Atypical porcine pestivirus (APPV) was recently reported to be associated with neurologic disorders in newborn piglets. Investigations of 1,460 serum samples of apparently healthy pigs from different parts of Europe and Asia demonstrate a geographically wide distribution of genetically highly variable APPV and high APPV genome and antibody detection rates.
Pestiviruses are highly variable RNA viruses within the family Flaviviridae. The recently discovered atypical porcine pestivirus (APPV) is capable of inducing neurologic disorder in its host, like other pathogens of this family (e.g., tick-borne encephalitis virus, Zika virus). Several recently published reports demonstrate that APPV is a prominent cause of virus-induced congenital tremor in pigs (1–4). Serum samples from healthy but viremic animals can induce birth of clinically affected offspring when experimentally transferred to sows during gestation (1,2). So far, APPV presence has been reported from the United States, some countries within Europe, and China (2,4–7). The economic relevance of APPV-related losses in pig production remains to be determined, but estimation revealed a drop in reproductive performance of 10% in an affected farm (4). Early data from the United States and Germany suggested a relatively high abundance (2.4%–22%) of APPV genomes in apparently healthy pigs (3,6,8) that likely play an important epidemiologic role as virus carriers. We investigated APPV genome and antibody abundance in healthy pigs from different parts of Europe and Asia. To provide insight into genetic diversity of this novel pathogen,
We tested 1,460 serum samples from Germany, Great Britain, Italy, Serbia, Switzerland, mainland China, and Taiwan by using an APPV-specific PCR and an indirect APPV ELISA, as previously described (3,9). The sample set comprised 369 serum samples from Germany that were previously screened for the presence of APPV genomes (3). For our study, serum samples were taken from apparently healthy pigs within the framework of national veterinary health management in concordance with national legal and ethical regulations.
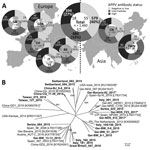
For APPV genome detection, we conducted a PCR targeting the nonstructural protein (NS) 3 encoding region and confirmed specificity of amplification by gel electrophoresis (3). We detected APPV genomes in domestic pigs from all investigated regions. In total, 130 (8.9%) of the 1,460 tested samples were APPV genome positive (Figure). Genome detection rates ranged from 2.3% (2/86 samples from Great Britain) to 17.5% (35/200 samples from Italy). Moreover, we demonstrated that APPV was abundant in Asia; we detected the APPV genome in 11/219 samples (5%) from mainland China and 22/200 samples (11%) from Taiwan.
We used individual samples with high genome loads to generate amplicons in a seminested PCR and subsequently performed Sanger sequencing (FlexiRun, LGC Genomics, Berlin, Germany). We generated 20 different APPV NS3 sequences from apparently healthy pigs of all countries (sequences deposited into GenBank under accession nos. MF279213–32). Genetic differences reflect geographic origin to a low degree (Figure); genetic variability even within a country is remarkably high (e.g., Germany and Italy). Genetic analyses including sequence data obtained from samples of diseased piglets revealed no correlation of pathogenicity with certain genetic variants (Figure).
In addition to the NS3 fragments, we determined APPV complete coding sequences (CDS) from 1 sample from a healthy pig from China (deposited into GenBank under accession no. MF167292) and 2 samples (accession nos. MF167290 and MF167291) obtained from pigs during outbreaks of congenital tremors in Germany (3,9). We performed next-generation sequencing as previously described (3). The outbreak isolates from Germany were almost identical (0.2% genetic distance) and were similar to an isolate from northern Germany (accession no. LT594521). The APPV from China had a unique 93-nt deletion in the NS5A encoding region. A similar genome (97.9% identity) is lacking this deletion (Guangxi Province; accession no. KY652092). The biological relevance of the deletion remains elusive, but classical pestiviruses show a remarkable genetic tolerance in this genomic region (10). The sequence data we obtained reveal a high genetic variability (up to 21% genetic distance), which is comparable to that of classical swine fever virus (Technical Appendix).
We applied an indirect APPV Erns antibody ELISA, as described (9), and classified the serologic status into low (S/p≤0.5), intermediate (0.5<S/p<1.0), or highly (S/p≥1.0) reactive. Due to the lack of reference material and a standard assay, we could not determine test parameters (e.g., sensitivity, specificity) at this stage. Nevertheless, the ELISA was a valuable tool for detecting seroconversion in infected pigs; >60% of the animals showed intermediate to high reactivity in the antibody ELISA (Figure), which is in line with high APPV genome detection rates. We detected similar frequencies of APPV antibody–positive samples for each region, independent of the genome detection rates (Figure). We found most of the viral genomes (≈86%) in samples with intermediate or low antibody status; few (≈14%) of the highly antibody-positive animals were viremic at the same time. This observation might indicate a degree of protection provided by the induced antibodies. Of the 40% of the pigs that were antibody negative, 10% were genome positive; possible explanations are that serum samples were taken either from acutely infected animals before induction of a detectable antibody response or from persistently infected animals lacking a specific humoral immune response due to a specifically acquired immunotolerance, a well-known consequence of intrauterine pestivirus infections.
Our findings indicate that the recently discovered APPV is abundant on several continents. APPV must be regarded as a pig pathogen of likely worldwide relevance.
Dr. Postel is a veterinarian and the head of the Laboratory for Molecular Biology of the European Union and World Organisation for Animal Health Reference Laboratory for Classical Swine Fever at the Institute of Virology of the University of Veterinary Medicine in Hannover, Germany. Research interests are molecular evolution and pathogenesis of pestiviruses and related viruses, characterization of novel pestivirus isolates, and diagnosis and control of classical swine fever.
Acknowledgment
We thank Tanja Frey and Birgitta Polley; the serum bank of the Federal Food Safety and Veterinary Office, Bern, Switzerland; the Animal and Plant Health Agency, Weybridge, UK; and the animal health service, Baden-Württemberg, Germany, for providing serum samples. We thank Daniela Indenbirken and Malik Alawi for their support in library preparation and analysis of next-generation sequencing data, and Elena Gräf for her support in Sanger sequencing. This work in part was financially supported by Boehringer Ingelheim.
References
- Arruda BL, Arruda PH, Magstadt DR, Schwartz KJ, Dohlman T, Schleining JA, et al. Identification of a divergent lineage porcine pestivirus in nursing piglets with congenital tremors and reproduction of disease following experimental inoculation. PLoS One. 2016;11:e0150104. DOIPubMedGoogle Scholar
- de Groof A, Deijs M, Guelen L, van Grinsven L, van Os-Galdos L, Vogels W, et al. Atypical porcine pestivirus: a possible cause of congenital tremor type A-II in newborn piglets. Viruses. 2016;8:271. DOIPubMedGoogle Scholar
- Postel A, Hansmann F, Baechlein C, Fischer N, Alawi M, Grundhoff A, et al. Presence of atypical porcine pestivirus (APPV) genomes in newborn piglets correlates with congenital tremor. Sci Rep. 2016;6:27735. DOIPubMedGoogle Scholar
- Schwarz L, Riedel C, Högler S, Sinn LJ, Voglmayr T, Wöchtl B, et al. Congenital infection with atypical porcine pestivirus (APPV) is associated with disease and viral persistence. Vet Res (Faisalabad). 2017;48:1. DOIPubMedGoogle Scholar
- British Veterinary Association. Congenital tremor associated with atypical porcine pestivirus. Vet Rec. 2017;180:42–3. DOIPubMedGoogle Scholar
- Hause BM, Collin EA, Peddireddi L, Yuan F, Chen Z, Hesse RA, et al. Discovery of a novel putative atypical porcine pestivirus in pigs in the USA. J Gen Virol. 2015;96:2994–8. DOIPubMedGoogle Scholar
- Yuan J, Han Z, Li J, Huang Y, Yang J, Ding H, et al. Atypical porcine pestivirus as a novel type of pestivirus in pigs in China. Front Microbiol. 2017;8:862. DOIPubMedGoogle Scholar
- Beer M, Wernike K, Dräger C, Höper D, Pohlmann A, Bergermann C, et al. High prevalence of highly variable atypical porcine pestiviruses found in Germany. Transbound Emerg Dis. 2016.PubMedGoogle Scholar
- Postel A, Meyer D, Petrov A, Becher P. Recent emergence of a novel porcine pestivirus: interference with classical swine fever diagnosis? Emerg Microbes Infect. 2017;6:e19. DOIPubMedGoogle Scholar
- Isken O, Langerwisch U, Schönherr R, Lamp B, Schröder K, Duden R, et al. Functional characterization of bovine viral diarrhea virus nonstructural protein 5A by reverse genetic analysis and live cell imaging. J Virol. 2014;88:82–98. DOIPubMedGoogle Scholar
Figure
Cite This ArticleTable of Contents – Volume 23, Number 12—December 2017
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Paul Becher, EU and OIE Reference Laboratory for Classical Swine Fever, Institute for Virology, University of Veterinary Medicine Hannover, Buenteweg 17, 30559 Hannover, Germany
Top